User:Rachelevelyn7
I am a third year science undergrad at the University of British Columbia. I am working towards a Bachelor's in Microbiology and Immunology.
Hello :)
Nod Factors
Introduction
Nodulation factors (Nod factors) act as signalling molecules between symbiotic bacteria and plants in response to flavonoids secreted by legume root hairs [1]. Rhizobium refers to bacteria that include: Rhizobia, Bradyrhizobia, Azorhizobia, and Sinorhizobia [2]. Members of these genera are capable of forming endosymbiotic interactions with leguminous plants [3]. These bacteria have the ability to fix nitrogen into a form that plants can utilize. The enzyme that is used for this process is nitrogenase and it has the ability to reduce atmospheric nitrogen (N2) to ammonia (NH3) at normal temperatures and atmospheric pressure [4]. Nod factors are secreted by rhizobia during a tightly regulated signalling pathway which results in formation of root nodules in leguminous plants, resulting in the rhizobia becoming bacteroids [2].
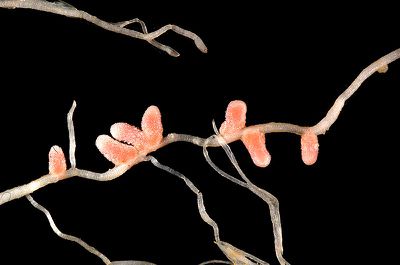
Rhizobium-Legume Symbiosis
The symbiotic interaction between rhizobia and leguminous plants plays a critical function in fixing inorganic nitrogen into biologically usable forms and, thus, is critical to agriculture and many ecosystems [6]. The estimated amount of nitrogen fixed by rhizobia-legume interactions is 15 Tg, which is equal to the amount that humans fix for the production of fertilizers [6]. The synthetic production of fertilizers follows the Haber-Bosch Process:
N2 + 3 H2 ⇌ 2 NH3
The Haber-Bosch Process requires a temperature of 450°C and a pressure of 500 atm. Both these requirements
are met by the burning fossil fuels and contribute to climate change [7].
In contrast, nitrogenase, the enzyme that rhizobia use to fix nitrogen, [5] uses the process:
N2 + 8 H+ + 16 ATP ⇌ 2 NH3 + 16 ADP + 16Pi + H2
The enzyme has the ability to work at normal temperature and atmospheric pressure [7].
Rhizobia supply reduced nitrogen to the leguminous plant while the plant provides rhizobia with sugars in the form of photosynthates [7]. Since both organisms benefit from this interaction, they share a mutualistic symbiosis.
Nodulation Process
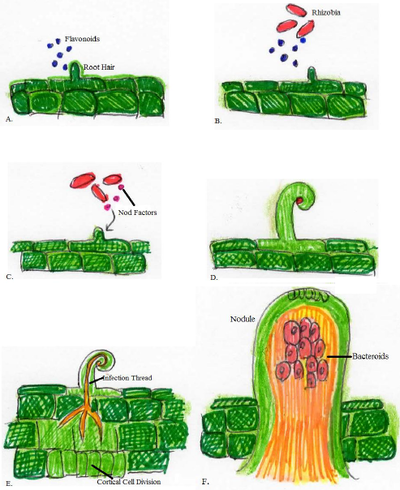
1) The leguminous plant releases flavonoids into the area surrounding the plant roots, known as the rhizosphere, when the available nitrogen in the soil is depleted [8].
2) Rhizobia in the surrounding soil secrete NodD, which is a protein that recognizes the flavonoids secreted by the plants [8].
3) Interaction with the flavonoids activates NodD, then NodD returns to the rhizobium to induce the transcription of nodABC genes [8].
4) nodABC proteins modify the Nod factor in response to the flavonoids released by the plant [8].
5) The Nod factors interact with the plant and cause root hair curling and root hair deformation. This leads to the formation of an infection thread into the root hair [8].
6) Nod factors cause cortical cell division in the primordium of the plant and form a nodule [8].
7) The rhizobia move through the plant root hair via the infection thread to the nodule within the plant [8].
8) Rhizobia switch into bacteroid form and are surrounded by a plant-derived symbiosome membrane [8].
9) Rhizobia begin nitrogen fixation and provide the legume with amino acids, such as glutamine and asparagine, or ureides, such as allantoin and allantoic acid, which are made in the nodule [9].
10) The legume provides simple sugars produced from plant catabolism to the rhizobia in the nodule [9].
Function of Nod Factors
Nod factors are the important signaling molecules in the symbiotic interaction between rhizobia and leguminous plants. When they bind to the root hairs of the leguminous plants, they cause root hair deformations, activation of plant genes, initiation of cortical cell division and nodule formation [9]. Each of these processes are necessary for the proper formation of root nodules, which enable the symbiotic interaction between rhizobia and leguminous plants. Nod factors control the specificity of the interaction between the two organisms and they specifically cause morphological and physiological changes in leguminous plant [1].
Additionally, Nod factors can affect other plants in the soil surrounding them. When Nod factors are present in mixed crop fields, they have the ability to stimulate seed germination, promote plant growth, increase photosynthetic rates, and increase grain yields of legume and non-legume crops [2]. Thus, Nod factors do not only benefit leguminous plants, but can contribute to the growth rate of many different types of crops.
Structure of Nod Factors
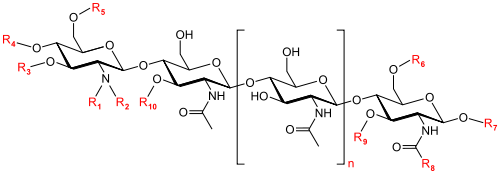
Nod factors are lipochito-oligosaccharides and have three to five N-acetyl-glucosamines [9]. The substitutions on the lipochito-oligosaccharides side chains determine specific recognition by the Nod factor receptors found in leguminous root hairs [10].
The specific structure of Nod factors is determined by modifications made by Nod genes, which are found in the rhizobium genome [9]. The Nod genes encode for proteins that modify Nod factors by adding or removing different chemical structures such as sulfates, fatty acids, acetyl groups, and methyl groups to the original lipochito-oligosaccharide structure [9].
The most extensively studied Nod genes are the common Nod genes (nodA, nodB and nodC). The enzymes encoded by the common Nod genes have specific functions. NodC is an enzyme that synthesizes the N-acetyl glucosamine backbone of the Nod factor [12]. NodB is a deacetylase that removes an acetyl group from the Nod factor and NodA is an acyltransferase that adds a fatty acid chain to the site deacetylated by NodB [12].
Host Specificity due to Nod Factors
There are two modes of specificity for rhizobium-legume symbiosis. The first is the interaction between the flavonoids, the chemical signals that are derived from the legumes [8]. The recognition of the flavonoids by the bacteria is dependent on the interactions with NodD [8]. When NodD is activated, it leads to the expression of nodABC as well as many other Nod genes, which results in the structural modification of Nod factors [13]. The second mode of specificity is dependent on the structure of the Nod factors themselves [14]. Since there are some bacteria that can make several structurally different Nod factors and therefore have different host plants, slight variations in the Nod factor structure leads to host specificity [11].
Genetic History
Several components of the rhizobium-legume interactions are similar to arbuscular mycorrhizal (AM) symbiosis. In both of these symbioses, specialized host membranes surround the microbes and form a symbiotic surface that facilitates the exchange of nutrients [1]. Another similarity is that AM fungi enter the root of their host plant in a comparable mechanism to rhizobia [16]. Also, the hyphae of the AM fungi spread within the cells of the inner cortex of the plant, this is mimicked by the rhizobium infection threads when forming nodules within the inner cortex of legumes [15]. Significantly, AM fungi produce lipochito-oligosaccharides, which are structurally similar to the Nod factors synthesized by rhizobia [15]. Thus, it has been hypothesized that these similar cellular processes are evidence that rhizobium-legume interactions was derived from the AM symbiosis with plant roots and the similar interactions between these two symbioses have been compiled into a common signalling pathway known as the common symbiosis (SYM) signalling pathway [16].
References
(1) Catoira, R., Galera, C. de Billy, F., Penmetsa, R. V., Journet, E.-P., Maillet, F., Rosenberg, C., Cook, D., Gough, C., and Denarie, J. “Four Genes of Medicago truncatula Controlling Components of a Nod Factor Transduction Pathway.” 2000. The Plant Cell, Volume 12. p. 1647-1665.
(2) Dakora, F. D. “Defining New Roles for Plant and Rhizobial Molecules in Sole and Mixed Plant Cultures Involving Symbiotic Legumes.” 2003. New Phytologist, Volume 158. p. 39-49.
(3) Guerts, R., Lillo, A., and Bisseling, T. “Exploiting an ancient signalling machinery to enjoy a nitrogen fixing symbiosis.” 2012. Current Opinion in Plant Biology. Volume 15. p. 438-443.
(4) Postgate, J. (1998). Nitrogen Fixation, 3rd Edition. Cambridge University Press. Cambridge UK.
(5) Zumft, W. G. and Mortenson, L. E. “The nitrogen-fixing complex of bacteria.” 1975. Biochem. Biophys. Acta. Volume 416. p. 1-52.
(6) Akcay, E. and Roughgarden, J. “Negotiation of Mutualism: Rhizobia and Legumes.” 2007. Proceedings: Biological Sciences, Volume 274. p. 25-32.
(7) Depret, G. and Laguerre, G. “Plant Phenology and Genetic Variability in Root and Nodule Development Strongly Influence Genetic Structure of Rhizobium Leguminosarum Biovar Viciae Populations Nodulating Pea.” 2008. New Phytologist, Volume 179. p. 224-235.
(8) Lindstrom, K., Terefework, Z., Suominen, L., and Lortet, G. “Signalling and Development of Rhizobium: Legume Symbioses.” 2002. Biology and Environment: Proceedings of the Royal Irish Academy, Volume 102B. p. 61-64.
(9) Lodwig, E. M., Hosie, A. H., Bourdes, A., Findlay, K., Allaway, D., Karunakaran, R., Downie, J. A., and Poole, P. S. “Amino-acid cycling drives nitrogen fixation in the legume-Rhizobium symbiosis.” 2003. Nature, Volume 422. p. 722-726.
(10) Streng, A., Camp, R., Bisseling, T., and Guerts, R. “Evolutionary origin of rhizobium Nod factor signaling.” 2011. Plant Signaling and Behaviour, Volume 6. p. 1510-1514
(11) Schultze, M., Quiclet-Sire, B., Kondorosi, E., Virelizier, H., Glushak, J. N., Endre, G., Gero, S. D., and Kondorosi, A. “Rhizobium meliloti Produces a Family of Sulfated Lipo-Oligosaccharides Exhibiting Different Degrees of Plant Host Specificity. 1992. Proceedings of the National Academy of Sciences of the United States of America, Volume 89. p. 192-196.
(12) Perret, X., Staehelin, C., and Broughton, W. J., “Molecular Basis of Symbiotic Promiscuity.” 2000. Microbiology and Molecular Biology Reviews, Volume 64. p. 180-201.
(13) Ardourel, M., Demont, N., Debelle, F., Maillet, F., de Billy, F., Prome, J-C., Denarie, J., and Truchet, G. “Rhizobium meliloti Lipooligosaccharide Nodulation Factors: Different Structural Requirements for Bacterial Entry into Target Root Hair Cells and Induction of Plant Symbiotic Developmental Responses.” 1994. The Plant Cell, Volume 6. p. 1357-1374.
(14) Vijn, I., das Neves, L., van Kammen, A., Franssen, H., and Bisseling, T. “Nod Factors and Nodulation in Plants.” 1993. Science, Volume 260. p. 1764-1765.
(15) Bonfante, P. and Requena, N. “Dating in the dark: how roots respond to fungal signals to establish arbuscular mycorrhizal symbiosis.” 2011. Current Opinion in Plant Biology, Volume 14. p. 451-457.
(16) Ivanov, S., Fedorova, E. E., Limpens, E., De Mita, S., Genre, A., Bonfante, P., and Bisseling, T. “Rhizobium-legume symbiosis shares an exocytotic pathway required for arbuscule formation. 2012. Proceedings of the National Academy of Sciences of the United States of America, Volume 109. p. 8316-8321.
