Bacteroide composition in the gut: Difference between revisions
No edit summary |
|||
| Line 4: | Line 4: | ||
==Genomic and Proteomic Advances== | ==Genomic and Proteomic Advances== | ||
Recently, genomic and proteomic analyses have provided insight into the mechanisms in which microbiota adapt and thrive within varying microenvironments. Sequencing B. thetaiotaomicron and B. fragilis show notable genomic commonalities. These Bacteroides, like eukaryotes, show low gene content for the size of their genome (Xu and Gordon, 2003). A large portion of the their genetic make-up consists of proteins containing over 1,000 amino acids, which, it seems, allow for a vast selection of relevant proteins for gene expression. Putative relevant proteins have been determined by homology to other known proteins. Of the 4779 proteins identified in the B. thetaiotaomicron proteome, 58% were ascribed a supposed function based on other known proteins, 18% were homologous to proteins without a known function, and 24% were not homologous to any protein identified in the public domain (Xu et al., 2003). The variety of functions is presumed to aid Bacteroides species in multiple ways: the proteome induces the mechanism by which some Bacteroides acquire and hydrolyze otherwise indigestible dietary polysaccharides, and includes an environment-sensing apparatus with extracytoplasmic function sigma factors and one- and two-component signal transduction systems (Xu et. al., 2003). | |||
<br><i>B. thetaiotaomicron</i> contains the capacity to utilize glycans derived from the colon. The majority (61%) of the necessary enzymes (glycohyrolases) potentially exist in the periplasm, and are necessary to the polysaccharide fermentation process that increases energy availability to the host. Other extracellular proteins have been shown to facilitate the binding of starches to the cell surface to be digested by other extracellular enzymes (Xu et al., 2003). | |||
Additionally, B. thetaiotaomicron was shown to contain a significant population of ECF-sigma factors, which according to other identified homologous genes, are cotranscribed with regulatory proteins that respond to external environmental stimulus and act on gene expression accordingly. | |||
The majority of one-component signal transduction systems were shown to be similar to other genes responsible for nutrient utilization, providing further evidence of Bacteroides’ genetically evolved fitness advantage in nutrient acquisition. | |||
Through genomic and proteomic sequencing, some of the mechanism underlying the adaptive capabilities of the Bacteroide species in the human gut have been identified. The expansive array of proteins available to them mediates their ability to respond to their environment, thrive within the rapidly changing microenvironment of the colon, and acquire and ferment complex carbohydrates for their own energy source as well as for the host. | |||
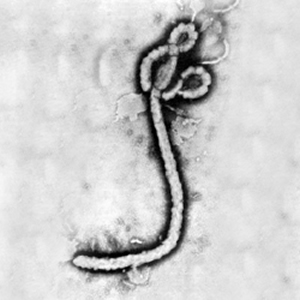
[[Image:Ebola virus 1.jpeg|thumb|300px|right|Electron micrograph of the Ebola Zaire virus. This was the first photo ever taken of the virus, on 10/13/1976. By Dr. F.A. Murphy, now at U.C. Davis, then at the CDC.]] | [[Image:Ebola virus 1.jpeg|thumb|300px|right|Electron micrograph of the Ebola Zaire virus. This was the first photo ever taken of the virus, on 10/13/1976. By Dr. F.A. Murphy, now at U.C. Davis, then at the CDC.]] | ||
Revision as of 00:27, 21 March 2015
This is the leader section -- here you can write a short description of your topic and why it is interesting. The goal is for this to quickly cover the main ideas of your topic and get the reader interested in reading the rest!
Genomic and Proteomic Advances
Recently, genomic and proteomic analyses have provided insight into the mechanisms in which microbiota adapt and thrive within varying microenvironments. Sequencing B. thetaiotaomicron and B. fragilis show notable genomic commonalities. These Bacteroides, like eukaryotes, show low gene content for the size of their genome (Xu and Gordon, 2003). A large portion of the their genetic make-up consists of proteins containing over 1,000 amino acids, which, it seems, allow for a vast selection of relevant proteins for gene expression. Putative relevant proteins have been determined by homology to other known proteins. Of the 4779 proteins identified in the B. thetaiotaomicron proteome, 58% were ascribed a supposed function based on other known proteins, 18% were homologous to proteins without a known function, and 24% were not homologous to any protein identified in the public domain (Xu et al., 2003). The variety of functions is presumed to aid Bacteroides species in multiple ways: the proteome induces the mechanism by which some Bacteroides acquire and hydrolyze otherwise indigestible dietary polysaccharides, and includes an environment-sensing apparatus with extracytoplasmic function sigma factors and one- and two-component signal transduction systems (Xu et. al., 2003).
B. thetaiotaomicron contains the capacity to utilize glycans derived from the colon. The majority (61%) of the necessary enzymes (glycohyrolases) potentially exist in the periplasm, and are necessary to the polysaccharide fermentation process that increases energy availability to the host. Other extracellular proteins have been shown to facilitate the binding of starches to the cell surface to be digested by other extracellular enzymes (Xu et al., 2003).
Additionally, B. thetaiotaomicron was shown to contain a significant population of ECF-sigma factors, which according to other identified homologous genes, are cotranscribed with regulatory proteins that respond to external environmental stimulus and act on gene expression accordingly.
The majority of one-component signal transduction systems were shown to be similar to other genes responsible for nutrient utilization, providing further evidence of Bacteroides’ genetically evolved fitness advantage in nutrient acquisition.
Through genomic and proteomic sequencing, some of the mechanism underlying the adaptive capabilities of the Bacteroide species in the human gut have been identified. The expansive array of proteins available to them mediates their ability to respond to their environment, thrive within the rapidly changing microenvironment of the colon, and acquire and ferment complex carbohydrates for their own energy source as well as for the host.
At right is a sample image insertion. It works for any image uploaded anywhere to MicrobeWiki. The insertion code consists of:
Double brackets: [[
Filename: Ebola virus 1.jpeg
Thumbnail status: |thumb|
Pixel size: |300px|
Placement on page: |right|
Legend/credit: Electron micrograph of the Ebola Zaire virus. This was the first photo ever taken of the virus, on 10/13/1976. By Dr. F.A. Murphy, now at U.C. Davis, then at the CDC.
Closed double brackets: ]]
Other examples:
Bold
Italic
Subscript: H2O
Superscript: Fe3+
Overall paper length should be 3,000 words, with at least 3 figures with data.
Section 2
Include some current research in each topic, with at least one figure showing data.
Section 3
Include some current research in each topic, with at least one figure showing data.
Further Reading
[Sample link] Ebola Hemorrhagic Fever—Centers for Disease Control and Prevention, Special Pathogens Branch
References
Edited by Maggie Schein, a student of Nora Sullivan in BIOL168L S2 (Microbiology) in The Keck Science Department of the Claremont Colleges Spring 2014.