Sulfolobus solfataricus: Difference between revisions
| Line 21: | Line 21: | ||
The ''S. solfataricus'' P2 genome has already been used extensively. It has been integral in the study of crenarchaeal biology, and has been compared to many euryarchaea to elucidate differences between the two groups. It has also provided more evidence for the separations of archaea, eukarya, and bacteria. The ''S. solfataricus'' P2 genome has also been manipulated for DNA studies of transcription, translation, the cell cycle, and other genetic events. Currently, the ''Sulfolobus solfataricus'' 98/2 strain is being sequenced. | The ''S. solfataricus'' P2 genome has already been used extensively. It has been integral in the study of crenarchaeal biology, and has been compared to many euryarchaea to elucidate differences between the two groups. It has also provided more evidence for the separations of archaea, eukarya, and bacteria. The ''S. solfataricus'' P2 genome has also been manipulated for DNA studies of transcription, translation, the cell cycle, and other genetic events. Currently, the ''Sulfolobus solfataricus'' 98/2 strain is being sequenced. | ||
Chrom.GIF | |||
==Cell Structure, Metabolism and Life Cycle== | ==Cell Structure, Metabolism and Life Cycle== | ||
Revision as of 04:58, 15 April 2009
Classification
Archaea; Crenarchaeota; Thermoprotei; Sulfolobales; Sulfolobaceae
Species
|
NCBI: Taxonomy |
Sulfolobus solfataricus
Description and Significance
Sulfolobus solfataricus grow in volcanic hot springs where they have ample sulfur and low pH. S. solfatricus is a very widely studied crenarchaeal organism. For this reason it is used as a model organism in archaeal research, with DNA replication, the cell cycle, chromosomal integration, and RNA processing is well researched and understood.
Proteins found in S. solfataricus are being investigated for use in biotechnology because they are active in a wide range of temperatures.
Genome Structure
The genome of S. solfataricus P2 was sequenced in 2001. It consists of a single circular chromosome containing 2,992,245 base pairs. There are 3,032 genes which encode 2,977 proteins. It has been shown that approximately one-third of the S. solfataricus P2 genome is unique. However, much of the genome (≈40%) is archaeal, while small portions overlap with bacteria and eukarya (12 and 2.3%, respectively). It has its own 743 open reading frames, and about 11% of the genome is mobile elements.
The S. solfataricus P2 genome has already been used extensively. It has been integral in the study of crenarchaeal biology, and has been compared to many euryarchaea to elucidate differences between the two groups. It has also provided more evidence for the separations of archaea, eukarya, and bacteria. The S. solfataricus P2 genome has also been manipulated for DNA studies of transcription, translation, the cell cycle, and other genetic events. Currently, the Sulfolobus solfataricus 98/2 strain is being sequenced.
Chrom.GIF
Cell Structure, Metabolism and Life Cycle
Interesting features of cell structure; how it gains energy; what important molecules it produces.
Ecology and Pathogenesis
Sulfolobus solfataricus is a thermoacidophile, and was first characterized in sulfur-rich volcanic springs in Yellowstone National Park. Strains of S. solfataricus have also been found in hot mud pools in the Solfatara crater north of Naples, Italy. It has also been discovered in El Salvador and the Dominica. While the optimum temperature for growth is ≈75oC, S. solfataricus can live in a range of 55-90oC. It can live in a pH range of 0.9-5.8, with its optimum being 2-3. However, S. solfataricus maintains its cytoplasmic pH at 6.5. It has also been shown that S. solfataricus requires an aerobic environment, in which it exists as a free-living sulfur chemolithotroph.
Strains of S. solfataricus are known to harbor plasmids that encourage bacteria-like conjugation between cells. However, the genes behind these mechanisms are dissimilar between S. solfataricus and bacteria.
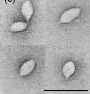
S. solfataricus can be infected by morphologically unusual viruses, such as SSV1 (Sulfolobus spindle-shaped virus). The virions formed by SSV1 often cluster together, forming rosette-like shapes. These viruses have been found in Yellowstone National Park, Iceland, and Japan.
References
Author
Page authored by Alvin Makohon-Moore and Antoni Malachowski, students of Prof. Jay Lennon at Michigan State University.