Human Chromosomal Integration of Latent State Human Herpes Virus 6 (HHV-6)
By: Kerri-Lynn Conrad
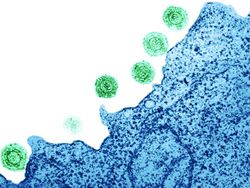
Introduction
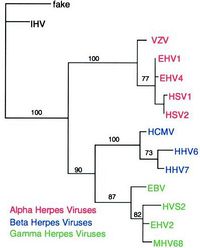
Human herpesvirus 6 (HHV-6) was first discovered in 1986 during a study of patients with AIDS-associated lymphoproliferative disorders. HHV-6 is a β-herpesvirus related to HHV-7 and HCMV, with greatest cell tropism for T lymphocytes. In 1998, it was discovered that HHV-6 was the causative agent of the nearly ubiquitous childhood disease exanthema subitum, commonly referred to as roseola (8). A combination of recent technological advances and epidemiological studies has shown HHV-6 to be unique among all human herpesviruses, as the virus can integrate into the telomeres of human chromosomes to achieve latency. Viral integration into human telomere sequences is associated with clinically observed vertical transmission of HHV-6 from parents to offspring.
Genomic Structure of HHV-6
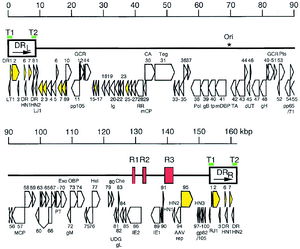
HHV-6 is an enveloped, linear double stranded DNA virus. The genome is 160 to 162 kB in size, with a 143-144 kB unique region (1, 2). The unique region is flanked by 8-9 kB terminal direct repeats at each end, which contain repeats of the hexanucleotide sequence GGGTTA, identical to the telomeric sequences of vertebrate chromosomes (3, 4).
The genome has a cluster of seven gene blocks that are conserved among all herpesviruses, as well as the US22 gene family, conserved in other β-herpesviruses such as CMV and HHV-7 (5, 6). Although the function of the US22 gene family is not fully understood, it has been found that members of this gene family may act as transactivators, enhancing and increasing HHV-6 viral gene expression (7).
The products of HHV-6 genes U27, U41, U43, U74, U77, and U73 are involved in viral replication. The protein product of gene U27 encodes the DNA polymerase processivity factor, binding to DNA polymerase and thus, enhances the rate of viral genome transcription. Gene U41 encodes a single stranded DNA binding protein, also facilitating DNA polymerase's interaction with the viral genome. The protein products of genes U43, U74, and U77 produce a viral helicase-primase complex that facilitates replication of the viral genome. Additionally, the origin-binding protein produced by gene U73 also aids in genomic DNA replication. HHV-6 attachment to host cells is facilitated by viral surface proteins gB and gH, encoded by viral genes U39 and U48, respectively (8).
Unique to HHV-6 among all other herpesviruses are genes U83 and U94. Gene U83 encodes a chemokine that uses signaling mechanisms such as calcium fluxes to recruit lymphocytes and macrophages for productive or latent infection (9). Gene U94 encodes protein product RepH6, which binds the human TATA-binding protein, where it has an effect on regulation of viral gene expression and viral DNA replication. Furthermore, RepH6 has been shown to play a role in latent stage infections, as it is transcribed in latently infected lymphocytes (10).
There are two variants of HHV-6; HHV-6A and HHV-6B. HHV-6A and HHV6B share an overall nucleotide sequence identity of approximately 90%. HHV-6A has 110 ORFs, while HHV-6B has 119 ORFs. Major genomic differences between the two strains lie in the direct repeat regions near the ends of the viral genome, as well as in the proteins encoded by genes U86-U100 in the early stage of infection. The differences observed in early stage viral proteins are the result of the splicing patterns unique to each viral variant (11, 12).
HHV-6 Replication Cycle
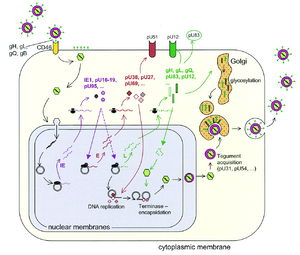
To initiate viral replication, HHV-6A and HHV-6B glycoproteins gH, gL, gQ, and gB attach to host cell surface receptor CD46. All nucleated cells in the human body contain CD46, and thus, the range of host cells for HHV-6 is expansive. The nucleocapsid of HHV-6 is then transported to the host cell's nucleus via the cellular microtubule network. Viral DNA is then released into the nucleus (12).
Expression of the viral genome requires utilization of host transcriptional and translational machinery. Like other herpesviruses, both variants of HHV-6 express initial early (IE), early (E), and late (L) stage proteins. IE proteins, encoded by viral genes U86, U89, U95, U16-U-19, and U3, are expressed immediately after infection and are involved in regulation of subsequent viral gene expression. Early proteins, encoded by viral genes such as U27, U41, U43/U74/U77, U73, and U94 are required for replication of the viral genome and virion assembly. Late stage proteins often become incorporated into mature virions. An example of a late HHV-6 protein is that encoded by viral gene U83, which helps to establish latent infection (8, 9, 12).
Viral DNA replication takes places in the host cell nucleus, by means of the rolling circle mechanism characteristic of herpesviruses. Virions are packaged and assembled within the host nucleus. Virus particles bud out of the nucleus into the cytoplasm, where acquire tegument proteins encoded by viral genes U31 and U54. After tegument acquisition, the virions travel to the Golgi, where they are enveloped with glycoproteins gH, gL, gQ, and gB. Mature HHV-6 particles are then released from the cell via exocytosis (Figure 4).
Epidemiology
Seroprevalence of HHV-6 is estimated to range between 90-95% in the adult population of the world. It has been shown that antibody titers and thus seropositivity can decrease with increasing age (13). Seroprevalence in the world is rapidly approaching 100%, with the exception of Morocco, where only 20% of individuals are seropositive (14).
Initial HHV-6 infection usually occurs within the first 6 to 15 months of life, with peak infection rates observed in infants 6 to 9 months of age, after the protection of maternal antibodies cease (15). Although there is no serologic test to distinguish HHV-6A and HHV-6B, it is believed that HHV-6B is more seroprevalent, causing the majority of clinically observed infections. HHV-6A seropositivity is usually observed in immunocompromised individuals or in adult patients who display clinical signs of exanthema subitum (roseola) (8, 12, 16).
HHV-6 transmission likely occurs through saliva, as HHV-6 can replicate readily in the epithelial cells of the salivary glands. This mode of transmission is especially likely for HHV-6B, as almost all saliva samples analyzed for HHV-6 contain the HHV-6B variant (5, 17). Germ line transmission of HHV-6 can occur, and is observed in approximately 2% of births (18).
Pathophysiology of HHV-6 Infection
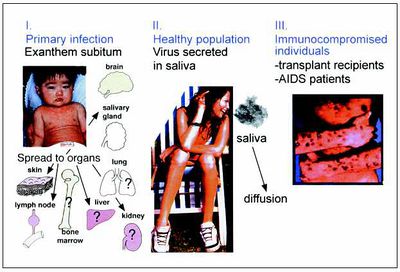
There are 3 recognized stages of HHV-6 infection. Initial HHV-6 infection most often occurs within children 6 to 15 months old and is generally the only symptomatic period most infected individuals will experience. Latent infection persists in the lymphocytes and monocytes of healthy adults and children. The virus shows no sign of pathology during this stage and individuals remain asymptomatic. Although infrequent, reactivation of the latently infected virus or re-infection in immunocompromised individuals can occur, causing a recurrence of symptoms [8, 12].
Initial Infection
HHV-6 infection observed in infants and young children is almost always caused by variant HHV-6B. HHV-6B can infect a wide range of host cells because of the prevalence of host cell receptor CD46. However, HHV-6B most often replicates in CD46 coated CD4+ T lymphocytes during initial infection(8).
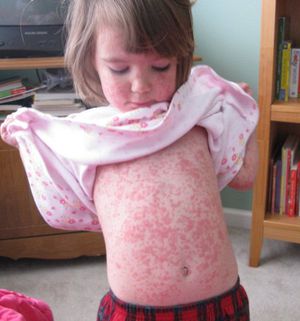
Initial infection is characterized by a febrile illness, with fever temperatures nearing or exceeding 40°C (104°F). HHV-6 infection accounts for 10-40% of hospital admissions of infants with high fevers (19). The fever can last for 3-7 days and then rapidly terminates. After cessation of the fever, approximately 17% of infected infants develop exanthema subitum, commonly known as roseola (20). The rash persists for 1-2 days, and is not a means of transmission. The rash is most often confined to the back, trunk, and neck, but can appear on the face, arms, and legs, generally causing no discomfort to the child (21).
In addition to high fever and in fewer cases, the development of exanthema subitum, HHV-6 infection can cause fatigue, gastrointestinal tract and respiratory tract symptoms. Approximately 10% of children in the initial stage of HHV-6 infection experience febrile seizures (12). High levels of viral titer have been isolated from the brain and cerebrospinal fluid, and can be associated with rare CNS symptoms, such as encephalitis (8, 12).
Latency in Healthy Children and Adults
HHV-6 latency and persistence in healthy, seropositive adults and children characterizes the second stage of HHV-6 infection. Viral replication continues in the epithelial cells of the salivary glands during this stage. This accounts for the abundance of HHV-6 isolates in saliva samples, and also represents the primary means of transmission to seronegative individuals (22).
Latency is achieved in host lymphocytes, monocytes, and possibly some organ tissues. HHV-6 DNA can be detected in the peripheral blood mononuclear cells (PBMCs) of 90% of seropositive individuals. HHV-6 achieves latency through integration into specific sites of the host chromosome (23).
Reactivation in Immunocompromised Individuals
The most acute clinical manifestations of HHV-6 infection are observed during reactivation of the virus in immunocompromised individuals. Such immunosuppression is often observed in individuals receiving organ transplants, as immunocompetency is reduced therapeutically to decrease chances of transplant rejection (12). Reactivation is also observed clinically in AIDS patients. HHV-6 has been shown to increase HIV replication by up-regulating certain cytokines and transactivating the long terminal repeats of the HIV genome, leading to increased viral loads and more aggressive AIDS onset (24).
Latency via Human Chromosomal Integration
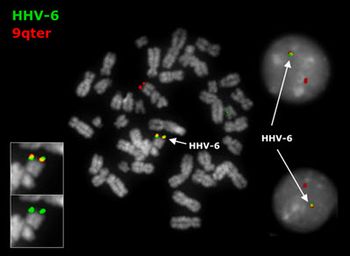
HHV-6 is unique among human herpesviruses in that it is able to persist in latent state by naturally integrating into regions of the host's chromosomes. HHV-6 chromosomal integration is observed in approximately 2% of seropositive individuals, and is often the cause of high levels of viral DNA in host serum and plasma. HHV-6A and HHV-6B have both been found to integrate into the human genome, yet independently; co-integration has never been observed. Integration of HHV-6A is observed in nearly a third of congenital infections, suggesting that germ line transmission of the virus results in a significant number of cases (25, 26). In cases of HHV-6 chromosomal integration, viral DNA can be isolated from every cell in the body, including skin fibroblasts, hair follicles, and blood cells (25). Immunocompetent patients with viral chromosome integration are identified by comparing the HHV-6 viral load in the host's blood and serum. Although viral loads will be substantial in both samples, concentration in serum will be at least 50-fold lower than the blood isolates (27). The entire HHV-6A genome does not integrate into the human genome. The mechanism of integration via homologous recombination results in the loss of 79 nucleotides from the HHV-6A genome (29. It has also been suggested that this mechanism of homologous recombination could result in the loss of host DNA, especially near the subtelomeric region, where the TERRA gene encoding telomerase is located (29). Loss of telomerase could result in chromosome shortening over time, resulting in the loss of host genetic material (29).
Once integrated into the subtelomeric region of the host chromosome, the HHV-6 virus persists in its latent state. However, cretain stressors and chemical compounds have been shown to induce viral replication. For example, Arbuckle et al. 2010 induced HHV-6A viral replication after treating HEK-293 cells with integrated HHV-6, as well as the PBMC's of patients, with known herpesvirus replication inducing compounds TPA and TSA. Although both compounds induced viral replication, TSA induced expression of latent viral gene U94 to a greater extent. Arbuckle et al. was also able to infect naive Molt-3 cells with reactivated HHV-6A integrated PBMCs. Syncytia formed between the infected T-cells and the Molt-3 cells, and soon after, Gardella gel analysis indicated the presence of replicating linear viral DNA and RNA. Thus, HHV-6A has a successful latent state, as after integration, it is able to replicate and infect new host cells (29).
HHV-6 has already been shown to integrate into 7 different chromosomal sites. In every case analyzed, chromosomal integration is always found in the telomere regions of the host chromosome, on either the p or q arm of one homolog of the chromosome, although not confined to a particular chromosome (25). Human telomeres contain repeats of the sequence TTAGGG. These repeats function to protect the ends of the chromosomes from degradation due to exonucleases. Additionally, these repeat sequences prevent shortening of the chromosome during replication. The genome of HHV-6 contains complementary GGGTTA repeats in the direct repeat regions. The repeats in the HHV-6 genome are only 21 to 80 base pairs in length, and are thus significantly shorter than the repeat sequences observed in human telomeres (28). The mechanism of HHV-6 integration is likely due to homologous recombination between the telomere repeat sequences of the human chromosome and the complementary repeat sequences of the HHV-6 genome. It has been previously shown that the telomere regions of the human chromosome are unstable and prone to such homologous recombination events (25). Another proposed mechanism of HHV-6 chromosomal integration involves the RepH6 viral protein encoded by gene U94. RepH6 can bind DNA site-specifically, and has also been shown to function as an endonuclease, helicase, and ATPase. Such functions of RepH6 mirror the chromosomal integration techniques of the related adeno-associated virus (AAV) rep gene. Therefore, RepH6 may facilitate chromosomal integration in a manner similar to the rep gene of AAV (25)
Several specific sites of HHV-6 chromosomal integration have been identified using FISH, chromosome specific PCR sequencing, and Gardella gel techniques (25, 26, 27, 29). A study of 9 British individuals, 7 adults, and 2 children, showed ubiquitous HHV-6 chromosomal integration. Only one site of HHV-6 integration was observed in each individual. This particular study identified chromosome sites 9q34.3, 10q26.3, 11p15.5, 17p13.3, and 19q 13.4 has sites of HHV-6 integration (25). A more recent study identified telomeric sites 17p13.3, 18q23, and 22q13.3 as sites of HHV-6 integration. In this study, sites of HHV-6A integration were conserved among family members, suggesting possibility of vertical transmission (29). Both studies were able to induce reactivation and latent viral replication via treatment with established viral reactivating compounds.
HHV-6 chromosomal integration has been observed in malignant lymphomas. It is suggested that integrated HHV-6 DNA could be oncogenic, modifying the expression of host cell proteins involved in cell growth and apoptotic signaling (30).
Although HHV-6 is the only human herpesvirus known to integrate into host chromosomes, Marek's disease virus, an avian herpesvirus, also integrates into host chromosomes (31). Marek's virus also integrates into host chromosomes via similar complement telomeric sequences. Phylogenetic analysis of HHV-6 and Marek's disease genomes show no close genetic relationship. However, the similar complement telomeric repeats GGGTTA are shared by both viruses, reinforcing these sites as the mechanism of chromosomal integration (25, 31).
Vertical Transmission of Integrated HHV-6 through Host Germ Line
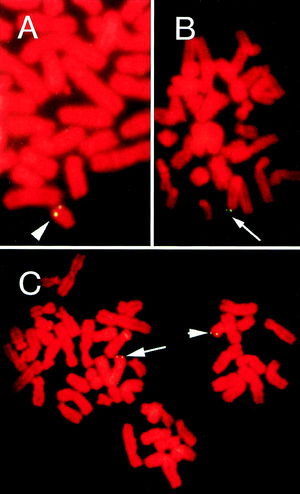
Although the primary route of HHV-6 transmission is thought to be from mother to child through saliva, several case studies have demonstrated that vertical transmission to offspring through the germ line is also possible. In a study of four families with high HHV-6 viral loads, FISH and chromosome specific PCR sequencing at viral-chromosomal junctions has shown that HHV-6A can be transmitted through the germ line (29). HHV-6B was also shown to be vertically transmitted in case studies of four families. Several individuals in these families suffered from similar autoimmune disorders and even lymphomas, suggesting clinical inheritance (32). FISH and hair follicle analyses of congenitally infected infants and their parents have shown that chromosomal integration of HHV-6 causes approximately 86% of such congenital infections. Statistically, this suggests that 1 out of every 116 live-born infants exhibits chromosomally integrated HHV-6 (26). It has also been shown that the offspring of individuals with chromosomally integrated HHV-6 can inherit the virus at two different loci, one from each parent, once more reinforcing the mechanism of vertical transmission (Figure 8, 33).
The Initial experiments detailing chromosomal integration of HHV-6 utilized FISH analyses. In one particular example, FISH analyses were able to show that the offspring of two individuals with HHV-6A integration at different chromosomal sites inherited HHV-6 at both sites (33). Although this experimental approach gave credence to the mechanism of germ line transmission of the integrated virus, FISH analysis cannot distinguish between actual integration and non-covalent linkages observed in the episomal latency of other herpesviruses (29). Therefore, Arbuckle et al., 2010 aimed to confirm the vertical transmission hypothesis by analyzing the peripheral blood mononuclear cells of families known to have integrated HHV-6A with techniques that could prove actual chromosomal integration. The study focused on four families with high viral loads, confirmed HHV-6A integration, with some members displaying central nervous system dysfunction (29). FISH analyses initially confirmed identical integration sites in parents and offspring. However, these results were validated with RT-PCR and subsequent sequence analysis of the viral-chromosomal junction. Parents and their offspring had the same HHV-6A integration site. All of these integration sites were located at the sub-telomeric region of the chromosome, however, the chromosome where this integration occurred varied from family to family, suggesting that HHV-6 does not have a preferential chromosome for its latent state. Rather, the virus needs only a sub-telomeric region with complementary telomeric repeats to integrate and begin its latent phase (29).
The implications of the germ line transmission of integrated HHV-6 are numerous. It is still unknown whether infants born with integrated HHV-6 have antibodies to the virus, or are still susceptible to infection with HHV-6 variants. Furthermore, HHV-6 integration into the human genome poses questions about the evolutionary significance of this unique latent state. Germ line transmission of the virus is evidence of conservation of the integrated virus in the human genome. We must look to future studies to illustrate the implications of this change to the human genome with regard to normal host gene expression, the possibility of the development of oncogenes, and the long term effects of latent state viral reactivation and replication on the host.
Future Work
Because HHV-6 is the only human herpesvirus known to integrate into the human genome, many questions regarding this mechanism of viral latency, its effect on host cell gene regulation, and possible pathological effects on the host need to be addressed. The molecular biology of the chromosomally integrated HHV-6 latent state is not fully understood. What underlying molecular mechanisms permit a virus to establish latency through integration rather than engaging in lytic cycle infection? Furthermore, the pathological consequences of chromosomal integration are of great interest. Because HHV-6 can insert itself into the genome, it may disrupt the expression of endogenous human genes, perhaps facilitating the development of cancer and other disease states. HHV-6's specific integration in the telomere repeats of the human genome may disrupt telomerase activity. Furthermore, disruption of normal telomere sequences via HHV-6 insertion can affect the expression of subtelomeric genes, which has been associated with mental retardation (29). Now that specific HHV-6 integration sites have been established, the pathological and clinical consequences of such recombinant events need to be further explored.
Another area of research interest is the relationship between HHV-6 and multiple sclerosis (MS) One study found that HHV-6 may play an important role in multiple sclerosis through a mechanism of molecular mimicry. It was found that there was high sequence identity between myelin basic protein (MBP), and residues 4-10 of the HHV-6 U24 gene. T-cells recognizing MBP were able to be activated by these viral residues. It is suggested that the presence and reactivation of HHV-6 in a patient can contribute to MS pathogenesis, as CD4+ cells are unable to discriminate between the viral residues and host myelin, leading to the destruction of insulating myelin protein, and the development of the autoimmune disorder (34). Further research and case studies are necessary to establish the link between HHV-6 and MS.
References
1 Dominguez, G., Dambaugh, T. R., Stamey, F. R., Dewhurst, S., Inoue, N. & Pellett, P. E. 1999. Human herpesvirus 6B genome sequence:
coding content and comparison with human herpesvirus 6A. J Virol 73:8040–8052.
2 Gompels, U. A., Nicholas, J., Lawrence, G., Jones, M., Thomson, B. J., Martin, M. E., Efstathiou, S., Craxton, M. & Macaulay, H. A. 1995. The DNA sequence of human herpesvirus-6: structure, coding content, and genome evolution. Virology 209:29–51.
3 Braun, D. K., Pellett, P. E., Hanson C. A. 1995. Presence and expression of human herpesvirus 6 in peripheral blood mononuclear cells of S100-positive, T cell chronic lymphoproliferative disease. J. Infect. Dis. 171:1351–1355.
4 Meyne, J., Ratliff, R. L., Moyzis, R. K. 1989. Conservation of the human telomere sequence (TTAGGG)n among vertebrates. Proc. Natl. Acad. Sci.USA 86:7049–7053.
5 Clark, D. A. 2000. Human herpesvirus 6. Rev Med Virol 10:155–173
6 Mendez J.C., Dockrell, D.H., Espy, M.J., Smith, T.F., Wilson, J.A., Harmsen, W.S., Ilstrup, D., Paya, C.V. 2001. Human beta-herpesvirus interactions in solid organ transplant recipients. J Infect Dis 183(2):179-184.
7 Nicholas, J., Martin M.E. 1994. Nucleotide sequence analysis of a 38.5-kilobase-pair region of the genome of human herpesvirus 6 encoding human cytomegalovirus immediate-early gene homologs and transactivating functions. J. Virol. 68:597–610
8 Dockrell, D. H. 2003. Human herpesvirus 6: molecular biology and clinical features. J. Med. Microbiol. 52:5–18.
9 Zou, P., Isegawa, Y., Nakano, K., Haque, M., Horiguchi, Y., Yamanishi, K. 1999. Human herpesvirus 6 open reading frame U83 encodes a functional chemokine. J Virol 73:5926–5933.
10 Rotola, A., Ravaioli, T., Gonelli, A., Dewhurst, S., Cassai, E. & DiLuca, D. 1998. U94 of human herpesvirus 6 is expressed in latently infected peripheral blood mononuclear cells and blocks viral gene expression in transformed lymphocytes in culture. Proc Natl Acad Sci USA 95:13911–13916.
11 Chou, S., Marousek G. I. 1994. Analysis of interstrain variation in a putative immediate-early region of human herpesvirus 6 DNA and definition of variant-specific sequences. Virology 198:370–376.
12 Bolle, L.D., Naesens L., Erik, D.C. 2005. Update on human herpesvirus 6 biology, clinical features, and therapy, Clin Microbiol Rev 18:217–245.
13 Brown, N. A., Sumaya, C. V., Liu, C.-R., Ench, Y., Kovacs, A., Coronesi, M., Kaplan, M. H. 1988. Fall in human herpesvirus 6 seropositivity with age. Lancet ii, 396.
14 Ranger S., Patillaud S., Denis F., Himmich A., Sangare A., M'Boup S. 1991.Seroepidemiology of human herpesvirus-6 in pregnant women from different parts of the world. J Med Virol 34:194-8.
15 Enders, G., Biber, M., Meyer, G., Helftenbein, E. 1990. Prevalence of antibodies to human herpesvirus 6 in different age groups, in children with exanthema subitum, other acute exanthematous childhood diseases, Kawasaki syndrome, and acute infections with other herpesviruses and HIV. Infection 18:12–15.
16 Wang, F. Z., Dahl, H., Ljungman, P., Linde, A. 1999. Lymphoproliferative responses to human herpesvirus-6 variant A and variant B in healthy adults. J. Med. Virol. 57:134–139.
17 Collot, S., Petit, B., Bordessoule, D., Alain, S., Touati, M., Denis, F., Ranger-Rogez, S. 2002. Real-time PCR for quantification of human herpesvirus 6 DNA from lymph nodes and saliva. J. Clin. Microbiol. 40:2445–2451.
18 van Loon, N. M., Gummuluru, S., Sherwood, D. J., Marentes, R., Hall, C. B. & Dewhurst, S. 1995. Direct sequence analysis of human herpesvirus 6 (HHV-6) sequences from infants and comparison of HHV-6 sequences from mother/infant pairs. Clin Infect Dis 21:1017–1019.
19 Portolani M., Cermelli C., Moroni A. 1993. Human herpesvirus-6 infections in infants admitted to hospital. J. Med.Virol. 39:146–51
20 Pruksananonda, P., Hall, C.B., Insel, R.A., McIntyre, K., Pellett, P.E., Long, C. E., Schnabel, K. C., Pincus, P. H., Stamey, F. R., Dambaugh, T. R. 1992. Primary human herpesvirus 6 infection in young children. N. Engl. J. Med. 326:1445–1450.
21 Stoeckle M.Y. The spectrum of human herpesvirus 6 infection: from roseola infantum to adult disease. Annu Rev Med 2000: 51: 423–430.
22 Di Luca, D., Mirandola, P., Ravaioli, T., Bigoni, B., Cassai, E. Distribution of HHV-6 variants in human tissues. Infectious Agents and Disease 1996;5:203-14.
23 Luppi, M., Barozzi, P., Morris, C.M., Merelli, E., Torelli, G. 1998. Integration of human herpesvirus 6 genome in human chromosomes. Lancet 352:1707-8.
24 Lusso, P., Gallo, R.C. 1995. Human herpesvirus 6 in AIDS. Immunol Today 16:67-71.
25 Nacheva, E.P., Ward, K.N., Brazma, D., Virgili, A., Howard, J., Leong, H.N., Clark, D.A. 2008. Human herpesvirus 6 integrates within telomeric regions as evidenced by five different chromosomal sites. Journal of Medical Virology, 80:1952–1958.
26 Hall, C.B., Caserta, M.T., Schnabel, K., Shelley, L.M., Marino, A.S., Carnahan, J.A. 2008. Chromosomal integration of human herpesvirus 6 is the major mode of congenital human herpesvirus 6 infection. Pediatrics 122(3): 513–520.
27 Ward, K.N., Leong, H.N., Nacheva, E.P., Howard, J., Atkinson, C.E., Davies, N.W., Griffiths, P.D., Clark, D.A. 2006. Human herpesvirus 6 chromosomal integration in immunocompetent patients results in high levels of viral DNA in blood, sera, and hair follicles. J Clin Microbiol 44:1571–1574.
28 Gompels, U. A., Macaulay, H.A. 1995. Characterization of human telomeric repeat sequences from human herpesvirus 6 and relationship to replication. J. Gen. Virol. 76:451-458.
29 Arbuckle, J.H., Medveczky, M.M., Luka, J., Hadley, S.H., Luegmayr, A., Ablashi, D., Lund, T.C., Tolar, J., DeMeirleir, K., Montoya, J.G., Komaroff, A.L., Ambros, P.F., Medveczky, P.G. 2010. The latent human herpesvirus-6A genome specifically integrates in telomeres of human chromosomes in vivo and in vitro. Proc Natl Acad Sci USA. 107:5563–5568.
30 Daibata, M., Taguchi, T., Taguchi, H., Miyoshi, I. 1998.Integration of human herpesvirus 6 in a Burkitt's lymphoma cell line. Br J Haematol 102:1307-1313
31 Kishi, M., Harada, H., Takahashi, M., Tanada, A., Hayashi, M., Nonoyama, M., Josephs, S.F., Buchbinder, A., Schachter, F., Ablashi, D., Wong-Staal, F., Salahuddin, S.Z., Gallo, R.C. 1988. A repeat sequence, GGGTTA, is shared by DNA of human herpesvirus 6 and Marek’s disease virus. J. Virol. 62:4824–4827.
32 Tanaka-Taya, K., Sashihara, J., Kurahashi, H., Amo, K., Miyagawa, H., Kondo, K., Okada, S., Yamanishi, K. 2004. Human herpesvirus 6 (HHV-6) is transmitted from parent to child in an integrated form and characterization of cases with chromosomally integrated HHV-6 DNA. J Med Virol 73:465-473.
33 Daibata, M., Taguchi, T., Nemoto, Y., Taguchi, H., Miyoshi, I. Inheritance of chromosomally integrated human herpesvirus 6 DNA. Blood 1999; 94:1545-1549.
34 Ablashi, D.V., Eastman, H.B., Owen, C.B., Roman, M.M., Friedman, J., Zabriskie, J.B., Peterson, D.L., Pearson, G.R., Whitman, J.E. 2003. Cross-reactivity with myelin basic protein and human herpesvirus-6 in multiple sclerosis. Annu Neurol. 53(2): 189–197.
Edited by Kerri-Lynn Conrad, student of Joan Slonczewski for BIOL 375 Virology, 2010, Kenyon College.
