Rhodococcus ruber: Difference between revisions
| Line 39: | Line 39: | ||
==Cell Structure and Metabolism== | ==Cell Structure and Metabolism== | ||
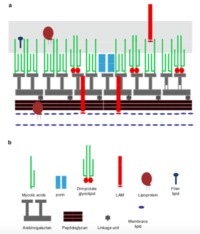
[[ File:Screen Shot 2020-04-29 at 4.19.14 AM.png| thumb| 200px| left| Figure _: ''Rhodococcus ruber'' cell wall structure [1] ]] | [[ File:Screen Shot 2020-04-29 at 4.19.14 AM.png| thumb| 200px| left| Figure _: ''Rhodococcus ruber'' cell wall structure [1] ]] | ||
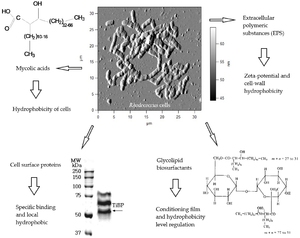
[[ File:Catalysts-09-00236-g002.png| thumb| 300px| right| Figure _: [] ]] | |||
It is a gram positive, non-motile, non-spore forming. | It is a gram positive, non-motile, non-spore forming. | ||
| Line 53: | Line 55: | ||
Have pronounced surfactant properties that facilitate the growth of the bacteria on hydrophobic substrates | Have pronounced surfactant properties that facilitate the growth of the bacteria on hydrophobic substrates | ||
==Ecological Impacts== | ==Ecological Impacts== | ||
Revision as of 09:13, 29 April 2020
Classification
Domain: Bacteria
Phylum: Actinobacteria
Class: Actinobacteria
Order: Corynbacteriales
Family: Nocardiaceae
Genus: Rhodococcus
Species
|
NCBI: [1] |
Rhodococcus ruber
Description and Significance
Historically, the genus Rhodococcus was first defined by Zopf in 1891. Nocardia rubra, or what is known today as Rhodococcus ruber, was isolated from a soil sludge by Kruse in 1896. However, there were issues with classification due to morphological and staining identification techniques. In 1977, Goodfellow and Alderson did a complete reclassification of the genus Rhodococcus, which resulted in Norcordia rubra being amended as Rhodococcus ruber.
The term Rhodococcus is from the combination of the Greek words “rhodon” and “coccus” meaning “the rose” and “the grain“ respectively. The morphological structure of R. ruber first starts as long rods, then breaks off into short rods and cocci throughout different growth phases. Rhodococcus ruber is a gram positive, non-motile, non-spore forming bacteria.
Genome Structure
12 genome studies have been reported for the following strains of Rhodococcus ruber: R. ruber str. YC-YT1, R. ruber str. YYL, R. ruber str. P14, R. ruber str. SD3, R. ruber str. R1, R. ruber str. DSM 43338, R. ruber str. P25, R. tuber IEGM 231, R. ruber str. OA1, R. ruber str. BKS 20-38, R. ruber str. Chol-4, and R. ruber str. NCRX 15591.
The genome of Rhodococcus ruber consists of one circular chromosome with an average size of 5.57 Mb. The genome size varies for every strain and larger genome sizes can be attributed to the presence of circular plasmids. The strains of R. ruber that contain plasmids are YC-TC1, YYL, and R1. The average amount of protein coding genes is 4932 and the average G+C content is 70.49%. A high guanine-cytosine content is characteristic of Rhodococcus species to provide DNA stability in varying conditions.
The genome sequences of R. ruber are helpful in identifying the genes necessary for metabolic pathways and cycles that R. ruber employs including the tricarboxylic acid cycle, pentose phosphate pathway, glycolysis, and glujconeogeneis. Additionally, the beta oxidation pathway and alkane degradation pathway are utilized. The genome highlights multiple metabolic gene clusters that are used for degradation of aromatic compounds, steroids, hydrocarbons, and xenobiotic polymers that R. ruber uses for carbon and energy sources. Some strains of R. ruber contain redundant genes which contributes to their versatile metabolic abilities with various nutrient sources.
Cell Structure and Metabolism
It is a gram positive, non-motile, non-spore forming.
Mycolic acids= Long chain fatty acids
Mycolic acids with chain lengths of C42-C48; much longer chains than other species of Rhodococcus
Cell envelope lipids
Have pronounced surfactant properties that facilitate the growth of the bacteria on hydrophobic substrates
Ecological Impacts
References
[1]| Alvarez, H. M. Biology of Rhodococcus; Springer, 2010; Vol. 16.
[2] | Bala
[3] | Farkas
[6] | Guevara
[8] | Krivorucho
[13] [Organism:noexp | Genome]
Author
Page authored by Hannah von Werder, student of Prof. Jay Lennon at Indiana University.