Persister Cells in E. coli
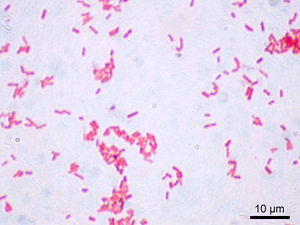
Bacterial communities form dormant, antibiotic tolerant cells called persister cells.5 These cells are not mutants, but phenotypic variants of the wild type.5 Persisters are selected to increase chances of survival in fluctuating environments.2. The diminished (or in some cases, overproduction) activity of key molecules in the cell, as well as non-growing or reduced growth rate, make cells in dormant state distinguishable.2 Persisters are found in all phases of cell growth and are non-dividing with a reduced metabolism.2,10 The slow growth of the persister cells protects them from antibiotics.1 Cells in the persister state are highly immune to bacterial antibiotics, which interfere with processes of key molecules.1 Persisters may hold a lot of explanation for biofilm tolerance to antibiotics.4,6 There has been extensive research performed on the persister cells in Escherichia coli (Figure 1) yet we still do not know why they do not make persisters in early-exponential growth, which indicates presence of specific persister genes.5 Persistence has been extremely difficult to research because persister subpopulations are constantly in flux and this subpopulation is a very small portion of the total bacteria population, for example, only one in one hundred thousand to one million cells of wild type Escherichia coli cells are persisters. 13
Types of Persistence
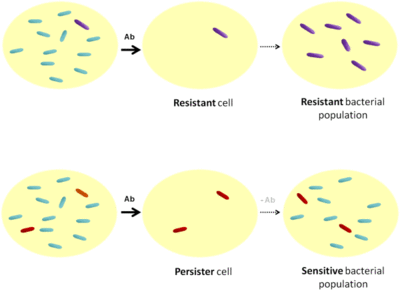
Persistence is triggered by environmental and intracellular stressors. While the process of differentiation is not completely understood, there is some evidence suggesting how and why cells become persistent. It should be noted that persistence is not the same as resistance (Figure 2). There are two different types of persistence.
Type I
Type I persisters are triggered to differentiate and become physiologically antibiotic tolerant, using strategies to resist persistence unless exposed to stressful conditions, such as starvation. This suggests that the bacteria sense these stressful environments and in preparation, push part of the population to become dormant. The rest of the population is left to thrive and risk death.1 Type I persisters are spawned during stationary phase and do not increase in numbers during growth phase. 2,13
Type II
Type II persisters are repetitively produced without environmental triggers and stressors and are slow growing.1 These “bet-hedging” strategies of persistence are believed to have evolved from antibiotic stressors and stress imposed on bacteria by prophages.7
Genetic Mechanisms
There are at least 37 known types of Toxin-Antitoxin (TA) systems in Escherichia coli, which are thought to be the genetic basis for the formation of persisters from normal cells.10 These toxin mechanisms usually consist of two genes encoding for stable toxins that disturb essential processes, and an unstable antitoxin that remedies the effects of the toxin.10 Toxin genes always code for proteins and the antitoxins code for either protein or RNA.3
Under type I conditions, the antitoxins are degraded by proteases in the cell, ClpXp or Lon, and the toxins are released to do their typical function in the cell.3 The most well depicted system is the Type II TA system, made up of a two-gene operon which encodes a stable toxin and an unstable antitoxin, MqsR/MqsRA. This TA system produces proteins that come together to form a complex that is not toxic. The antitoxin mqsA can be found just below the mqsR in the same operon, and encodes a slightly longer amino acid protein that neutralizes mqsR toxicity. 3 Evidence suggests that the mqsRA TA system plays an important part in many of Escherichia coli cellular processes. Deletion of mqsR and the mqsRA operon decreased persister cell formation, while mqsR overexpression increased persister formation. This is important because mqsA has also been shown to serve as a transcriptional regulator of several Escherichia coli genes, including general response regulators, as well as the mqsRA operon.3
The first candidate persister gene identified was the hipA gene encoded by the hipBA locus. HipA is a high peristance gene that encodes for toxin-antitoxin modules and overexpression of this gene leads to persistent phenotype and growth arrest.9 It inhibits protein synthesis and leads to tolerance of many drugs, possibly even persister formation.5 HipA is widely thought to be the seed of persistent infections. 8
Anti-Persister Therapies
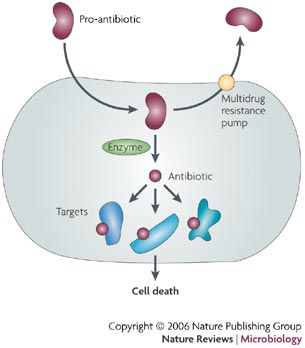
It seems unlikely to be able to come up with the perfect anti-resister antibiotic. Kim Lewis has suggested an antibiotic that is a benign compound which has a bacterial enzyme that is converted to a antiseptic compound once it enters the cytoplasm (Fig 3). The goal is for this compound to bind covalently to many bacterial components and in effect, killing the cell. 11
There are some antimicrobials which are similar to the ideal anti-persister and utilize broad-spectrum compounds that kill anaerobic bacteria by converting to antiseptic-type molecules covalently binding to cells inside the cytoplasm. 11
Because there are so many genes involved in persistence, it is next to impossible to develop an antibiotic that targets the persister cells genetic mechanisms. Some researchers have found that it may be better to target persister cells using a bactericidal approach, utilizing the stressful environment which drives cells to become dormant (and in general, avoid the affects of antibiotics). If these processes could be tracked, they could be targeted as potential weak spots for the persister cells. 12
Further Reading
[1] Antibiotic/Antimicrobial Resistance Information from the Center of Disease Control and Prevention
[2] Cancer Persister Cells
References
Edited by Michelle Suarez, a student of Nora Sullivan in BIOL168L (Microbiology) in The Keck Science Department of the Claremont Colleges Spring 2014.