Quasispecies Theory and Novel HIV Therapeutics Based on Lethal Mutagenesis
A Viral Biorealm page on the family Quasispecies Theory and Novel HIV Therapeutics Based on Lethal Mutagenesis
The human immunodeficiency virus (HIV) is the causative agent of acquired immunodeficiency syndrome (AIDS), the second most common infectious cause of death globally. As of the writing of this wiki, HIV is estimated to have infected more than 75 million people globally and has been the proximate cause of death of 36 million people [1,2].
Since the first reported cases of AIDS and the discovery of HIV as its cause in the early 1980's, researchers and clinicians have struggled to develop and administer effective therapeutics to combat HIV/AIDS, and the production of a viable vaccine remains an unrealized goal [3]. The difficulty of developing effective HIV therapeutics and vaccines is due largely to the extraordinary mutation rate of HIV, which enables the virus to rapidly evade the selective pressures imposed by antiretroviral medications and potential vaccines by generating a large and genetically diverse population through mutagenesis [4]. A comprehensive knowledge of the genetic diversity characteristic of HIV populations in infected individuals - what have been termed viral quasispecies - is therefore essential for the discovery and delivery of effective HIV medications and vaccines.
Novel therapeutics based on fundamental components of quasispecies theory, most notably the concepts of error catastrophe and lethal mutagenesis, have been developed and constitute a unique approach toward an effective cure for AIDS and the global eradication of HIV.
Quasispecies Theory and HIV
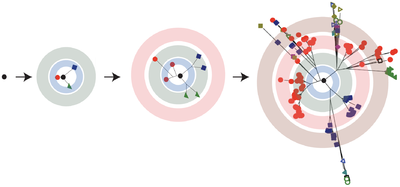
The near-infinite population sizes, high rates of mutagenesis, and rapidity of replication of RNA viruses preclude their description by classical population genetics, in which the fitness of a specific phenotype is principally the product of its ability to reproduce successfully [5]. In RNA viruses such as HIV, extremely high rates of error-prone mutation during the course of viral replication create large swarms of genetically diverse variants. Collectively, this swarm of similar-yet-divergent variants, which stems from an original infection event by a founder RNA virus, forms a quasispecies (Figure 1); [6]. Within a quasispecies, phenotypic fitness depends on both an individual variant's reproductive ability and its capacity for mutation. Understanding the link between quasispecies theory and its effect on the intrahost behavior of HIV may help elucidate therapeutic strategies targeting the attenuation or elimination of retroviral pathogenesis in infected individuals [7].
To be considered a quasispecies, a virus must meet the following criteria:
1. Individual variants within the population must have diverged through mutation from the master sequence of a founder virus.
2. Individual variants within the population must respond interactively to selective pressures within the host.
3. Each variant must contribute to the larger identity and attributes of the total population [7].
Genetic Diversity and HIV Quasispecies
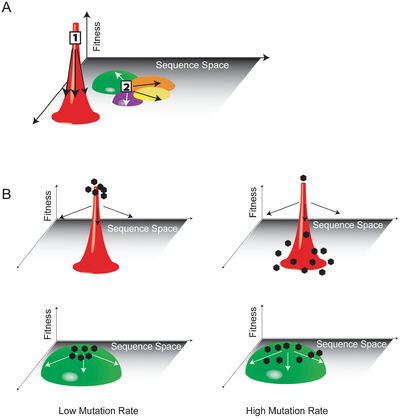
The immense genetic diversity present within its quasispecies is one of the main characteristics of HIV infection within a host. The great majority of HIV infections stem from the transmission of a single virion - the master sequence around which the quasispecies is organized [8]. The generation of HIV quasispecies in infected hosts is driven principally by the low replicative fidelity of HIV reverse transcriptase, the rapid kinetics of HIV replication, aspects of the HIV replicative cycle itself that are conducive to mutagenesis, and viral genetic recombination. Reverse transcription of the HIV RNA genome results in 1-2 mutations per virion, or about 10-4 mutations at each transcribed nucleotide [7,9]. Current evidence suggests that, within a given RNA viral quasispecies, at least every possible point mutation and a large number of double mutations are produced during each cycle of replication [10]. Furthermore, the genetic diversity of the viral population present within a single infected host after full progression to AIDS has been estimated to rival the annual genetic diversity of influenza [4]. The vast genetic diversity of the HIV quasispecies residing within infected hosts has important implications for viral fitness. Cellular and tissue tropism, immune escape, and virulence are significant determinants of viral fitness in addition to replicative capacity [11]. The ability to produce an expansive array of genetic variants - to probe a large area of sequence space (a geometric model of sequence variation in which genetic divergence is represented by physical distance) - allows HIV to rapidly adapt to selective pressures, infect a wide breadth of cells, and reproduce successfully (Figure 2).
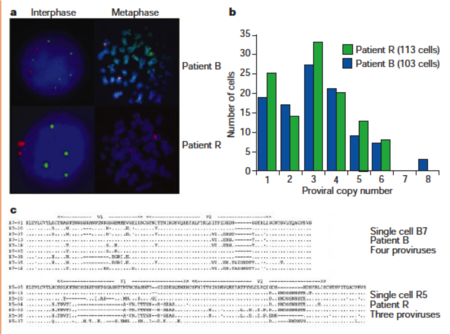
Genetic recombination during the HIV replication cycle constitutes an important contribution to the genetic diversity present within an HIV quasispecies.
During the HIV replication cycle, transcription of the HIV genome by host cellular RNA polymerase II provides a minor source of transcriptional errors [35]. A more significant source of viral mutation occurs directly downstream of genomic transcription, during the packaging of the transcribed viral genome into individual virions at the host cell plasma membrane [36]. Retroviruses such as HIV house two copies of the viral genome within a single virion, which are linked non-covalently at the dimerization initiation site [37]. During the reverse transcription of the HIV genome within a subsequently infected host cell, reverse transcriptase engages in template switching between both strands, potentially producing a chimeric, recombinant DNA sequence. In order for this recombination event to produce novel genetic diversity within the HIV quasispecies, a host cell must be co-infected with at least two virions containing genetically divergent RNA dimers. Furthermore, once produced via recombinant reverse transcription, chimeric replicants must be packaged together within a single virion. Despite the seeming unlikelihood of these events occurring with great enough frequency to produce a large number of chimeric, recombinant HIV genomes, experimental evidence suggests that the in vivo rate of viral co-infection can be extremely high. A 2002 study by Andreas Jung and colleagues utilizing fluorescent in situ hybridization of HIV-infected spleen cells found that most infected cells contain at least two genetically distinct proviruses (Figure 3); [38]. Experimental evidence has also been gathered that the co-packaging of HIV genomes occurs regardless of the virus' ability to fully integrate into the host cell genome [39]. Taken together, these observations suggest that the recombination of HIV genomes during the retroviral replication cycle represents an important contribution to the breadth of genetic diversity present within HIV quasispecies.
Master Sequences and Progeny Virions
An important question which arises from the observation that the intrahost genetic variation of HIV quasispecies stems from the transmission of a single founder virus, containing a master sequence, is whether or not progeny virions augment, either directly or indirectly, the fitness of the founder virus. A 2014 study conducted by Jonathan Carlson and colleagues investigating the heterosexual HIV transmission bottleneck - that is, the severe selection bias that occurs during HIV infection which results in the transmission of only a single HIV variant out of the entire quasispecies - found that a virion's likelihood of transmission is independent of several factors which increase intrahost viral fitness. In their study, Carlson et al. utilized a statistical model which was demonstrated to accurately quantify the probability of transmission for a given virion based on its relationship to a consensus sequence derived from a large cohort of infected couples. A striking finding of this modeling study was that amino acid sequences that are correlated with a virion's increased immune escape from human leukocyte antigen alleles are not likely to be found within the genome of founder viruses, implying that the benefit of this fitness advantage is offset by its replicative cost [40]. Such immune escape adaptations are not likely to be required by a founder virus during the infection of an immunologically naive host, in which neutralizing antibodies and other infection-mediating factors of the adaptive immune response have not yet been produced against HIV infection. Another observation in favor of the hypothesis that subsequent progeny variants provide functions that support the propagation, and ultimate transmission, of virions bearing a master sequence concerns the glycosylation of the HIV Env protein. Env alleles which produce lesser degrees of glycosylation and whose genomes are more similar to ancestral sequences are favored for transmission as components of founder virus genomes. High levels of Env glycosylation are associated with increased immune escape, as glycosylation provides a steric boundary (the 'glycocalyx') against neutralizing host factors [40]. In toto, these observations suggest that the propagation and transmission of founder viruses is independent of factors which play a large role in mediating viral fitness within non-immunologically naive hosts.
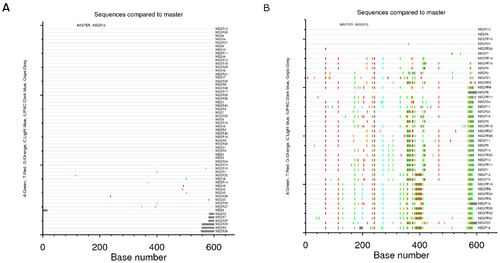
Another investigation into the genetic links between founder and progeny virions was conducted in 2013 by Pierre Frange et al. This study collated sequencing data gathered via single genome amplification, from eight recently infected pairs of HIV-infected patients. Their results provide further evidence in support of the existence of the sexual transmission HIV bottleneck; while Frange et al. found striking and significant variation within the genetic diversity of donors, the sequencing data from newly-infected patients' revealed the presence of only a single HIV variant (Figure 4); [44].
Mutational Coupling
In an HIV quasispecies, the phenomenon of mutational coupling allows for the persistence of genetic diversity at the expense of the replicative ability of some individual variants. Mutational coupling occurs when a variant with a relatively low replicative ability persists because of its proximity in sequence space to a variant with a relatively higher replicative ability (Figure 2); [12]. Variants deemed unfit according to traditional population genetics may nevertheless exhibit high fitness if they are likely to undergo mutations that will produce de novo variants with greater replicative ability (their mutational couple). The consequences of mutational coupling on the course of HIV infection have been depicted graphically. In Figure 2, the "fitness landscape" of a quasispecies is represented as containing peaks (high fitness, smaller area of sequence space), valleys (low fitness, larger areas of sequence space), and intermediate areas of relative flatness. Quasispecies with a high degree of mutational coupling are better able to occupy greater areas of sequence space - and achieve a higher overall degree of fitness - than those confined to discrete peaks in the fitness landscape. The robust mutagenesis and correspondingly high degree of mutational coupling in HIV enables its effective occupation of sequence space and facilitates its ability to propagate within infected hosts [7].
Error Threshold and Error Catastrophe
Although HIV quasispecies are well-suited to occupy large swathes of sequence space through the rapid generation of mutant virions, a trade-off exists between the genetic diversity actually expressed within a host and the virus' ability to adapt to selective pressures. The small size (9 kilobases long), complex protein products, and the presence of three open reading frames of the HIV genome all limit the amount of viable variants that may be replicated in successive generations. There is a high likelihood that significantly deleterious mutations will arise from each cycle of HIV replication [9,13]. The 'error threshold' of a viral quasispecies describes the maximum rate of mutation at which the quasispecies can persist within a host without becoming overcome by deleterious mutations and diverging permanently from the master sequence [14]. Experimental evidence on the effect of induced mutagenesis in vesicular stomatitis virus (VSV) and HIV suggests that RNA viruses produce mutations at rates very near their error thresholds [15]. Of particular interest, a study by Crotty et al. examined the effects of the endogenous anti-HIV mutagen class APOBEC3 and exogenous mutagens on the adaptive fitness of HIV in vivo. Crotty and colleagues found that even slight increases in HIV mutational rate, induced by any of the mutagens used, severely attenuated the virus' replicative ability and resulted in the rapid accumulation of non-functional virions; HIV progression beyond the replicative error threshold resulted in error catastrophe, the collapse of viral fitness [16]. These results imply that HIV infection may be effectively treated by stimulating the rate of viral mutation, a strategy known as lethal mutagenesis [17].
Endogenous and Exogenous Mutagens Induce HIV Lethal Mutagenesis
APOBEC3
The human APOBEC3 proteins are cytokines which induce lethal mutagenesis in retroviruses. The best-studied of these cytokines is APOBEC3G, which combats HIV infection in primary CD4+ T lymphocytes [18]. APOBEC3G incorporates into budding virions in HIV infected cells and interferes with viral replication upon the subsequent infection of neighboring T cells [19]. Following the production of single-stranded, negative-sense DNA by reverse transcriptase, APOBEC3G deaminates cytidine bases in the viral DNA, producing a uracil base at each deamination site. Uracil-studded DNA is subsequently targeted for degradation by endogenous apurinic/apyrimidinic endonuclease and uracil N-glycosylase or is transcribed to a hyper-mutated positive-sense DNA strand that produces excessive stop codons within HIV genomic open reading frames (Figure 5). The hyper-mutated virions generated following the successful mutagenic action of APOBEC3G cross the error threshold and are rendered non-viable by deleterious mutations and error catastrophe [20-22].
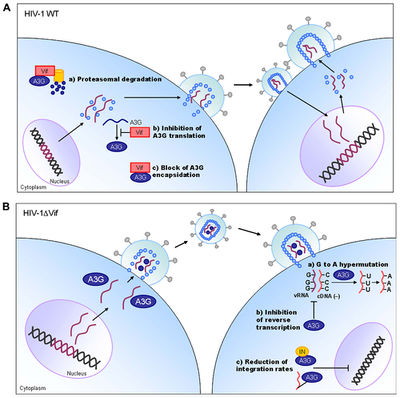
The lethal mutagenic activity of APOBEC3G illustrates a compelling approach for the control of HIV infectivity and viability. Indeed, several studies have reported that APOBEC3 cytokines are capable of decreasing HIV infectivity by 90% within one cycle of replication [23-25]. APOBEC3 lethal mutagens are not without their limitations, however. A 2009 study by Jern and colleagues implicated APOBEC3 cytokines in the induction of sub-lethal mutagenesis during HIV replication, which may ultimately accelerate the adaptation of HIV to selective pressures (i.e. HIV therapeutics) and stimulate drug resistance [26]. Additionally, the endogenous activity of APOBEC3 cytokines is limited by the activity and adaptation of the HIV Vif accessory protein, which inhibits the antiviral action of APOBEC3 by stimulating its degradation in proteasomes [22]. Although the APOBEC3 cytokines exert a strong selective pressure against HIV via lethal mutagenesis, these endogenous antiretrovirals are nevertheless insufficient to combat HIV infection by themselves. Fortunately, exogenous mutagens have been developed which are effective against HIV infection in pre-clinical evaluations.
5-AZC
5-AZC (5-azacytidine) is a ribonucleoside analog that has been demonstrated to limit the infectivity of HIV [26]. In infected cells, 5-AZC stimulates HIV mutagenesis during the reverse transcription of negative-sense viral DNA. First, 5-AZC-diphosphate is reduced by endogenous ribonucleotide reductase to 5-aza-2'-deoxycytidine diphosphate (5-aza-dCDP), which is then phosphorylated to yield 5-aza-2'-deoxycytidine triphosphate (5-aza-tCDP). 5-aza-tCDP is incorporated into the negative-sense viral DNA during transcription and undergoes spontaneous cytosine ring-opening and deformylation, producing a guanylurea compound [27]. This guanylurea compound then base pairs with deoxycytidine (C) deoxyribonucleosides. Following the integration of mutated viral DNA into the host genome, 5-aza-tCDP is removed from the viral positive-sense DNA by endogenous DNA machinery and replaced with guanosine (G) nucleosides. During the subsequent transcription of viral RNA, the newly-inserted G nucleosides effect G-to-C transversion mutations in the synthesized positive-sense viral RNA [28]. The extensive G-to-C transversion mutations lead to error catastrophe and the collapse of HIV's replicative ability (Figure 6).
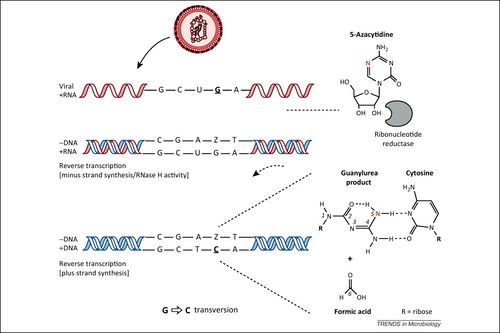
Decitabine and Gemcitabine
Two clinically-approved drugs for the treatment of cancer, decitabine (5-aza-2'-deoxycytidine) and gemcitabine (2'-deoxy-2', 2'-difluorocytidine), have been demonstrated to induce HIV lethal mutagenesis. Decitabine is a cytosine analog that induces G-to-C transversion mutations by a manner similar to that of 5-AZC. Gemcitabine is an irreversible ribonucleoside reductase inhibitor that augments the incorporation of decitabine into negative-sense viral DNA. When administered in tandem by Clouser and colleagues, decitabine and gemcitabine effected a 73% decrease in HIV infectivity. Clouser et al. reported that the application of the two drugs to HIV infected cells in vitro led to shifts in the HIV mutation spectrum secondary to increased rates of mutagenesis [29]. Importantly, the observed decrease in HIV infectivity could be attributed to error catastrophe following lethal mutagenesis by decitabine and gemcitabine and not the non-viability of affected viral DNA, because no decline in the rate of viral DNA synthesis was observed [28].
Combination decitabine and gemcitabine therapy for the treatment of HIV infection has recently performed favorably in pre-clinical evaluations. Both drugs were demonstrated to be non-toxic at therapeutic doses and successfully inhibited HIV infection in a murine AIDS model [30,31].
KP1461/KP1212
KP1461 is the prodrug of KP1212, a deoxycytidine nucleoside analog that induces A-to-G and G-to-A nucleoside transversion mutations when incorporated into viral DNA by a mechanism analogous to that of 5-AZC and decitabine. In phase II clinical trials, KP1212 reduced HIV titers to undetectable levels in cell culture after 9-24 rounds of viral replication. However, this reduction in HIV titers failed to persist, and all enrolled patients were seropositive for HIV by the end of the 125 day study period [32]. A 2014 study of KP1212 by Li and colleagues used advanced spectroscopic methods - including variable temperature NMR and 2D infrared spectroscopy - to further elucidate the mechanism of action of KP1212. Enol-keto tautaumerization, with enolic forms predominating, of KP1212 was found to influence the extent of A-to-G and G-to-A transversion mutations induced by the drug. This recent study may provide crucial insight into the mechanistic challenges faced by exogenous lethal mutagens and enable the translation of KP1212 into further clinical trials [33].
Lethal Mutagenic Agents and Drug Resistance
One important question concerning the safety and efficacy of lethal mutagenic agents is whether or not they may contribute to the propagation of drug-resistant HIV variants. Although there is a dearth of experimental evidence addressing this question in the context of lethal mutagenic drugs, studies investigating the action of APOBEC proteins and drug resistance may offer insight into the potential pitfalls of lethal mutagenic therapy. A multitude of studies have been conducted, both in silico and in vitro, which implicate APOBEC in the dissemination of HIV variants possessing augmented immune escape mutations [4]. In a 2008 study, Mulder and colleagues were able to isolate drug-resistant strains from cell cultures infected with HIV engineered to produce mutations in the Vif allele that partially impaired the function of ABOBEC3G [41]. Although these drug resistant HIV variants were only produced upon recombination with wild-type virions, the possibility that unchecked or accelerated mutagenesis may contribute the emergence of drug-resistant HIV strains cannot be excluded, since lethal mutagenic therapeutics are based on the activity of the APOBEC proteins.
Future Prospects for HIV Therapeutics Based on Lethal Mutagenesis
The induction of HIV lethal mutagenesis by novel therapeutics in infected hosts offers an exciting and important advance towards the attainment of a safe and effective HIV cure. The development of these therapeutics represents a concrete achievement based on the understanding and analysis of quasispecies theory as it pertains to HIV. The quasispecies model was first articulated by the German chemists Manfred Eigen and Peter Schuster in the 1970s [42]. The subsequent AIDS epidemic and the resulting breakneck pace of research into the mechanisms and nature of HIV infection and replication have combined fruitfully with quasispecies theory to produce a host of promising therapeutic agents within an astonishingly brief period. The concept of lethal mutagenesis is directly inspired by the notion of error threshold embedded within quasispecies theory. Although no therapeutics based on lethal mutagenesis are currently in the U.S. drug development pipeline, previously undertaken proof-of-concept research into lethal mutagenesis has nevertheless provided important insight into the nature of the HIV quasispecies and has elucidated a potential avenue for future drug design [43].
References
[1] "The Top 10 Causes of Death." World Health Organization. Web. 08 Nov. 2014. http://www.who.int/mediacentre/factsheets/fs310/en/
[2] "HIV/AIDS." World Health Organization. Web. 07 Nov. 2014. http://www.who.int/gho/hiv/en/
[3] Nabel, G. 2001. Challenges and opportunities for development of an AIDS vaccine. Nature. 410, 1002-1007. http://www.nature.com/nature/journal/v410/n6831/full/4101002a0.html
[4] Smythe, R. et al. 2012.The origin of genetic diversity in HIV-1. Virus Research. 169, 415-429. http://www.ncbi.nlm.nih.gov/pubmed/22728444
[5] Holland, J. et al. 1982. Rapid evolution of RNA genomes. Science. 215, 1577–1585. http://www.ncbi.nlm.nih.gov/pubmed/7041255
[6] Biebricher, C.K. and Eigen, M. 2006. What is a quasispecies? Curr. Top. Microbiol. Immunol. 299, 1–31. http://www.ncbi.nlm.nih.gov/pubmed/16568894
[7] Lauring, A.S. and Andino, R. 2010. Quasispecies Theory and the Behavior of RNA Viruses. PLoS Pathog 6(7): e1001005. http://www.plospathogens.org/article/info%3Adoi%2F10.1371%2Fjournal.ppat.1001005
[8] Abrahams, M.R. et al. 2009. Quantitating the multiplicity of infection with human immunodeficiency virus type 1 subtype C reveals a non-poisson distribution of transmitted variants. J. of Virol. 83, 3556-3567. http://www.ncbi.nlm.nih.gov/pubmed/19193811
[9] Slonczewski, Joan, and John Foster. "Microbiology: An Evolving Science" February 20, 2008. W.W. Norton and Company, New York. http://www.wwnorton.com/college/biology/microbiology2/
[10] Vignuzzi M, Stone JK, Andino R (2005) Ribavirin and lethal mutagenesis of poliovirus: molecular mechanisms, resistance and biological implications. Virus Res 107: 173–181. http://www.ncbi.nlm.nih.gov/pubmed/15649563
[11] Domingo E, Holland JJ (1997) RNA virus mutations and fitness for survival. Annu Rev Microbiol 51: 151–178. http://www.ncbi.nlm.nih.gov/pubmed/9343347
[12] Wilke CO (2005) Quasispecies theory in the context of population genetics. BMC Evol Biol 5: 44. http://www.biomedcentral.com/1471-2148/5/44
[13] Overbaugh J, Bangham CR (2001) Selection forces and constraints on retroviral sequence variation. Science 292: 1106–1109. http://www.ncbi.nlm.nih.gov/pubmed/11352065
[14] Biebricher CK, Eigen M (2005) The error threshold. Virus Res 107: 117–127. http://www.ncbi.nlm.nih.gov/pubmed/15649558
[15] Lee CH, Gilbertson DL, Novella IS, Huerta R, Domingo E, et al. (1997) Negative effects of chemical mutagenesis on the adaptive behavior of vesicular stomatitis virus. J Virol 71: 3636–3640. http://www.ncbi.nlm.nih.gov/pubmed/9094637
[16] Crotty S, Cameron CE, Andino R (2001) RNA virus error catastrophe: direct molecular test by using ribavirin. Proc Natl Acad Sci U S A 98: 6895–6900. http://www.pnas.org/content/98/12/6895.full
[17] Dapp, M.J. et al. 2013. Back to the future: revisiting HIV-1 lethal mutagenesis. Trends in Microbiol. 21, 56-62. http://www.ncbi.nlm.nih.gov/pubmed/23195922
[18] Greene, W. et al. 2013. Molecular biology of HIV: implications for new therapies. Sande's HIV/AIDS. Ch. 3. http://www.sciencedirect.com/science/book/9781455706952
[19] Stopak, K. et al. 2003. HIV-1 Vif blocks the antiviral activity of APOBEC3G by impairing both its translation and intracellular stability. Mol Cell. 12, 591-601. http://www.ncbi.nlm.nih.gov/pubmed/14527406
[20] Mariani, R. et al. 2003. Species-specific exclusion of APOBEC3G from HIV-1 virions by Vif. Cell. 114, 21-31. http://www.ncbi.nlm.nih.gov/pubmed/12859895
[21] Marin, M. et al. 2003. HIV-1 Vif protein binds the editing enzyme APOBEC3G and induces its degradation. Nat. Med. 9, 1398-403. http://www.ncbi.nlm.nih.gov/pubmed/14528301
[22] Sheehy, A.M. et al. 2003. The antiretroviral enzyme APOBEC3G is degraded by the proteasome in response to HIV-1 Vif. Nat. Med. 9, 1404-7. http://www.ncbi.nlm.nih.gov/pubmed/14528300
[23] Harris, R.S. et al. 2003. DNA deamination mediates innate immunity to retroviral infection. Cell. 113, 803-809. http://www.ncbi.nlm.nih.gov/pubmed/12809610
[24] Mangeat, D. et al. 2003. Broad antiretroviral defence by human APOBEC3G through lethal editing of nascent reverse transcripts. Nature. 424, 99-103. http://www.ncbi.nlm.nih.gov/pubmed/12808466
[25] Zhang, H. et al. 2003. The cytidine deaminase CEM15 induces hypermutation in newly synthesized HIV-1 DNA. Nature. 424, 94-98. http://www.ncbi.nlm.nih.gov/pubmed/12808465
[26] Jern, P. et al. 2009. Likely role of APOBEC3G-mediated G-to-A mutations in HIV-1 evolution and drug resistance. PLoS Pathog. 5, e1000367. http://www.plospathogens.org/article/info%3Adoi%2F10.1371%2Fjournal.ppat.1000367
[27] Yu, X. et al. 2003. Induction of APOBEC3G ubiquitination and degradation by an HIV-1 Vif-Cul5-SCF complex. Science. 302, 1056-60. http://www.ncbi.nlm.nih.gov/pubmed/14564014
[28] Bouchard, J. et al. 1990. 5-Azacytidine and 5-azadeoxycytidine inhibit human immunodeficiency virus type 1 replication in vitro. Antimicrob. Agents Chemother. 34, 206-209. http://www.ncbi.nlm.nih.gov/pubmed/1691617
[29] Rogstad DK, et al. 2009. Chemical decomposition of 5-aza-2'-deoxycytidine (Decitabine): kinetic analyses and identification of products by NMR, HPLC, and mass spectrometry. Chem. Research in Tox. 22, 1194–1204. http://www.ncbi.nlm.nih.gov/pubmed/19480391
[30] Clouser CL, et al. 2010. Exploiting drug repositioning for discovery of a novel HIV combination therapy. J. of Virol. 84, 9301–9309. http://www.ncbi.nlm.nih.gov/pubmed/20610712
[31] Clouser CL, et al. 2011. Analysis of the ex vivo and in vivo antiretroviral activity of gemcitabine. PLoS One. 6:e15840. http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0015840
[32] Clouser CL, et al. Activity of a novel combined antiretroviral therapy of gemcitabine and decitabine in a mouse model for HIV-1. Antimicrob Agents Chemother.2012;56:1942–1948. http://www.ncbi.nlm.nih.gov/pubmed/22271861
[33] Mullins J. et al. 2011. Mutation of HIV-1 genomes in a clinical population treated with the mutagenic nucleoside KP1461. PLoS One. 6, e15135. http://www.ncbi.nlm.nih.gov/pubmed/21264288
[34] Li, D. et al. 2014. Tautomerism provides a molecular explanation for the mutagenic properties of the anti-HIV nucleoside 5-aza-5,6-dihydro-2'-deoxycytidine. PNAS. 111, E3252–E3259. http://www.ncbi.nlm.nih.gov/pubmed/25071207
[35] Zhang et al. 2004. Host RNA polymerase II makes minimal contributions to retroviral frame-shift mutations. J. Gen. Virol. 85:2389-2395. http://www.ncbi.nlm.nih.gov/pubmed/15269381
[36] Ivanchenko et al. 2009. Dynamics of HIV-1 assembly and release. PLoS Path. 5 (11) (2009), p. e1000652. http://www.plospathogens.org/article/info%3Adoi%2F10.1371%2Fjournal.ppat.1000652
[37] Paillart et al. 1994. Mutational analysis of the bipartite dimer linkage structure of human immunodeficiency virus type 1 genomic RNA. J. Biol. Chem. 269:27486-27493.
[38] Jung et al. 2002. Multiply infected spleen cells in HIV patients. Nature. 418:144. http://www.ncbi.nlm.nih.gov/pubmed/12110879
[39] Englund et al. 1995. Integration is required for productive infection of monocyte-derived macrophages by human immunodeficiency virus type 1. J. Virol. 69:3216-3219. http://www.ncbi.nlm.nih.gov/pubmed/7707554
[40] Carlson et al. 2014. Selection bias at the heterosexual HIV-1 transmission bottleneck. Science. 345. http://www.ncbi.nlm.nih.gov/pubmed/25013080
[41] Mulder et al. 2008. Cytidine deamination induced HIV-1 drug resistance. PNAS. 105:5501-5506. http://www.ncbi.nlm.nih.gov/pubmed/18391217
[42] Nowak, A. M. 2002. From quasispecies to universal grammar. Z. Phys. Chem. 5-20. http://www.ped.fas.harvard.edu/people/faculty/publications_nowak/ZPhysChem02.pdf
[43] "Treatments for HIV & AIDS." AIDSMEDS. Web. 17 Dec. 2014. http://www.aidsmeds.com/list.shtml
[44] Frange P, Meyer L, Jung M, Goujard C, Zucman D, et al. (2013) Sexually-transmitted/founder HIV-1 cannot be directly predicted from plasma or PBMC-derived viral quasispecies in the transmitting partner. PLoS ONE 8(7): e69144. doi:10.1371/journal.pone.0069144.
http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0069144
Page authored by Benji Dossetter, B.A. '15 for BIOL 375 Virology, Kenyon College, December, 2014.
