Mycoplasma pulmonis
A Microbial Biorealm page on the genus Mycoplasma pulmonis
Classification
Higher order taxa
Bacteria; Firmicutes; Mollicutes; Mycoplasmataceae; Mycoplasma
Species
Mycoplasma pulmonis
Description and significance
Mycoplasma was first isolated in 1898 from a species that is a bovine pathogen. M.pulmonis falls under genus mycoplasma that are easily distinguished from other bacteria due to their small size (0.3-0.8 micron in diameter) and lack of cell wall. Mycoplasma is believed to descend from gram-positive bacteria. The organism is covered by lipoprotein based cytoplasmic membrane. Its cell membrane is rich in protein components that consists of structurally adaptive lipoproteins used to invade host immune system, attach to the host cells, and pathogenic invasion. The primary habitats of human and animal mycoplasma are in mucous membranes of the respiratory and urogenital tracts, eyes, mammary glands and the joints.
They are the smallest cells known that are capable of self replication. Also, lack of rigid cell wall allows them to have plasticity so mycoplasma don't have an exact cell shape. This allows them to pass through cell's porin and attack the cell. Also, lack of a cell wall causes the organism to be a difficult target for antibiotics so it is harder to treat the disease caused by this bacteria. M.pulmonis has its genome completely sequenced because they serve as model for minimal cell structure, they are also very small and cause pathogenicity in other organism. Studying the bacteria increases knowledge on the mechanism of their attack to host cell (pathophysiology), evolution, and studies to produce vaccine/ treatment for the disease.
Genome structure
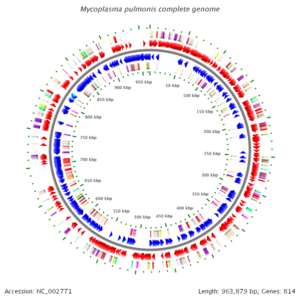
Mycoplasma pulmonis is among the first organism to have its genome sequenced due to its small size. Mycoplasma has the smallest of the prokaryotic genomes and it lacks complex gene-regulatory systems. M. pulmonis has a single circular 963,879 bp chromosome with a G + C content of 26.6 mol%. The genome contains 782 putative coding sequences (CDSs)with 482 CDS that could have function assigned and 92 CDS that have sequence matched to hypothetical protein. The genome also have a single set of rRNA genes and 29 tRNAs genes.
The length of the tandem repeat region of the variable surface antigen (Vsa) protein determines the biofilm formation of M.pulmonis. When Vsa proteins are short with few tandem repeats, they form a biofilm that attaches the polystyrene and glass. However, when Vsa protein are long with many tandem repeats, M. pulmonis are found floating on the medium as microcolonies.
An interesting feature of M. pulmonis is the actin like structure used for gliding and attachment to host cell with contributions from Protein Gli521 and Gli349. Protein Gli521 has its gene encoded downstream of the Gli349 gene and it is composed of 4,738 amino acids. The protein generates force for movement during gliding. As for Protein Gli349, it assists in binding of the bacteria to host cells.
M. pulmonis also have a unique coding in their genome, TGA encodes for trytophan in Mycoplasma but it is a stop codon for other eubacteria. Surprisingly, several of the genes previously reported to be essential for a self-replicating minimal cell are missing in the M. pulmonis genome although this genome is larger than the other mycoplasma genomes fully sequenced until now.
Cell structure and metabolism
M. pulmonis have a minimal cell structure due to loss of metabolic genes during evolution. They are pathogenic bacteria so they need a mechanism or structure to help them survive in the host cell. The organism has actin like structure that help them to glide and bind onto the surface of the host target cell. Gliding motility is also related to their ability to form microcolonies on the cell. Other than the actin like structure for motility, they also have cholesterols that function in transport and cell wall stability. M. pulmonis also has a stabilizer in their cell membrane to prevent osmotic shock.
The majority of mycoplasmas are deficient in genes coding for components of intermediary and energy metabolism and thus generate ATP using electron transport phosphorylation or glycolysis. They are facultative aerobes because they can live in environment with or without oxygen present. Also, they lost most of their genes involved in amino acid and cofactor biosynthesis, synthesis of the cell wall, lipid metabolism, and cellular processes during evolution. Therefore they serves as model for minimal structure needed for cell metabolism. The bacteria consists of circular chromosome, plasma membrane and ribosomes.
Pathology
Mycoplasma pulmonis is the agent of murine respiratory mycoplasmosis (MRM), a pneumonia-like slow progressing disease that sometimes does not develop clinical symptoms. It can also cause genital infections which are associated with conditions ranging from decreased fertility to fetal wastage both in rodents and humans. It is hypothesized that genetic predisposition and environment are the determinant for the disease caused by M. pulmonis.
The predicted virulence factors for M. pulmonis are hemolysin, secreted nucleases and a glyco-protease. They do not lyse the erythrocytes but instead they rapidly lyse in erythrocytes with cytoskeletal deficiency because it increases diffusion for glycophorin. Trypsin on the cell of M. pulmonis plays part in hemaglutination of the red blood cells (RBCs) because it cleaves the glycophorin on the membrane of RBCs. Bovin serum albumin also thought to assist in the hemaglutination of the erythrocytes.
The species is also capable of gliding motility in which the mechanism is still unknown. However, it is hypothesized that they glide over the surface by employing actin-like cytoskeletal components and motility proteins specific for the microbial group.
Application to Biotechnology
The complete genome sequence of the organism enables it to serve as the model organism for bacterial evolution related to pathogenicity. It is also useful as model for studying mycoplasmal infections.
Current Research
1. Evidence for type III restriction and modification systems in Mycoplasma pulmonis. J Bacteriol. 2007 Mar;189(6):2197-202. Epub 2007 Jan 5.
Mycoplasma pulmonis is predicted to code for type III restriction and modification (R-M) enzymes.
2. A stochastic mechanism for biofilm formation by Mycoplasma pulmonis.
J Bacteriol. 2007 Mar;189(5):1905-13. Epub 2006 Dec 1.
Biofilm formation of M. pulmonis depends on the length of Vsc protein.
3. Neutrophil histamine contributes to inflammation in Mycoplasma pneumonia. J Exp Med. 2006 Dec 25;203(13):2907-17. Epub 2006 Dec 11. Mycoplasma directly stimulated histamine production by naive neutrophils and strongly upregulated mRNA encoding histidine decarboxylase, the rate-limiting enzyme in histamine synthesis.
References
1.Bischoff, Joe, Mikhail Domrachev, Scott Federhen, Carol Hotton, Detlef Leipe, Vladimir Soussov, Richard Sternberg, and Sean Turner. "Mycoplasma Pulmonis." NCBI. 1 May 2007 <http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&id=2107&lvl=3&p=mapview&p=has_linkout&p=blast_url&p=genome_blast&keep1&srchmode=1&unlock&lin=s
2.Wl, Simmons, Bolland Jr, Daubenspeck Jm, and Dybviq K. "A Stochastic Mechanism for Biofilm Formation by Mycoplasma Pulmonis." os. PubMed. 2 May 2007 <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=pubmed&cmd=Retrieve&dopt=AbstractPlus&list_uids=17142389&query_hl=5&itool=pubmed_docsum>
3."Mycoplasma Pulmonis UAB CTIP Project At Genoscope." NCBI. 2 May 2007 <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=genomeprj&cmd=Retrieve&dopt=Overview&list_uids=100>.
4.Chambaud, Isabelle, Roland Heilig, Stéphane Ferris, Valérie Barbe, and Delphine Samson. "The Complete Genome Sequence of the Murine Respiratory Pathogen Mycoplasma Pulmonis." PubMedCentral os 29.10 (2001): 2145-2153. PubMed. 2 May 2007 <http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=55444>.
5.Minion, F C., and J D. Goguen. "Trypsin-Sensitive, Bovine Serum Albumin-Dependent Hemolysis Activity in Mycoplasma Pulmonis." Infection and Immunity 49 (1985): 440-442. PubMed Central. 4 June 2007 <http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=262037>.
6. M, Miyata, and Yamamoto H. "Gliding Mutants of Mycoplasma Mobile: Relationships Between Motility and Cell Morphology, Cell Adhesion and Microcolony Formation." Microbiology 146 (2000): 1311-1320. PubMed Central. 4 June 2007 <http://www.ncbi.nlm.nih.gov/sites/entrez?Db=pubmed&Cmd=ShowDetailView&TermToSearch=10846210&ordinalpos=1&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_RVAbstractPlus>.
7. S, Seto, Uenoyama A, and Miyata M. "Identification of a 521-Kilodalton Protein (Gli521) Involved in Force Generation or Force Transmission for Mycoplasma Mobile Gliding." Journal of Bacteriology 187 (2005): 3502-3510. PubMed Central. 4 June 2007 <http://www.ncbi.nlm.nih.gov/sites/entrez?Db=pubmed&Cmd=ShowDetailView&TermToSearch=15866938&ordinalpos=1&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_RVDocSum>.
Edited by Grace Wanto
KMG