Xanthomonas citri
Classification
Higher order taxa
Domain: Bacteria
Phylum: Proteobacteria
Class: Gammaproteobacteria
Order: Xanthomonadales
Family: Xanthomonadanceae
Species
Xanthomonas citri
Description and significance
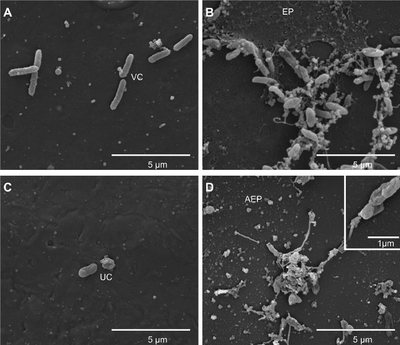
X. citri is part of the Xanthomonas genus of phytopathogenic bacteria that inhabit the phyllosphere of citrus plants species and crops and produce the citrus canker disease. The bacteria are named for their pathogenic behaviors that target citrus fruits and crops. At least three forms of the disease exists, with one of them, Canker A, being the most destructive form.
Although the geographical origin and the original discoverer of X. citri is debatable; Lee (1918) stated that the species originated from southern China, however, Fawcett and Jenkins (1933) reported that the species originated from parts of India and Java. Fawcett and Jenkins’ findings suggest that X. citri originated from tropical areas in Asia. X. citri was first described in 1915, after it was found in the Gulf States of USA (1). Severe infectious forms of the disease can cause premature fruit drop, defoliation, blemished fruit, and reduced fruit quality. In addition, warm, humid, or cloudy climate combined with heavy rainfall and strong wind exacerbates the impacts of the disease.
X. citri can also produce biofilms on both biotic and abiotic surfaces of organisms, particularly on cankers of diseased plants. The bacteria utilize flagella-mediated motility throughout biofilm formation, and sliding and swimming motility (without flagella) through diffusible signal factor (DSF) (2). In addition, X. citri utilizes type IV pilus for twitching motility, biofilm development, and adherence (3). X. citri secretes proteins through the type III secretory system into the extracellular matrix (4). Bacterial colonies of X. citri are circular, convex, mucoid, and shiny (21). The bacteria are gram-negative, rod-shaped, aerobic (5), and nonsporeforming. They also produce yellow, water-insoluble pigments due to the presence of carotenoid nostoxanthin, and oxidatively metabolize sugars (6).
Nutrition and growth
X. citri requires methionine or cysteine for growth, and is inhibited by 0.02% triphenyltetrazolium chloride. X. citri growth develops in 46-72 hours and colonies develop as creamy-yellow colors on PDA or straw-yellow on nutrient agar. The identity of isolated bacteria may be confirmed through its pathogenicity to grapefruit leaves or by utilizing polymerase chain reaction (PCR) techniques using specific primers (8).
L-glutamate is an important nitrogen source and nutrition in X. citri and has a significant role in xanthan gum production (exopolysaccharide used in industrial and biotechnological applications) by significantly increasing its production (9).
Some natural field strains of X. citri have been reported to have the capability of using lactose as a sole carbon source (in xanthan gum production) (10).
For optimal laboratory study of X. citri, the bacteria may be stored in 4-8°C until processing (11) and requires incubation at 28°C in NB liquid medium for 16-20 hours with a pH of approximately 5.4 (12).
X. citri waste by-product contain approximately 15.2% cellulose, 18.2% hemicellulose, 24.6% pectin, 15-20% soluble sugars, 6.5-8.0% proteins, 3.8% starch, and 3.5% ashes (13).
Genome and genetics
X. citri belongs to the major prokaryotic branch of bacteria (6). There are a few distinct bacteria closely related to X. citri, such as Xanthomonas vesicatoria (14) and Xanthomonas campestris pv. malvacearum by DNA-DNA hybridization (15).
The complete circular genome sequencing of X. citri strain AW12879 was determined by using 454 pyrosequencing and Illumina technology. The genome contains a chromosome with DNA GC content of 64.71% and genome size of 5,320,000 base pairs, and also contains two plasmids (circular), pXcaw19 with DNA GC content of 63.07% and 18,869 base pairs and pXcaw58 with DNA GC content of 61.85% and 58,317 base pairs. The plasmid pXcaw19 is not similar to plasmids ‘’X. Citri subsp. citri’’ strain 306. On the other hand, the plasmid strain pXcaw58 has about 35% similarity with pXAC64 (16).
X. citri strain AW12879 has 4,760 annotated CDS, of which 3,457 can be assigned to a COG functional category. The genome has two rRNA operons (identified with FgenesB) and 54 tRNA genes (identified with tRNAscan-SE (16).


Pathology
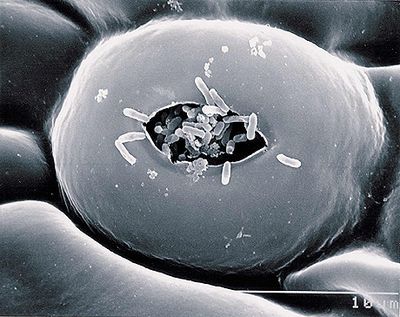
X. citri is best known as causing the disease citrus canker in fruits. There are various mechanisms for the bacterial propagation of this disease. X. citri grow out of lesions in leaves, stems, and fruit, and if free moisture exists on the lesions, the bacteria oozes out and spreads its infection across the surface of the lesion. The bacteria may also penetrate out through stomatal pores or wounds caused by thorns, insects, and blowing sand, which are aided by wind-driven rain (most common method of penetration) (17).
X. citri remain alive in the margins of lesions, until the leaves or fruit fall onto the ground. The bacteria also die upon exposure to dry surface after oozing through lesions onto surfaces; accelerated death also occurs as a result of sun exposure (17).
Sanitation, windbreaks, and leafminer control with frequent copper spray application have been considered as being plausible and important methods of treatment for plants and trees infected by X. citri. Suggestions for improving methods of infection control include the ideas that applying several weeks of copper spray application may decrease the infection, using windbreaks may greatly reduce the speed of bacterial infection spread and severity, and that using windbreaks may enhance the copper spray’s impact. In addition, leafminer control may decrease infection in young trees and certain cultivars (17).
Current Research
Foliar Application of Biofilm
In one recent study, scientists framed their research based on the idea that biofilm formation is important during early infection of X. citri on host leaves. They focused on testing their hypothesis that small molecules that inhibit biofilm formation of X. citri would enhance control of the citrus disease. Through their investigation and study, they found that D-leucine (amino acid involved in energy and metabolism) and 3-indolyl acetonitrile (known for inhibiting biofilm substance on Escherichia coli and Pseudomonas aeruginosa) inhibited biofilm formation of X. citri on different abiotic surfaces and host leaves. The scientists observed that the biofilm inhibition occurred at a concentration lower than their set minimum inhibitory concentration. In addition, they found that when D-leucine and IAN are applied individually or combined with copper, in greenhouse assays, such combination reduced the number of canker lesions and bacterial populations of X. citri on citrus host leaves (18).
Biofilm formation and Type III Protein Secretion
Many bacterial plant pathogens utilize a Type III protein secretion system (T3SS) to secrete effector proteins to colonize their hosts. Research was conducted to investigate the role of T3SS in X. citri subsp. citri in biofilm formation. The scientists who conducted this study compared the biofilm formation capacity of various X. citri T3SS mutants with a wild type strain. The scientists discovered that the T3SS pathway was necessary for biofilm formation in X. citri. Lack of T3SS in T3SS mutants caused changes in protein expression related to metabolic processes, energy generation, exopolysaccharide production, bacterial motility, and outer membrane proteins. Alterations of EPS production and bacterial motility were also present. T3SS was shown to increase X. citri virulence, and thus indicated T3SS's novel role in biofilm formation in the bacteria (19).
References
1. Das A.K. "Citrus canker—A review". J Appl Hort. June 2003. 5(1): 52-60.
7. http://www.apsnet.org/publications/apsnetfeatures/Pages/citruscanker.aspx
8. http://www.cabi.org/isc/datasheet/56921
17. Gottwald and Graham. "Citrus canker". "The Plant Health Instructor". 2005.
21. Xanthomonas axonopodis pv. citri. EPPO Bulletin. Aug 2005. (35): 289-294.
Authored by GJ, a student of CJ Funk at John Brown University
