Gemmata obscuriglobus
Classification
Prokarya; Planctomyces; Planctomycetia; Planctomycetales; Planctomycetaceae; Gemmata; Gemmata obscuriblobus
Species
Gemmata obscuriglobus
|
The National Center for Biotechnology Information taxonomy page for Gemmata obscuriglobus: [1] |
Gemmata obscuriglobus is a spherical budding bacteria isolated from a freshwater dam in Queensland, Australia. This bacteria is interesting because it exhibits several structures and cellular processes that are typically reserved for only eukaryotic cells, yet are classified as prokaryotes. Some of these interesting features include a membrane-bounded nucleoid, the ability to perform endocytosis on fully folded proteins, it's ability to reproduce by a unique way of budding, and its lack of a cell wall composed of peptidoglycan.
Genome Structure
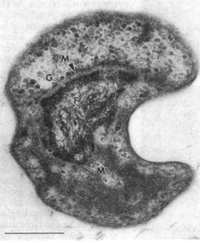
The genome of Gemmata obscuriglobus is made up of DNA contained in the fibrillar, membrane-bounded nucleoid which is then surrounded by a another membrane as stated before. The genome consists of 9.16 million base pairs consisting of 8,080 genes that produce 7,989 proteins discovered thus far. The DNA does not contain chromosomes, but a circular piece of DNA that is characteristic to most all prokaryotes.
text text text text text
Cell Structure, Metabolism and Life Cycle
Cellular Structure
The cellular structure of the Gemmata obscuriglobus species is unique from other bacteria species in that it has many features and structures that are only seen in eukaryotes, yet is classified as a prokaryote. The most distinguishing feature of this species and other related species is its membrane-bounded nucleoid. The DNA is contained in a fibrillar nucleoid area enclosed in a membrane, which is then enclosed in an outer membrane. Bacteria species have been found to have some associations between their DNA and membranes, but never as complete or complex as this species and related species. The dual-membrane structure surrounding the nucleoid is termed "nuclear body" signifying its difference from the eukaryotic nucleus. This species can also perform endocytosis where the cell uptakes fully folded proteins into the cytoplasm. This process has always thought to have been restricted to eukaryotes. Bacteria have means of taking in material from the extracellular environment, but not molecules as large as fully folded proteins nor by the means of transport. Gemmata obscuriglobus have receptors on their membrane that mediate this in an energy-dependent process. These receptors are non-specific. The ability to perform receptor mediated endocytosis is traceable back to the eukaryotic ancestor, LECA. Another amazing feature about this species is its complete lack of peptidoglycan and a cell wall. This is unusual for a prokaryote, because a cell wall is so crucial in support, reproduction, and protection in most bacteria species. For support, Gemmata obscuriglobus uses glutamate. This lack of peptidoglycan is responsible for G. obscuriglobus' resistance to a broad spectrum of antibiotics. This is because most antibiotics work to either prevent or damage the peptidoglycan layer in bacteria.
Metabolism
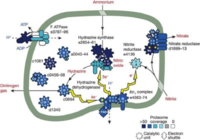
Gemmata obscuriglobus is an anaerobic ammonium oxidizer and metabolizes ammonium like other anammox bacteria.
Life Cycle
The life cycle of this species is also unique in its own nature. It reproduces by budding, a process in which the daughter cell grows from the mother cell unequally, and breaks off when it becomes mature. Budding is common in some eukaryotes, such as many species of fungi, but not typical of bacteria nor archaea. The mechanism by which this species undergoes budding is also very different. DNA transfer from parent to offspring begins shortly after formation of the daughter bud. DNA is transferred via a connecting passage that exists between the two until maturation and then separation occurs. The young bud's DNA is naked at this point, but is soon bound by membrane derived from both the mother and daughter cell. After separation the daughter can then live freely and undergo budding itself. Each mature cell can bud multiple times in its lifetime. An important difference between this species and other bacteria apart from the mechanism is the absence of the FtsZ gene. Most bacteria reproduce by binary fission in which the cell duplicates its genome, doubles in size, and splits into two cells equal in size at the septum with help of the FtsZ protein.
Ecology, Pathogenesis, and Evolutionary Importance
Organisms within the phyla Planctomyces are aquatic and can be found in marine water, fresh water, brackish water, and even acidic bog water. The first time Gemmata obscuriglobus was isolated was in 1991 from a freshwater dam in Queensland, Australia. Being that they are ammonium oxidizers, they convert ammonium and nitrite to dinitrogen gas and water completing the loop of the nitrogen cycle. As much as 50% of the oceans nitrogen gas productivity is a result of this microbial process, and many of the species that do this are within the phyla Planctomyces.
This organism is nonpathogenic. However, Planctomycetes are most closely related to the phyla Chlamydiae and Verrucomicrobia which contain species that cause disease in humans.
With all these similarities to eukaryotes, how do we know they are prokaryotes? At first researchers did not know where to classify this organism on the tree of life, but after genomic analysis of the 16S rRNA gene, it has been confirmed a prokaryote. This signifies that all these eukaryotic features have evolved independently from their eukaryotic counterparts and could possibly give us better insight to the evolution of eukaryotes.
References
Cayrou C, Raoult D, Drancourt M. “Broad-spectrum antibiotic resistance of Planctomycetes organisms determined by Etest”. Journal of Antimicrobial Chemotherepy. 2010. Vol. 65 no. 10 pp. 2119-2122.
Fuchsman CA, Rocap G. “Whole-genome reciprocal BLAST analysis reveals that planctomycetes do not share an unusually large number of genes with Eukarya and Archaea”. Applied and environmental microbiology. 2006. vol. 72 no. 10 p. 6841-6844.
Fuerst JA, Sagulenko E. “Protein uptake by bacteria: An endocytosis-like process in the planctomycete Gemmata obscuriglobus”. Communicative & Integrative Biology. 2010. 3(6). p. 572-575.
Fuerst JA, Sagulenko E. "Keys to eukaryality: Planctomycetes and ancesteral evolution of cellular complexity". Frontiers in Microbiology. 2012. Vol. 3 no. 167.
Fuerst JA, Webb RI. “Membrane-bounded nucleoid in the eubacterium Gemmata obscuriglobus”. Proceedings of the Nation Academy of Sciences of the United States of America. 1991. Vol. 88, pp. 8184-8188.
Lee KC, Webb RI, Fuerst JA. “The cell cycle of the planctomycete Gemmata obscuriglobus with respect to cell compartmentalization” BMC Cell Biology. 2009. Vol. 10 .
Lonhienne TG, Sagulenko E, Webb RI, Lee KC, Franke J, Devos DP, Nouwens A, Carroll BJ, Fuerst JA”. Endocytosis-like protein uptake in the bacterium Gemmata obscuriglobus.” Proceedings of the Nation Academy of Sciences of the United States of America. 2010. Vol. 107 no. 29 p. 12883–12888.
Pearson A, Budin M, Brocks JJ. “Phylogenetic and biochemical evidence for sterol synthesis in the bacterium Gemmata obscuriglobus”. Proceedings of the Nation Academy of Sciences of the United States of America. 2003. Vol. 100 no. 26 pp. 15352-15357.
Strous M, Pelletier E, et al. “Deciphering the evolution and metabolism of an anammox bacterium from a community genome”. Nature. 2006. Vol. 440 pp. 790-794.
Author
Page authored by Michael Bowe and Ryan Quenneville, students of Profs. Ned Walker and Kazem Kashefi at Michigan State University.
<-- Do not remove this line-->