Rhizobium Nodulation and Evolution
Introduction
Lack of Nitrogen is a limiting factor for the growth of plants as Nitrogen incorporated in the biomass. Legumes plants have overcome this problem by incorporating nitrogen fixing bacteria, Rhizobium, inside their legumes. In the Rhizobium-plant relationship, the plant provides carbohydrates to the Rhizobium species while the Rhizobium provides fixed Nitrogen for the plant.
The ecological habitat of Rhizobia species varies from Arid deserts (Zahran 1999) to Tropical rainforests (Woomer et. al. 1988). Many different factors affect the growth of Rhizobium species, such as pH of soil, soil temperature (Woomer et. al. 1988), and amount and frequency of precipitation (Higashi et al. 2008). As these factors differ based on geography, various species of rhizobium have evolved appropriately.
The Ability of Rhizobia to fix nitrogen can help in preventing denudation, increasing crop harvest and reducing de-nitrification. This ability has made rhizobia an important part of modern agriculture and a hot topic for research. The aim of this paper is to review the compounds involved in nodulation and the evolution of Rhizobium species.
Nodulation
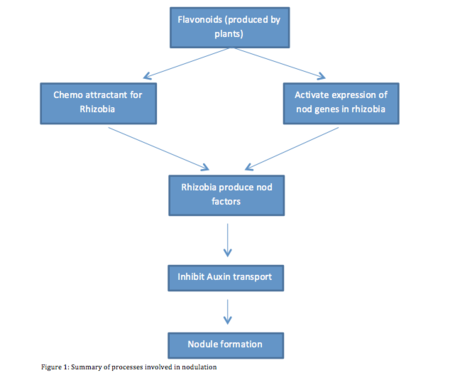
Nodulation is a complex process, which involves chemical conversation between the Rhizobium and host. During the nodulation process, both Rhizobia and legumes plant interact closely. A group of ubiquitous and diverse molecules called Flavonoids are among the chemicals that facilitate the interactions between Rhizobia and legumes plant. Flavonoids play several roles during the different stages of nodulation. They concentrate rhizobia near the roots and activate the synthesis of the Nod signal, which is essential for nodule formation. Another function of Flavonoids is the regulation of Auxin (Subramanian et. al. 2007). The Amount of Auxin present during nodulation directly affects nodulation and determines the shape of the nodule (spherical or cylindrical). A summary of the chemical conversation is schematically illustrated in Figure 1.
Another Important chemical group that facilitates the close interaction of Rhizobia and legumes is Lipochitooligosaccharides, which are made from Beta-1,4 N-acetyl-D-glucosamine attached to a fatty acyl chain. The various forms of Lipochitooligosaccharides are the result of different oligosaccharide backbone length, the molecule groups carried by the organism and the type of fatty acid. Lipochitooligosaccharides are responsible for a variety of important functions, such as root hair deformation, calcium spiking, nodule development and much more. There is compatibility between Lipochitooligosaccharides structure made by rhizobia and the host plant receptors; nodulation does not occur in the absence of specific Lipochitooligosaccharides features (Cooper 2007).
Although Lipochitooligosaccharides have been found to be extremely important for development of nodules, some rhizobium organisms have been observed to make nodules even in the absence of the Lipochitooligosaccharides gene. By sequencing the DNA of Photosynthetic Bradyrhizobium strains extracted from the nodules of plants in North America and Africa, it was found that the genome of Bradyrhizobium includes neither Nod A nor Nod C genes. Nod A is an Acetyl Transferase and Nod C is responsible for Oligomerization of N-acetyl-glucosamine. The Bradyrhizobium does not have two of the three enzymes essential for the synthesis of Lipochitooligosaccharides. It has been found that another molecule derived from purine which acts like a cytokine, is responsible for the formation of nodules in Bradyrhizobia (Giraud et. al. 2007). This finding suggests that Lipochitooligosaccharides are not the only chemicals involved in the chemical conversation.
Evolution of Rhizobium species
There are evidences suggesting that Rhizobium bacteria acquired their symbiotic capabilities through horizontal gene transfer (Martínez-Romero 2009).
One evidence supporting this hypothesis emerges when analyzing the ribosomal RNA and constructing a phylogenetic tree. Organisms of the same families have similar ribosomal RNA because these sequences are conserved and variations in these sequences could be lethal. This dictates that organisms with a common ancestor must have similar ribosomal RNA. Hence, they are excellent in producing a phylogenetic tree. By sequencing ribosomal RNA, it has been observed that rhizobium organism do not group together. In fact, they are dispersed among organisms of different families (Amadou et. al. 2008). If the symbiotic characteristic of rhizobium organisms were a result of vertical gene transfer, one would expect all of the organisms with a common ancestor to have rhizobia characteristics. This observation indicates that the symbiotic characteristic attributed to rhizobia organisms is not a result of vertical gene transfer.
In addition, essential genes that are responsible for the symbiotic characteristic of rhizobium species are present on plasmids. These plasmids can be transferred from one organism to another (Brom et. al. 2004). Moreover, it was shown that when a plasmid is transferred from one organism to another, the rhizobia characteristic is transferred to the host. In an experiment symbiotic plasmids where taken from Rhizobium tropici and placed inside the organism Ensifer adhaerens, a non-nodule forming bacterium that attaches to other bacteria and can cause lysis. After E. adhaerens received the plasmid from R. tropici, it surprisingly formed nodules on the bean plants (Rogel et. al. 2001).
To sum up, Rhizobium organisms do not have a common ancestor and the symbiotic characteristic can be transferred from one organism to another. These findings suggest that the symbiotic activity of Rhizobia species is a result of Horizontal gene transfer.
Refrences
1. Amadou, C., A. Lajus, M. Marchetti, V. Poinsot, Z. Rouy, B. Servin, M. Saad, C. Schenowitz, V. Barbe, J. Batut, C. Médigue, G. Pascal, C. Masson-Boivin, S. Mangenot, M. Glew, C. Bontemps, D. Capela, S. Carrère, S. Cruveiller, and C. Dossat. 2008. Genome sequence of the beta-rhizobium Cupriavidus taiwanensis and comparative genomics of rhizobia. Genome Res. 18:1472-1483. doi: 10.1101/gr.076448.108.
2. Brom, S., L. Girard, C. Tun-Garrido, A. García-de los Santos, P. Bustos, V. González, and D. Romero. Transfer of the Symbiotic Plasmid of Rhizobium etli CFN42 Requires Cointegration with p42a, Which May Be Mediated by Site-Specific Recombination. .
3. Cooper, J. E. 2007. Early interactions between legumes and rhizobia: disclosing complexity in a molecular dialogue. J. Appl. Microbiol. 103:1355-1365. doi: 10.1111/j.1365-2672.2007.03366.x.
4. Giraud, E., Y. Prin, G. Bena, L. Hannibal, J. Fardoux, M. Kojadinovic, L. Vuillet, A. Lajus, S. Cruveiller, Z. Rouy, S. Mangenot, L. Moulin, B. Segurens, C. Dossat, W. L. Franck, W. Chang, E. Saunders, D. Bruce, P. Richardson, P. Normand, B. Dreyfus, D. Pignol, D. Vallenet, G. Stacey, D. Emerich, A. Verméglio, C. Médigue, M. Sadowsky, V. Barbe, E. Cytryn, J. Avarre, M. Jaubert, D. Simon, and F. Cartieaux. 2007. Legumes symbioses: absence of Nod genes in photosynthetic bradyrhizobia. Science (New York, N.Y.). 316:1307-1312. doi: 10.1126/science.1139548.
5. Higashi, S., T. Uchiumi, and M. Abe. 1983. Elimination of Rhizobium infectivity by temperature treatment. J. Gen. Appl. Microbiol. 29:281-285.
6. Martínez-Romero, E. 2009. Coevolution in Rhizobium-legume symbiosis? DNA Cell Biol. 28:361-370. doi: 10.1089/dna.2009.0863.
7. Rogel, M. A., I. Hernández-Lucas, L. D. Kuykendall, D. L. Balkwill, and E. Martinez-Romero. 2001. Nitrogen-fixing nodules with Ensifer adhaerens harboring Rhizobium tropici symbiotic plasmids. Appl. Environ. Microbiol. 67:3264-3268. doi: 10.1128/AEM.67.7.3264-3268.2001.
8. Subramanian, S., G. Stacey, and O. Yu. 2007. Distinct, crucial roles of flavonoids during legume nodulation. Trends Plant Sci. 12:282-285. doi: 10.1016/j.tplants.2007.06.006.
9. Woomer, P., P. W. Singleton, and B. B. Bohlool. 1988. Ecological indicators of native rhizobia in tropical soils. Appl. Environ. Microbiol. 54:1112-1116.
10. Zahran, H. H. 1999. Rhizobium-Legume Symbiosis and Nitrogen Fixation under Severe Conditions and in an Arid Climate. Microbiology and Molecular Biology Reviews. 63:968-968.