Simian Immunodeficiency Virus – Research subjects for vaccine and drug development and models for the origin of HIV and AIDS
Overview

Simian immunodeficiency viruses (SIV) are a group within the Lentiviridae genus of retrovirus family, (Retroviridae) whose genome is ribonucleic acid (RNA) but reverse transcribe to deoxyribonucleic acid (DNA) intermediates before making viral proteins. This viral group is found naturally infecting more than 40 African monkeys (or Old World monkeys – Cercopithecidae family) and 2 species of great apes, namely chimpanzee (Pan troglodytes) and gorilla (Gorilla gorilla)[3]. SIV are the most closely related viruses to the human immunodeficiency virus type 1 and 2 (HIV–1 and HIV–2), which cause acquired immunodeficiency syndrome (AIDS) in humans [4]. Despite being called “immunodeficiency viruses”, SIV do not usually result in any AIDS-like syndromes in many of their natural African monkey host communities, such as sooty mangabeys (Cercocebus atys) and Colobus monkeys (Colobinae subfamily) even with high viral counts in infected individuals and high prevalence of infection (up to 59% of a community can be infected)[2][5][6][7]. This suggests that SIV have a long history of coevolution with these natural hosts. However, there has been evidence suggesting recent host jumps of SIV from their long-adapted hosts to new host species such as chimpanzees in the wild, macaques in captives (an example of Asian monkeys who have much less exposure to SIV than the African ones) and humans, where they developed into new strains through combinations and mutations and became pathogenic in the new host [8][9][10][11]. In macaques (Macaca spp.) and chimpanzees (Pan troglodytes), SIV infection can cause simian acquired immunodeficiency syndrome (SAIDS) similar to AIDS in human[8][12]. Thus, researches into SIV, in both their natural reservoir and laboratory model primates, not only reveals the origin of HIV and AIDS, but also help further understandings in HIV’s pathology and developments of HIV and AIDS preventions and treatments.
Taxonomy
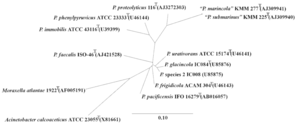

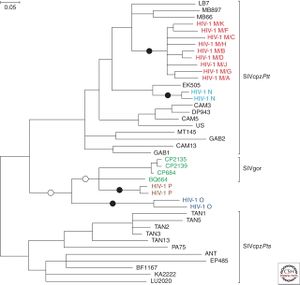
SIV are defined as lentiviruses (which itself belongs to the subfamily Orthoretrovirinae in the Retroviridae virus family) that specifically infect non-human primates (simians). The Lentiviridae genus includes a wide variety of primate infecting viruses and a few other viral species that infect felines, cows, horses, goats, and sheep[19][6][20]. Infection of lentiviruses are characterized by long dormant periods (infection may never progress to disease phase as in the case of some natural host monkeys), progressive disease development, permanent damage of tissues, organs, or systems, and, in many cases, lethal outcomes[19]. SIV are a polyphyletic group in the primate lentivirus that also include HIV-1 and HIV-2 (Figure 2)[4]. The SIV species in chimpanzee (SIVcpz) is closely related to the HIV-1 groups M and N[21], while HIV-1 group O is more closely related to the SIV species in gorillas (SIVgor)[22]. And the SIV species that is naturally non-pathogenic in sooty mangabeys (SIVsm) has been shown to be ancestral to HIV-2 and SIV species that cause lethal SAIDS in Asian macaque monkey species [19][23]. A full phylogeny of SIV along with different types of HIV and other non-primate lentiviruses are demonstrated in Figure 2 (species relatedness was determined by the molecular clock gene pol)[4].
Structure - SIV in non-human primates as an effective and accessible animal model for HIV research
SIV share characteristic traits in morphology and morphogenesis with other members of the lentivirus genus[24]. Genetically, the closest-related viral species to SIV are HIV-1 and HPV-2[21][22][23], the other two members of the primate lentivirus subgenus[4] and the most well-studied species among lentiviruses. Thus, according to a model of lentivirus structured based on HIV-1 and HIV-2, an SIV virion outside of a host cell, along with other lentiviruses’ virions, are either spherical or pleomorphic. SIV virion likely has an outer-membrane derived from host protein with viral spike proteins that bind specifically with host species of each SIV species. Inside the outer-membrane is a protein capsid containing a tightly coiled positive (+) single-stranded RNA genome and a few crucial viral proteins for controlling the host cell’s defense system, reverse transcription, and integration into the host genome, such as reverse transcriptase, integrase, and tRNA lyase. Once inside host cells, SIV and other lentiviruses do not exist as distinct virus particles but reverse transcribe their RNA genomes into double-stranded DNA and integrate into the host genome with finesse to not cause any lethal mutations or trigger any oncogenes in host cells. Then, the viral double-stranded DNA would exploit the host’s transcribing and translating system to transcribe viral proteins and assemble and disperse progeny virions[25].
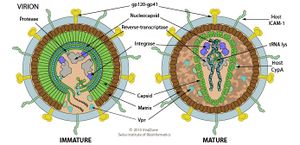
SIV in non-human primates as effective and accessible animal models for HIV research
Due to the close resemblance and relatedness of SIV to HIV, the most similar anatomies of SIV’s hosts (non-human primates) and human, and the ability of some SIV species to induce simian AIDS in the common primate model organisms, the Asian macaques (Macaca spp.), with rapid disease progression and pathology similar to human AIDS, SIV effectively serve as models to study HIV and AIDS in animals[26][12][10]. However, since the SIV species that infect Asian macaques are in the lineage SIVsmm/HIV-2 (Figure 2), which is relatively distant from the common global pandemic HIV-1 species, the macaques model still has certain differences in disease progression and drug response from the more common HIV-1[26] that researchers need to keep in mind when adopting this model organism for HIV researches. On the other hand, the closest animal model to HIV-1, and probably HIV-1-caused AIDS, has been shown to the SIVcpz found in the natural reservoir of the central chimpanzee subspecies (Pan troglodytes troglodytes), which has its own distinct type of SIVcpz infecting only that one subspecies[2]. Previously, all SIV found in natural reservoirs were not thought to be pathogenic, however, fecal sampling for SIV antibodies and behavioral studies in feral chimpanzees revealed that SIVcpz infection in wild chimpanzees can be pathologic and can progress to SAIDS with histopathology similar to what is seen in human’s AIDS[8]. Moreover, transmission of SIVcpz in the wild is also well-documented to be mostly sexual transmission and mother-to-infants transmission, similar to what is observed in humans[8][27]. One limit to this SIV model is that chimpanzees (Pan troglodytes) and their counterparts, the bonobos (Pan paniscus), are rare, endangered species, thus limiting researchers’ ability to study them in laboratory settings. However, chimpanzee species and subspecies are the best-studied primate species in the wild with many groups studying chimpanzee and bonobos communities for multiple decades[28]. Thus, studying virology and pathology of SIVcpz in the wild and documenting behaviors and social interactions of infected chimpanzee communities in the wild can produce great insights into the virology, transmission, pathology, and social factors related to the transmission of HIV-1 among humans
Evolution of SIV
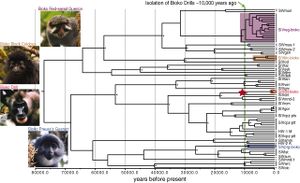
Molecular efforts in dating the age of the SIV, or more precisely, the primate lentivirus lineage, based on the modeled mutation rate of HIV-1 and HIV-2 (the most well-studied primate lentiviruses) have bred great controversies. Dating methods using nucleotide molecular clocks that use the mutation rate of HIV-1 and HIV-2 as standard references suggest the most recent common ancestor of the primate lentivirus to be around a few hundreds to a few thousands of years before present[30][9]. This contradicts the pathological and epidemiological evidences suggesting SIV might have evolved in a host-dependent manner to be non-pathogenic in some African monkeys [2][5][6][7]. However, a study in 2010[29] combining the viruses’ molecular clock dating and phylogeny analysis with the host species’ phylogenic studies and biogeographical history suggests that the SIV lineage has existed for at least 32,000 years. Another research into the existence of SIV in the ring-tailed lemur of south-western Madagascar showed the existence of antigens against SIV in ring-tailed lemurs here. If there was no cross-contamination of SIV from mainland and SIV in primates evolved in a host-dependent way from a common ancestor (as seen in the case of green monkeys (Chlorocebus spp.))[7], this can push the possible date of the oldest common ancestor of the SIV lineage to 85 million years ago, which is the approximate date of the last primate common ancestor and is also around the time the total separation of Madagascar from the Indian continent happened[31].
This direct and indirect evidence of SIV’s historic existence among the primates, especially those of the more ancient orders (monkeys and lemurs) who SIV have evolved natural co-existing relationships with, suggest a terrible bias in the HIV-based molecular clock assay, which points to a possible, drastic increase in HIV comparing to their more ancient SIV counterparts. On the other hand, it’s also HIV’s notoriously rapid mutation rate that is the main obstacle against the development of vaccines against HIV and the primary cause of drug resistance in treatments. Thus, further research into the infections and mutating and replicating activities of older, naturally non-pathogenic SIV species can help us understand the genetic difference that enables the rapid mutation rate in their human-infecting counterparts, as well as develop vaccines against HIV. Moreover, considering the long existence of SIV (at least 32,000 years ago), it is likely that humans have come into contact with SIV multiple times and cross-transmission of SIV to human are likely to have occurred many times throughout the past 32,000 years. Thus, there must be recent social and biological factors that the rapid spread of HIV only during the past 100 years or so[32][33].
The origin of Human Immunodeficiency Virus (HIV) from SIV
Despite both being called HIV and causes AIDS in human, HIV-1 and HIV-2 are only distantly related in the primate lentivirus group (Figure 2)[4][34]. Each type of HIV belongs to a different lineage of primate lentivirus that contains other SIV species, some of which are considered to be the predecessors or each type of HIV[3]. Additionally, within HIV-1, there are also 3 major epidemic groups M, N, an O, with groups M and N having a different SIV predecessor from group O[3][21][35]. These are consistent with the widely accepted knowledge that HIV strains have arisen multiple times through multiple cross-transmission events from non-human primates.
Surprisingly, HIV-2, which was initially found in West Africa[36], was the viral species whose origin was first determined. In the beginning, the SIV species in captive rhesus macaques (Macaca mulatta) (SIVmac) was determined to be the most closely related to human’s HIV type-2. However, feral rhesus macaques were not found to carry SIV, and the infect macaques in captive died quickly of SAIDS[10], which suggested that they had only acquired SIV recently. In 1989, the SIV species closely related to the human’s HIV type 2 was discovered naturally infecting the sooty mangabeys (Cercocebus atys) of West Africa. Despite its host-jumped forms being gravely virulence in their new hosts (the Asian macaques), the sooty mangabey’s SIV (SIVsmm) is not virulent in its natural host. In captive environments, SIVsmm must have spread from its natural host to various species of macaques, and eventually human[37]. The interspersed origins of different subtypes of HIV-2 among the SIVsmm/HIV-2 lineage suggest that there had been multiple cross-species transmission events of SIVsmm and other SIV species in captive macaques (Macaca sp.) onto humans. However, only the subtype A and B became pandemic in human[2]. Even though soot mangabeys are widely accepted as the natural reservoir for HIV-2 predecessors, HIV-2 might not have resulted directly from cross-transmission from sooty mangabeys. Genetic comparison shows that the SIV species from Asian stump-tail macaques in captive (SIVstm), not SIVsmm, is the most closely related to HIV-2[38][4]. Thus, HIV-2 seems to be a result of secondary cross-transmission facilitated by the close and frequent interactions between humans and monkeys in the huge monkey export industry that serves pharmaceutical testing in the mid-20th century.
HIV-1’s origin, on the other hand, is more complicated. Among the epidemic HIV-1 groups, group M is the most prevalent, accounting for the majority of infections[3]. Despite the discoveries of SIVcpz strains closely related to HIV-1 group M in 1989[39] and multiple suspicions of the chimpanzee origin of HIV-1 group M (then known only as HIV-1) that follow suits[9], there were still hesitant to conclude on the origin of HIV-1 group M’s origin as no natural infection of chimpanzees with SIVcpz had been detected at that time[2]. The origin of HIV-1 group M, as well as group N, which was discovered later in 1998[40], was only confirmed in the latter half of the 2000s to be from the specific strain of SIVcpz of the central chimpanzee subspecies (Pan troglodytes troglodytes) (SIVcpzPtt)[2][21]. Genetic analysis showed that both HIV-1 group M and N are very closely related to SIVcpzPtt variants from southern Cameroon, and the 2 subtypes of HIV-1 are interspersed among the variants of SIVcpzPtt (Figure 5)([21][41]. This confirmed that HIV-1 group M and N originated from SIVcpzPtt in the central chimpanzee subspecies from two distinct host-jump events. Such findings also made it possible to track the approximate location of the cross-transmission [3][21][41]. Nonetheless, it is unclear how exactly human acquired HIV-1 type M and N from chimpanzees, but it was possibly in the context of bushmeat hunting and consumption, which is the most likely situation where humans can have cutaneous or mucous exposure to infective blood or mucosal fluids of SIVcpzPtt-infected chimpanzees [3]. Not only facilitated the cross-transmission of SIVcpzPtt to humans, hunting activities are also possible channels for the chimpanzees to acquire their SIV species in the first place. SIVcpz likely hasn’t had a very long host-dependent evolution time comparing to non-pathogenic SIV strains endemic in Old World monkeys (like green monkeys or mangabeys) because it still causes grave health and reproductive damages to its infected chimpanzee communities, and the distribution of SIVcpz among chimpanzees is much less even among communities [2][21][8]. Thus, SIVcpz, despite having infected its host for a much longer period of time than HIV[3][9], must be a rather recent (comparing to SIV in Old World monkeys) acquisition of chimpanzees. Genetic analysis showed that SIVcpz is a recombination of SIV species in red-capped mangabeys (Cercocebus torquatu) and greater spot-nosed monkeys (Cercopithecus nictitans), that likely resulted from a co-infection of SIV and SIV due to the chimpanzees’ hunting and consumption of those monkeys ([42] reviewed in Sharp et. al, 2010[3]).
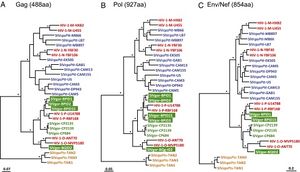
HIV-1 group O, as well as the minor group P (only 2 infections identified so far[43]), however, has been recently determined to have originated from the SIV species infecting gorillas (Gorilla gorilla) (SIVgor) instead of SIVcpz. This discovery was enabled by the development and application of a non-invasive sampling technique to detect antibodies against SIV in fecal and urine samples and to further construct complete provirus from the antibodies’ sequences (reviewed in Sharp & Hahn, 2010[2])[21][22][43], which allows for large-scale studies of SIV prevalence and pathology in endangered ape species such as the chimpanzees (Pan troglodytes) and gorillas in the wild. In the large-scale study of SIV in central Africa gorilla communities, D’arc’s team[43] showed that: like the case of SIVcpz, SIVgor is also restricted to infecting only a certain population of wild gorillas in the southern western lowlands of southern Cameroon. And among the variants of SIVgor found in this area, the research team found variant strains of SIVgor, SIVgor-BQID2 and SIVgor-BPID1 & BPID15, that are the closest related viruses to HIV-1 group O and P, respectively, based on full-length genomic sequencing and phylogenic comparisons using different highly conserved genes in lentivirus (Figure 6)[43]. The amount of divergence between these SIVgor strains and their most closely related HIV-1 groups are comparable to what was observed among HIV-1 group M and N and their possible SIVcpzPtt predecessor strains (Figure 6)[43][3]. The fact that HIV-1 group O and P are interspersed among the SIVgor/HIV-1 O & P clade (Figure 6) tells us that this two groups of HIV-1, similar to the case of group M and N, arose from 2 different cross-transmission events from gorillas to humans. Genetic analyses also show that SIVgor and HIV-1 groups and P belong in the larger lineage of SIVcpz/HIV-1. However, SIVgor, along with strains of HIV-1 group O and P, forms a parallel clade to the SIVcpzPtt/HIV-1 M and N clade (Figure 6). Both of these clades are more closely related to each other than to the clade of SIVcpz in the western chimpanzee subspecies Pan troglodytes schweinfurthii (SIVcpzPts) which seems to have not given rise to any group of HIV-1 until this point (Figure 6)[43]. These evidences further demonstrate that the SIVgor itself might also have developed from cross-transmission of SIVcpzPtt[22][43]. Furthermore, the fact that all strains of SIVgor cluster together into a monophyletic group in phylogenic analyses based on DNA and protein sequences suggests that not only did SIVgor arise from cross-transmission of SIVcpzPtt to gorilla, but there was only one such cross-transmission event that gave rise to all the SIVgor strains in western lowland gorillas[22][43]. Unlike the case of SIVcpz and HIV, there are no prominent hypothetical scenarios for the gorillas’ acquiring of SIVgor, as gorilla is strictly herbivorous and secluded, preferring to avoid other primates (reviewed in D’arc et. al., 2015,[43]). However, transmission is still possible due to the overlapping foraging range of western lowland gorillas and many troops of central chimpanzees (Pan troglodytes troglodytes) (reviewed in D’arc et. al., 2015[43]). Transmission from gorillas to humans and the emergence of HIV-1 group O and P may have been due to human hunting activities, however, no possible explanations have been officially proposed. It is also probable that gorillas may suffer SAIDS and other pathogenic impacts due to SIVgor, however, due to the even lower prevalence of SIVgor comparing to that in chimpanzees (2% overall prevalence comparing to 6% in central and 13% in eastern chimpanzees) (reviewed in D’arc et. al., 2015[43]) and the less accessible habitat range of gorillas, there has been much less research into possible pathogenicity of SIVgor in gorillas.
Host adaptations of SIV into apes (chimpanzees and gorillas) and human (as HIV)
Despite SIV’s widespread infections among the primate species, especially the Old World monkeys (Cercopithecidae family), infection of SIV and their successor HIV in human has only occurred relatively recently and only in certain communities[43] (reviewed in Sharp & Hahn, 2010[2]). This probably is a result of the highly complex and developed immune system of primates, especially the apes (including human), with various restriction factors in the innate immune system that inhibit the infection, replication, and transmission of viruses, which SIV and HIV have to neuter or antagonize in order to successfully replicate and spread within a host (reviewed in Sharp & Hahn, 2010[2]). Three primate interferons that have been shown to hinder SIV/HIV cross-species infection are APOBEC3G (A3G) (interferes with reverse transcription[44]), TRIM5α (interferes with viral uncoating[45]), and tetherin (inhibit progeny viral particles from budding out of infect cells[46]).
In the cross-transmission of SIVcpzPtt from chimpanzees (Pan troglodytes) into gorillas (Gorilla gorilla), A3G interferon seems to have been the most challenging restricting factor. In chimpanzees, the A3G interferon is antagonized by the SIVcpz’s vif protein and eventually degraded. However, once in gorillas, the vif-mediated degradation by SIVpz’s vif protein variants is entirely inactive or only minimally active against gorillas’ A3G. In addition, gorillas’ A3G is only specifically susceptible to the SIVgor’s vif protein. This suggests that mutations in the vif gene that modified SIVcpz's vif protein to be more compatible with gorilla's tetherin was vital for the host-jump from chimpanzees to gorillas. This also further explains why there was only one successful cross-transmission of SIV from chimpanzees to gorillas in the past, even though central wetland gorillas have habitat ranges that overlap with many troops of central chimpanzees (Pan troglodytes troglodytes)[43].
Another difference from the HIV-1 M&N/SIVcpzPtt lineage within the clade of HIV-1 O&P/SIVgor is that HIV-1 group O neuter human’s tetherin using the protein nef[47] similar to that seen in most SIV species, even SIVcpz and SIVgor[43], but different from HIV-1 group M and N, which use the vpu protein for this function[48]. Because the tetherin motif that the original nef proteins of SIVcpz, SIVgor, and most other SIV species target is deleted in human, there must have been important changes to the nef protein of HIV-1 group O, and possibly group P as well, so that it effectively targets another region of human’s tetherin and facilitate host-jumping from gorillas to humans[43].
Nonetheless, changing the nef protein to target another region of tetherin was not the only approach to adapting against human’s immune system for SIV when host-jumping to humans. In HIV-1 group M and N and SIV species of spot-nosed, mona, mustached, and Dent’s monkeys uses vpu protein instead of the nef protein as the antagonist to host’s tetherine[48][49]. While nef protein of HIV-1 group M shows potent antagonistic activity against human’s tetherin, the nef protein of HIV-1 group N only shows marginal activity[48]. This can explain why HIV-1 group M causes a global pandemic and HIV-1 group N has only infected 20 people as of 2011[3][50]. Moreover, species of HIV and SIV have multiple pathways to neuter their primate hosts’ tetherins (reviewed in Sharp & Hahn, 2010[2]). This suggests that the selection for an effective response to antagonize host’s tetherin plays at least a significant role in the immediate host-specific selection after SIV or HIV species have transmitted cross-species.
Conclusion
Overall, since HIV directly originated from some species of SIV (as discussed above), SIV in non-human primates are the closest natural animal models to human’s HIV. However, employment of non-human primate SIV models in studying HIV pose substantial challenges. The animal models that, possibly, most closely resemble the major HIV strains, HIV type 1 groups M and O, are chimpanzees and gorillas, which are greatly endangered primate species. Thus, they cannot be used for laboratory researches, even invasive testing on feral individuals are greatly discouraged. Nonetheless, much can be learned from studying the biology, pathology, and epidemiology of SIV infections in these simian models in wild communities, which, conveniently, are already of great research interests among the anthropology and ecology scientific communities. This can be accomplished thanks to advanced animal tracking systems and non-invasive testing methods recently developed. Due to the limits of an undergraduate intermediate-level topic-review project, only the evolution and structure of SIV, along with their connections to HIV and HIV research, are discussed on this page at the moment. Yet, SIV is a vast research topic receiving great efforts and attentions from the scientific communities. Researches withing the past five decades have generated a large amount of knowledge in the topic, a lot of which has not been discussed on this page at the moment. For future development of this wiki page, the primary author suggests additions of sections such as “Pathology of SIV – Evolution of AIDS” or “Transmission of SIV, Vaccine against SIV – Possible SIV models to develop HIV vaccine”.
References
- ↑ Murphey-Corb, Michael, Louis N. Martin, S. R. S. Rangan, Gary B. Baskin, Bobby J. Gormus, Robert H. Wolf, W. Abe Andes, Melanie West, and Ronald C. Montelaro. 1986. “Isolation of an HTLV-III-Related Retrovirus from Macaques with Simian AIDS and Its Possible Origin in Asymptomatic Mangabeys.” Nature 321(6068):435–37.
- ↑ 2.00 2.01 2.02 2.03 2.04 2.05 2.06 2.07 2.08 2.09 2.10 2.11 Sharp, Paul M., and Beatrice H. Hahn. 2010. “The Evolution of HIV-1 and the Origin of AIDS.”
- ↑ 3.00 3.01 3.02 3.03 3.04 3.05 3.06 3.07 3.08 3.09 3.10 Sharp, Paul M., and Beatrice H. Hahn. 2011a. “Origins of HIV and the AIDS Pandemic.” Cold Spring Harbor Perspectives in Medicine 1(1):a006841.
- ↑ 4.0 4.1 4.2 4.3 4.4 4.5 4.6 Perrone, Rosalba, Enrico Lavezzo, Giorgio Palu, and Sara Richter. 2017. “Conserved Presence of G-Quadruplex Forming Sequences in the Long Terminal Repeat Promoter of Lentiviruses.” Scientific Reports 7.
- ↑ 5.0 5.1 Santiago, Mario L., Friederike Range, Brandon F. Keele, Yingying Li, Elizabeth Bailes, Frederic Bibollet-Ruche, Cecile Fruteau, Ronald Noë, Martine Peeters, John F. Y. Brookfield, George M. Shaw, Paul M. Sharp, and Beatrice H. Hahn. 2005. “Simian Immunodeficiency Virus Infection in Free-Ranging Sooty Mangabeys (Cercocebus Atys Atys) from the Taï Forest, Côte d’Ivoire: Implications for the Origin of Epidemic Human Immunodeficiency Virus Type 2.” Journal of Virology 79(19):12515–27.
- ↑ 6.0 6.1 6.2 Peeters, Martine, and Valerie Courgnaud. 2002. “Overview of Primate Lentiviruses and Their Evolution in Non-Human Primates in Africa.” HIV Sequence Compendium.
- ↑ 7.0 7.1 7.2 Beer, Brigitte E., Elizabeth Bailes, Robert Goeken, George Dapolito, Cheik Coulibaly, Stephen G. Norley, Reinhard Kurth, Jean-Pierre Gautier, Annie Gautier-Hion, and Dominique Vallet. 1999. “Simian Immunodeficiency Virus (SIV) from Sun-Tailed Monkeys (Cercopithecus Solatus): Evidence for Host-Dependent Evolution of SIV within the C. LhoestiSuperspecies.” Journal of Virology 73(9):7734–44.
- ↑ 8.0 8.1 8.2 8.3 8.4 Keele, Brandon F., James Holland Jones, Karen A. Terio, Jacob D. Estes, Rebecca S. Rudicell, Michael L. Wilson, Yingying Li, Gerald H. Learn, T. Mark Beasley, Joann Schumacher-Stankey, Emily Wroblewski, Anna Mosser, Jane Raphael, Shadrack Kamenya, Elizabeth V Lonsdorf, Dominic A. Travis, Titus Mlengeya, Michael J. Kinsel, James G. Else, Guido Silvestri, Jane Goodall, Paul M. Sharp, George M. Shaw, Anne E. Pusey, and Beatrice H. Hahn. 2009. “Increased Mortality and AIDS-like Immunopathology in Wild Chimpanzees Infected with SIVcpz.” Nature 460(7254):515–19.
- ↑ 9.0 9.1 9.2 9.3 Sharp, P. M., E. Bailes, F. Gao, B. E. Beer, V. M. Hirsch, and B. H. Hahn. 2000. “Origins and Evolution of AIDS Viruses: Estimating the Time-Scale.” Biochemical Society Transactions 28(2):275–82.
- ↑ 10.0 10.1 10.2 Gardner, Murray B., and Paul A. Luciw. 1989. “Animal Models of AIDS.” The FASEB Journal 3(14):2593–2606.
- ↑ Barré-Sinoussi, Françoise. 1996. “HIV as the Cause of AIDS.” The Lancet 348(9019):31–35.
- ↑ 12.0 12.1 Letvin, N. L., K. A. Eaton, W. R. Aldrich, P. K. Sehgal, B. J. Blake, S. F. Schlossman, N. W. King, and R. D. Hunt. 1983. “Acquired Immunodeficiency Syndrome in a Colony of Macaque Monkeys.” Proceedings of the National Academy of Sciences of the United States of America 80(9):2718–22.
- ↑ http://www.freepik.com/free-vector/business-team-outlines-pack_831669.htm#term=human
- ↑ http://www.flaticon.com/free-icon/monkey_47138
- ↑ http://www.freepik.com/free-vector/cat-silhouettes-set_718091.htm#term=cat
- ↑ http://www.flaticon.com/free-icon/horse-standing-black-shape_35907
- ↑ http://www.freepik.com/free-vector/cows-and-bull-silhouettes_788343.htm
- ↑ http://www.freepik.com/free-vector/pack-of-farm-animal-silhouettes_1058750.htm#term=sheep&page=1&position=29
- ↑ 19.0 19.1 19.2 19.3 Shuljak, B. F. 2006. “Lentiviruses in Ungulates. I. General Features, History and Prevalence.” Bulgarian Journal of Veterinary Medicine 9(3):175–81.
- ↑ Pedersen, N. C., J. K. Yamamoto, T. Ishida, and H. Hansen. 1989. “Feline Immunodeficiency Virus Infection.” Veterinary Immunology and Immunopathology 21(1):111–29.
- ↑ 21.0 21.1 21.2 21.3 21.4 21.5 21.6 21.7 Keele, Brandon F., Fran Van Heuverswyn, Yingying Li, Elizabeth Bailes, Jun Takehisa, Mario L. Santiago, Frederic Bibollet-Ruche, Yalu Chen, Louise V Wain, Florian Liegeois, Severin Loul, Eitel Mpoudi Ngole, Yanga Bienvenue, Eric Delaporte, John F. Y. Brookfield, Paul M. Sharp, George M. Shaw, Martine Peeters, and Beatrice H. Hahn. 2006. “Chimpanzee Reservoirs of Pandemic and Nonpandemic HIV-1.” Science (New York, N.Y.) 313(5786):523–26.
- ↑ 22.0 22.1 22.2 22.3 22.4 Pedersen, N. C., J. K. Yamamoto, T. Ishida, and H. Hansen. 1989. “Feline Immunodeficiency Virus Infection.” Veterinary Immunology and Immunopathology 21(1):111–29.
- ↑ 23.0 23.1 Apetrei, Cristian, Amitinder Kaur, Nicholas W. Lerche, Michael Metzger, Ivona Pandrea, Johnny Hardcastle, Shelley Falkenstein, Rudolf Bohm, Jeffrey Koehler, Vicki Traina-Dorge, Tessa Williams, Silvija Staprans, Gail Plauche, Ronald S. Veazey, Harold McClure, Andrew A. Lackner, Bobby Gormus, David L. Robertson, and Preston A. Marx. 2005. “Molecular Epidemiology of Simian Immunodeficiency Virus SIVsm in U.S. Primate Centers Unravels the Origin of SIVmac and SIVstm.” Journal of Virology 79(14):8991.
- ↑ Desrosiers, Ronald C. 1990. “The Simian Immunodeficiency Viruses.” Annual Review of Immunology 8(1):557–78.
- ↑ https://viralzone.expasy.org/264?outline=all_by_species
- ↑ 26.0 26.1 Policicchio, Benjamin B., Ivona Pandrea, and Cristian Apetrei. 2016. “Animal Models for HIV Cure Research.” Frontiers in Immunology 7:12.
- ↑ Rudicell, Rebecca S., James Holland Jones, Emily E. Wroblewski, Gerald H. Learn, Yingying Li, Joel D. Robertson, Elizabeth Greengrass, Falk Grossmann, Shadrack Kamenya, and Lilian Pintea. 2010. “Impact of Simian Immunodeficiency Virus Infection on Chimpanzee Population Dynamics.” PLoS Pathogens 6(9).
- ↑ Pusey, Anne E., Lilian Pintea, Michael L. Wilson, Shadrack Kamenya, and Jane Goodall. 2007. “The Contribution of Long‐term Research at Gombe National Park to Chimpanzee Conservation.” Conservation Biology 21(3):623–34.
- ↑ 29.0 29.1 Worobey, Michael, Paul Telfer, Sandrine Souquière, Meredith Hunter, Clint A. Coleman, Michael J. Metzger, Patricia Reed, Maria Makuwa, Gail Hearn, Shaya Honarvar, Pierre Roques, Cristian Apetrei, Mirdad Kazanji, and Preston A. Marx. 2010. “Island Biogeography Reveals the Deep History of SIV.” Science 329(5998):1487.
- ↑ Wertheim, Joel O., and Michael Worobey. 2009. “Dating the Age of the SIV Lineages That Gave Rise to HIV-1 and HIV-2.” PLoS Computational Biology 5(5):e1000377–e1000377.
- ↑ Goodman, Morris, Danilo A. Tagle, David H. A. Fitch, Wendy Bailey, John Czelusniak, Ben F. Koop, Philip Benson, and Jerry L. Slightom. 1990. “Primate Evolution at the DNA Level and a Classification of Hominoids.” Journal of Molecular Evolution 30(3):260–66.
- ↑ Lemey, Philippe, Oliver G. Pybus, Bin Wang, Nitin K. Saksena, Marco Salemi, and Anne-Mieke Vandamme. 2003. “Tracing the Origin and History of the HIV-2 Epidemic.” Proceedings of the National Academy of Sciences of the United States of America 100(11):6588–92.
- ↑ Worobey, Michael, Marlea Gemmel, Dirk E. Teuwen, Tamara Haselkorn, Kevin Kunstman, Michael Bunce, Jean-Jacques Muyembe, Jean-Marie M. Kabongo, Raphaël M. Kalengayi, Eric Van Marck, M. Thomas P. Gilbert, and Steven M. Wolinsky. 2008. “Direct Evidence of Extensive Diversity of HIV-1 in Kinshasa by 1960.” Nature 455(7213):661–64.
- ↑ Chakrabarti, Lisa, Mireille Guyader, Marc Alizon, Muthiah D. Daniel, Ronald C. Desrosiers, Pierre Tiollais, and Pierre Sonigo. 1987. “Sequence of Simian Immunodeficiency Virus from Macaque and Its Relationship to Other Human and Simian Retroviruses.” Nature 328(6130):543–47.
- ↑ Van Heuverswyn, Fran, Yingying Li, Cecile Neel, Elizabeth Bailes, Brandon F. Keele, Weimin Liu, Severin Loul, Christelle Butel, Florian Liegeois, Yanga Bienvenue, Eitel Mpoudi Ngolle, Paul M. Sharp, George M. Shaw, Eric Delaporte, Beatrice H. Hahn, and Martine Peeters. 2006. “SIV Infection in Wild Gorillas.” Nature 444(7116):164.
- ↑ Campbell-Yesufu, Omobolaji T., and Rajesh T. Gandhi. 2011. “Update on Human Immunodeficiency Virus (HIV)-2 Infection.” Clinical Infectious Diseases : An Official Publication of the Infectious Diseases Society of America 52(6):780–87.
- ↑ Hirsch, Vanessa M., Robert A. Olmsted, Michael Murphey-Corb, Robert H. Purcell, and Philip R. Johnson. 1989. “An African Primate Lentivirus (SIVsmclosely Related to HIV-2.” Nature 339(6223):389–92.
- ↑ Khan, A. S., T. A. Galvin, L. J. Lowenstine, M. B. Jennings, M. B. Gardner, and C. E. Buckler. 1991. “A Highly Divergent Simian Immunodeficiency Virus (SIVstm) Recovered from Stored Stump-Tailed Macaque Tissues.” Journal of Virology 65(12):7061–65.
- ↑ Huet, Thierry, Rémi Cheynier, Andreas Meyerhans, Georges Roelants, and Simon Wain-Hobson. 1990. “Genetic Organization of a Chimpanzee Lentivirus Related to HIV-1.” Nature 345(6273):356–59.
- ↑ Simon, François, Philippe Mauclère, Pierre Roques, Ibtissam Loussert-Ajaka, Michaela C. Müller-Trutwin, Sentob Saragosti, Marie Claude Georges-Courbot, Françoise Barré-Sinoussi, and Françoise Brun-Vézinet. 1998. “Identification of a New Human Immunodeficiency Virus Type 1 Distinct from Group M and Group O.” Nature Medicine 4(9):1032–37.
- ↑ 41.0 41.1 Heuverswyn, Fran Van, Yingying Li, Elizabeth Bailes, Cecile Neel, Benedicte Lafay, Brandon F. Keele, Katharina S. Shaw, Jun Takehisa, Matthias H. Kraus, Severin Loul, Christelle Butel, Florian Liegeois, Bienvenue Yangda, Paul M. Sharp, Eitel Mpoudi-Ngole, Eric Delaporte, Beatrice H. Hahn, and Martine Peeters. 2007. “Genetic Diversity and Phylogeographic Clustering of SIVcpzPtt in Wild Chimpanzees in Cameroon.” Virology 368(1):155–71.
- ↑ Goodall, Jane. "The chimpanzees of Gombe: Patterns of behavior." Cambridge Mass (1986).
- ↑ 43.00 43.01 43.02 43.03 43.04 43.05 43.06 43.07 43.08 43.09 43.10 43.11 43.12 43.13 43.14 43.15 D’arc, Mirela, Ahidjo Ayouba, Amandine Esteban, Gerald H. Learn, Vanina Boué, Florian Liegeois, Lucie Etienne, Nikki Tagg, Fabian H. Leendertz, Christophe Boesch, Nadège F. Madinda, Martha M. Robbins, Maryke Gray, Amandine Cournil, Marcel Ooms, Michael Letko, Viviana A. Simon, Paul M. Sharp, Beatrice H. Hahn, Eric Delaporte, Eitel Mpoudi Ngole, and Martine Peeters. 2015. “Origin of the HIV-1 Group O Epidemic in Western Lowland Gorillas.” Proceedings of the National Academy of Sciences of the United States of America 112(11):E1343–52.
- ↑ Sheehy, Ann M., Nathan C. Gaddis, Jonathan D. Choi, and Michael H. Malim. 2002. “Isolation of a Human Gene That Inhibits HIV-1 Infection and Is Suppressed by the Viral Vif Protein.” Nature 418(6898):646–50.
- ↑ Stremlau, Matthew, Christopher M. Owens, Michel J. Perron, Michael Kiessling, Patrick Autissier, and Joseph Sodroski. 2004. “The Cytoplasmic Body Component TRIM5α Restricts HIV-1 Infection in Old World Monkeys.” Nature 427(6977):848–53.
- ↑ Neil, Stuart J. D., Trinity Zang, and Paul D. Bieniasz. 2008. “Tetherin Inhibits Retrovirus Release and Is Antagonized by HIV-1 Vpu.” Nature 451(7177):425–30.
- ↑ Kluge, Silvia F., Katharina Mack, Shilpa S. Iyer, François M. Pujol, Anke Heigele, Gerald H. Learn, Shariq M. Usmani, Daniel Sauter, Simone Joas, Dominik Hotter, Frederic Bibollet-Ruche, Lindsey J. Plenderleith, Martine Peeters, Matthias Geyer, Paul M. Sharp, Oliver T. Fackler, Beatrice H. Hahn, and Frank Kirchhoff. 2014. “Nef Proteins of Epidemic HIV-1 Group O Strains Antagonize Human Tetherin.” Cell Host & Microbe 16(5):639–50.
- ↑ 48.0 48.1 48.2 Sauter, Daniel, Michael Schindler, Anke Specht, Wilmina N. Landford, Jan Münch, Kyeong-Ae Kim, Jörg Votteler, Ulrich Schubert, Frederic Bibollet-Ruche, Brandon F. Keele, Jun Takehisa, Yudelca Ogando, Christina Ochsenbauer, John C. Kappes, Ahidjo Ayouba, Martine Peeters, Gerald H. Learn, George Shaw, Paul M. Sharp, Paul Bieniasz, Beatrice H. Hahn, Theodora Hatziioannou, and Frank Kirchhoff. 2009. “Tetherin-Driven Adaptation of Vpu and Nef Function and the Evolution of Pandemic and Nonpandemic HIV-1 Strains.” Cell Host & Microbe 6(5):409–21.
- ↑ Schmökel, Jan, Daniel Sauter, Michael Schindler, Fabian H. Leendertz, Elizabeth Bailes, Marie-Christine Dazza, Sentob Saragosti, Frederic Bibollet-Ruche, Martine Peeters, Beatrice H. Hahn, and Frank Kirchhoff. 2011. “The Presence of a Vpu Gene and the Lack of Nef-Mediated Downmodulation of T Cell Receptor-CD3 Are Not Always Linked in Primate Lentiviruses.” Journal of Virology 85(2):742–52.
- ↑ Delaugerre, Constance, Fabienne De Oliveira, Caroline Lascoux-Combe, Jean-Christophe Plantier, and François Simon. 2011. “HIV-1 Group N: Travelling beyond Cameroon.” The Lancet 378(9806):1894.
Authored by Minh Pham for BIOL 238 Microbiology, taught by Joan Slonczewski, 2020, Kenyon College.
