Pythium oligandrum
Classification
Kingdom: Chromalveolata
Phylum: Heterokontophyta
Class: Oomycetes
Order: Pythiales
Family: Pythiaceae
Species
|
NCBI: Taxonomy [1] |
Pythium oligandrum
Description and Significance
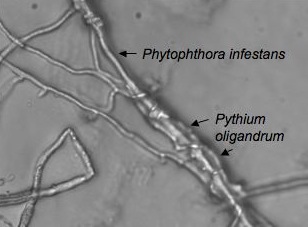
Oomycetes form a diverse group of fungus-like eukaryotic microorganisms, in which modern research evidently proved that they are more closely related to heterkonts than fungi, as once believed. These microorganisms are classified as water moulds and encompass a large host range of organisms such as fish, crustaceans, vertebrates and plants. Specifically within that class are parasitic and mycoparasitic species such as Phytophthora infestans and P. oligandrum, respectively. The multitudes of saprophytic oomycetes inhabit primarily aquatic and moist soil habitats. They have a positive effect on nutrient cycling through their key role in the decomposition and recycling of organic matter (8). Interestingly, P. oligandrum can curb the pathogenic effects of parasitic oomycetes such as Phytophthora infestans.
Furthermore, P. oligandrum is a nonpathogenic oomycete that can be utilized as a biocontrol agent to provide damping-off and root disease resistance to various soil born plant pathogens. It also ability to cannibalize other oomycetes such as Fusarium and Phytophthora, making it a crucial defense for the plant roots. Furthermore, this ability can be utilized to fight off fungal infections such as ringworm and athlete's foot.
Genome Structure
P. oligandrum has not yet been sequenced. Organisms within in the Oomycete class are known to have a wide range in their genome size, with the Pythium genus having the smallest among the oomycete(6). Little is known regarding the organization of the genome due to limited genome sequencing within the class. It is interesting to note that although oomycetes are considered to by eukaryotes, keystone sequences like the TATA box have been missing in some oomycete gene sequences (7). The chromosome number is not known in P. oligandrum, but it is estimated to have somewhere around 18 (8).
Cell Structure, Metabolism and Life Cycle
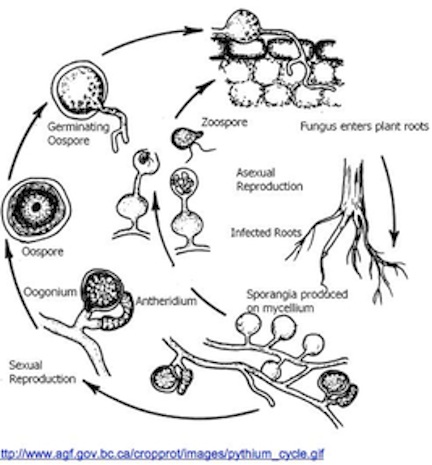
Pythium oligandrum has two distinctive life cycles. One of which is asexual and the other is sexual. In the asexual lifecycle, P. oligandrum exists as a zoospore. A zoospore is a flagellated cell containing genetic material. The zoospore of P. oligandrum is diploid and contains the full genome. In the case of P. oligandrum, the zoospore is an important life stage because of the zoospores ability to move through water via its flagella. When conditions are suitable, the zoospore undergoes mitotic divisions and matures. Pythium oligandrum exists in its sexual life stage about 20% of the time(1). This life cycle is characterized by thick-walled oospores. Oospores are a result of the fertilization of oogonium by antheridium, which is the male sex organ of the organism. The maturation of the oospores leads to the production of mycelium and a mature organism capable or reproduction and completing the cycle. Oospores are characterized by their thick cell walls and are relatively large cells.
Pythium oligandrum is a eukaryotic organism. They were once thought to be a type of fungi, but they lack chitin and are made of cellulose, hence are currently categorized as a protist. Their oospores are spiny walled, which distinguished them from other species of Pythium. Their mature stage is multi-cellular and filamentous. P. oligandrum is found in the rhizosphere of various plant roots. P. oligandrum is parastic and heterotrophic. It get its nutrients from other organisms by lysing them and "sucking out" what it needs for its biochemistry. In addition, it was wrongly classified as a fungi because it is a decomposer and undergoes the process of nitrification, in which nitrogen compounds undergo oxidation.
NH3 + O2 + H2O → NO3− + 5H+ + 4e−
Ecology and Pathogenesis
P. oligandrum exemplifies a symbiotic relationship within the rhizosphere of various plant roots such as tomatoes and sugar beet plants. This relationship consists of P. oligandrum inhibiting the pathogenic capabilities of various harmful oomycetes and so maintains the plants roots from foreign attack. In return P. oligandrum will obtain the necessary nutrients from the plant roots to thrive. Furthermore, P. oligandrum is considered an “aggressive mycoparasite” that forms parasitic relationships with plants and other microorganisms. For the latter reason, it can actually be beneficial to plants because it can provide as a sort of defense from other plant pathogens esp. fungi. For this reason, P. oligandrum is a potential biocontrol agent. However, it's aggressive cannibalistic behavior can cause damping-off. This can potentially kill seeds because the organism attacks the first roots formed during germination. However, because it infects fungi, it can ruin mushroom crops such as those crops of the common mushroom(2) (Agaricus bisporus)
References
1. "Review report for the active substance Pythium oligandrum M1." EUROPEAN COMMISSION HEALTH & CONSUMERS DIRECTORATE-GENERAL. Web. <http://ec.europa.eu/food/plant/protection/evaluation/existactive/M1.pdf>.
2. Godfrey et al. 2003. Identification of Pythium oligandrum using species-specific ITS rDNA PCR oligonucleotides. The British Mycological Society. 107(7):790-796. Science Direct. 21. April 2011.
3. Le Floch, G., Tambong, J., Vallance, J., Tirilly, Y., Lévesque, A. and Rey, P. (2007), Rhizosphere persistence of three Pythium oligandrum strains in tomato soilless culture assessed by DNA macroarray and real-time PCR. FEMS Microbiology Ecology, 61: 317–326. doi: 10.1111/j.1574-6941.2007.00348.x
4. Holmes, K. A., Nayagam, S. D. and Craig, G. D. (1998), Factors affecting the control of Pythium ultimum damping-off of sugar beet by Pythium oligandrum. Plant Pathology, 47: 516–522. doi: 10.1046/j.1365-3059.1998.00253.x
5. Reproductive life cycle picture: British Columbia: ministry of agriculture . Web. 21 Apr 2011. <http://www.gov.bc.ca/agri/>.
6. Kamoun, Sophien. "Molecular Genetics of Pathogenic Oomycetes." Eukaryot Cell 2. (2003): 191-199. Web. 23 Apr 2011.
7. Pieterse, C. M. J., P. van West, H. M. Verbakel, P. W. H. M. Brasse, G. C. M. van den Berg Velthuis, and F. Govers as cited in: Kamoun, Sophien. "Molecular Genetics of Pathogenic Oomycetes." Eukaryot Cell 2. (2003): 191-199. Web. 23 Apr 2011.
8. Martin, F.N as cited in Kamoun, Sophien. "Molecular Genetics of Pathogenic Oomycetes." Eukaryot Cell 2. (2003): 191-199. Web. 23 Apr 2011.
9. Falih, A. and Wainwright, M. (1995), Nitrification in vitro by a range of filamentous fungi and yeasts. Letters in Applied Microbiology, 21: 18–19. doi: 10.1111/j.1472-765X.1995.tb00996.x
Author
Page authored by Thomas Fox and Fuad Hajiabdi, student of Prof. Jay Lennon at Michigan State University.
<-- Do not remove this line-->