Pseudomonas: Difference between revisions
No edit summary |
|||
| Line 13: | Line 13: | ||
===Species:=== | ===Species:=== | ||
''Pseudomonas abietaniphila; P. agarici; P. agarolyticus; P. alcaliphila; P. alginovora; P. andersonii; P. antarctica; P. asplenii; P. azelaica; P. batumici; P. borealis; P. brassicacearum; P. chloritidismutans; P. cremoricolorata; P. diterpeniphila; P. filiscindens; P. frederiksbergensis; P. gingeri; P. graminis; P. grimontii; P. halodenitrificans; P. halophila; P. hibiscicola; P. hydrogenovora; P. indica; P. japonica; P. jessenii; P. kilonensis; P. koreensis; P. lini; P. lurida; P. lutea; P. marginata; P. meridiana; P. mesoacidophila; P. pachastrellae; P. palleroniana; P. parafulva; P. pavonanceae; P. proteolyica; P. psychrophila; P. psychrotolerans; P. pudica; P. rathonis; P. reactans; P. rhizosphaerae; P. salmononii; P.thermaerum; P. thermocarboxydovorans; P. thermotolerans; P. thivervalensis; P. umsongensis; P. vancouverensis; P. wisconsinensis; P. xanthomarina; P. xiamenensis; P. | ''Pseudomonas abietaniphila; P. agarici; P. agarolyticus; P. alcaliphila; P. alginovora; P. andersonii; P. antarctica; P. asplenii; P. azelaica; P. batumici; P. borealis; P. brassicacearum; P. chloritidismutans; P. cremoricolorata; P. diterpeniphila; P. filiscindens; P. frederiksbergensis; P. gingeri; P. graminis; P. grimontii; P. halodenitrificans; P. halophila; P. hibiscicola; P. hydrogenovora; P. indica; P. japonica; P. jessenii; P. kilonensis; P. koreensis; P. lini; P. lurida; P. lutea; P. marginata; P. meridiana; P. mesoacidophila; P. pachastrellae; P. palleroniana; P. parafulva; P. pavonanceae; P. proteolyica; P. psychrophila; P. psychrotolerans; P. pudica; P. rathonis; P. reactans; P. rhizosphaerae; P. salmononii; P.thermaerum; P. thermocarboxydovorans; P. thermotolerans; P. thivervalensis; P. umsongensis; P. vancouverensis; P. wisconsinensis; P. xanthomarina; P. xiamenensis; P. sp''. <br /> | ||
'''P. aeruginosa group:''' ''P. aeruginosa; P. alcaligenes; P. anguilliseptica; P. citronellolis; P. flavescens; P. jinjuensis; P. mendocina; P. nitroreducens; P. oleovorans; P. pseudoalcaligenes; P. resinovorans; P. straminae <br /> | |||
'''P. chloroaphis group:''' ''P. aurantiaca; P. chlororaphis; P.s fragi; P. lundensis; P. taetrolens <br /> | |||
'''P. fluorescens group:''' ''P. azotoformans; P. brenneri; P. cedrina; P. congelans; P. corrugata; P. costantinii; P. extremorientalis; P. fluorescens; P. fulgida; P. gessardii; P. libanensis; P. mandelii; P. marginalis; P. mediterranea; P. migulae; P. mucidolens; P. orientalis; P. poae; P. rhodesiae; P. synxantha; P. tolaasii; P. trivialis; P. veronii <br /> | |||
'''P. pertucinogena group:''' ''P. denitrificans; P. pertucinogena <br /> | |||
'''P. putida group:''' ''P. fulva; P. monteilii; P. mosselii; P. oryzihabitans; P. plecoglossicida; P. putida <br /> | |||
'''P. stutzeri group:''' ''P. balearica; P. luteola; P. stutzeri <br /> | |||
'''P. syringae group:''' ''P. avellanae; P. cannabina; P. caricapapyae; P. cichorii; P. coronafaciens; P. fuscovaginae; P. tremae; P. viridiflava'' | |||
==Description and Significance== | ==Description and Significance== | ||
Revision as of 18:01, 8 June 2006
|
NCBI: |
Classification
Higher order taxa:
Bacteria; Proteobacteria; Gammaproteobacteria; Pseudomonadales; Pseudomonadaceae
Species:
Pseudomonas abietaniphila; P. agarici; P. agarolyticus; P. alcaliphila; P. alginovora; P. andersonii; P. antarctica; P. asplenii; P. azelaica; P. batumici; P. borealis; P. brassicacearum; P. chloritidismutans; P. cremoricolorata; P. diterpeniphila; P. filiscindens; P. frederiksbergensis; P. gingeri; P. graminis; P. grimontii; P. halodenitrificans; P. halophila; P. hibiscicola; P. hydrogenovora; P. indica; P. japonica; P. jessenii; P. kilonensis; P. koreensis; P. lini; P. lurida; P. lutea; P. marginata; P. meridiana; P. mesoacidophila; P. pachastrellae; P. palleroniana; P. parafulva; P. pavonanceae; P. proteolyica; P. psychrophila; P. psychrotolerans; P. pudica; P. rathonis; P. reactans; P. rhizosphaerae; P. salmononii; P.thermaerum; P. thermocarboxydovorans; P. thermotolerans; P. thivervalensis; P. umsongensis; P. vancouverensis; P. wisconsinensis; P. xanthomarina; P. xiamenensis; P. sp.
P. aeruginosa group: P. aeruginosa; P. alcaligenes; P. anguilliseptica; P. citronellolis; P. flavescens; P. jinjuensis; P. mendocina; P. nitroreducens; P. oleovorans; P. pseudoalcaligenes; P. resinovorans; P. straminae
P. chloroaphis group: P. aurantiaca; P. chlororaphis; P.s fragi; P. lundensis; P. taetrolens
P. fluorescens group: P. azotoformans; P. brenneri; P. cedrina; P. congelans; P. corrugata; P. costantinii; P. extremorientalis; P. fluorescens; P. fulgida; P. gessardii; P. libanensis; P. mandelii; P. marginalis; P. mediterranea; P. migulae; P. mucidolens; P. orientalis; P. poae; P. rhodesiae; P. synxantha; P. tolaasii; P. trivialis; P. veronii
P. pertucinogena group: P. denitrificans; P. pertucinogena
P. putida group: P. fulva; P. monteilii; P. mosselii; P. oryzihabitans; P. plecoglossicida; P. putida
P. stutzeri group: P. balearica; P. luteola; P. stutzeri
P. syringae group: P. avellanae; P. cannabina; P. caricapapyae; P. cichorii; P. coronafaciens; P. fuscovaginae; P. tremae; P. viridiflava
Description and Significance
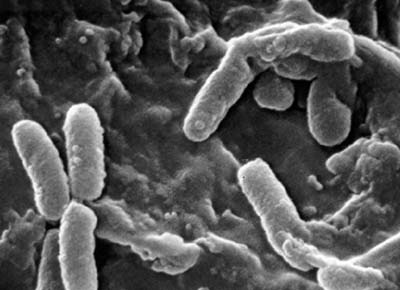
Pseudomonas bacteria can be found in many different environments such as soil, water, and plant and animal tissue. Many different species of this bacteria are opportunistic pathogens that affect humans, animals, and plants. Pseudomonas aeruginosa, called the "epitome" of opportunistic pathogens, almost never infects uncompromised tissues; however, it can infect practically any type of tissue if that tissue has some type of compromised defenses (Todar).
Genome Structure
Pseudomonas syringae pathovar tomato DC3000, the tomato and Arabidopsis thaliana pathogen, has genome 6.5 megabases in size that is compromised of a circular chromosome and two plasmids, which all encode for 5,763 ORFs. 298 established and putative virulence genes were found including those that encode for 31 comfirmed and 19 predicted proteins dealing with secretion systems. Many of these virulence genes are proximal to mobile elements; these collectively make up 7% of the whole genome. The genome also had many genes involved with the movement of nutrients, especially sugars, as well as genes dealing with cell attachment to plant surfaces. Over 12% of the the genes were found to deal with regulation; this may be due to the bacterium's need "for rapid adaptation to the diverse environments encountered during epiphytic growth and pathogenesis" (Buell et al. 2003). While many similarities were found among the genomes of Pseudomonas syringae pathovar tomato DC3000, Pseudomonas putida, and Pseudomonas aeruginosa, 1,159 genes were found to be unique to DC3000, 811 of which have no known function. (Buell et al. 2003)
Cell Structure and Metabolism
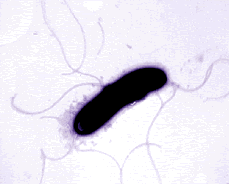
Pseudomonas bacteria are generally aerobic rod-shaped bacteria that are known for their metabolic diversity (DOE Joint Genome Institute). Some species of this bacteria, such as P. aeruginosa, are opportunistic pathogens that secrete extracellular proteases and adhere and invade host tissue. More than half of the clinical isolates of Pseudomonas bacteria produce pyocyanin, a blue-green pigment. The Pseudomonas fluorescens group are nonpathogenic saprophytes that also produce a pigment, particularly under conditions of low iron availability. This pigment is a soluble, greenish, fluorescent pigment that led to the group's name. These bacteria are generally obligate aerobes; however, some strains can utilize NO3 instead of O2 as an electron acceptor. They have multiple polar flagella that assist in the bacteria's movement. Because they have simple nutritional requirements, they "grow well in mineral salts media supplemented with any of a large number of carbon sources" (DOE). Some researches seek to exploit P. fluorescens to partially or completely degrade pollutants such as styrene, TNT, and polycyclic aromatic hydrocarbons. Several strains of this bacteria also have the ability to suppress plant diseases by "protecting the seeds and roots from fungal infection" (DOE). This ability is due to secondary metabolites produced by these bacteria such as antibiotics, siderophores, and hydrogen cyanide as well as the ability of these bacteria to rapidly colonize the rhizosphere and out-compete some of pathogens (DOE).
Ecology
Pseudomonas bacteria can be found in soil, marshes, coastal marine habitats, and plant and animal tissue; generally, these bacteria can tolerate a variety of physical conditions. Some of these species, such as Pseudomonas aeruginosa, are opportunistic pathogens. Cystic fibrosis patients, burn victims, people with cancer, and patients who are in intensive care units for extended periods of time are most at risk for contracting a disease caused by a P. aeruginosa infection (Pseudomonas Gemone Project).
Pathology
Pseudomonas aeruginosa is common in patients with compromised host defense mechanisms and is the "most common pathogen isolated from patients who have been hospitalized longer than one week" (Qarah and Cunha). It is common for this bacteria to cause nosocomial infections like pneumonia, urinary tract infections, and bacteremia; infections caused by this bacteria can become complicated and may even be life threatening. It is both invasive and toxigenic and has three stages of infection: the first is bacterial attachment and colonization, the second is local infection, and the third is bloodstream dissemination and systemic disease. The bacteria produce extracellular proteases that assist adherence and invasion and are important in the organism's virulence.
References
General:
- Pseudomonas Genome Project
- Todar, Kenneth: "Pseudomonas aeruginosa." University of Wisconsin-Madison Department of Bacteriology.
- DOE Joint Genome Institute: Pseudomonas fluorescens
Genome:
- Buell, Robin C., Vinita Joardar, Magdalen Lindeberg, Jeremy Selengut, Ian T. Paulsen, Michelle L. Gwinn, Robert J. Dodson, Robert T. Deboy, A. Scott Durkin, James R. Kolonay, Ramana Madupu, Sean Daugherty, Lauren Brinkac, Maureen J. Beanan, Daniel H. Haft, William C. Nelson, Tanja Davidsen, Nikhat Zafar, Liwei Zhou, Jia Liu, Qiaoping Yuan, Hoda Khouri, Nadia Fedorova, Bao Tran, Daniel Russell, Kristi Berry, Teresa Utterback, Susan E. Van Aken, Tamara V. Feldblyum, Mark D'Ascenzo, Wen-Ling Deng, et al. 2003. "The complete genome sequence of the Arabidopsis and tomato pathogen Pseudomonas syringae pv. tomato DC3000." PNAS, vol. 100, no. 18. (10181-10186)
Pathology:
- Qarah, Samar and Burke A. Cunha. 2003. "Pseudomonas aeruginosa infections." Emedicine.
- Toner, Charles B. and Stephen Krivda. 2003. "Pseudomonas folliculitits." Emedicine.
