Agrobacterium Infection through Gene Transfer
Introduction

Agrobacterium is a genus of bacteria compose of gram negative soil bacteria. Many bacteria of this genus are known to cause a number of diseases in plants. Moreover, Agrobacterium are known for causing diseases through the use of horizontal gene transfer [5]. Tumors are the resulting symptoms produced in a wide variety of Agrobacterium caused plant diseases [5]. The most well-known of these diseases is crown gall disease caused by Agrobacterium tumefaciens. Other species of Agrobacterium that induce disease in plants include A. rhizogenes, known to cause ‘hairy root’ disease, as well as A. rubi that produces cane gall formation in raspberry and blackberry [8, 11]. All these bacteria infect their host by transferring their DNA into the plants. The different species of Agrobacterium use the same process and mechanisms for infection and transferring DNA. This ability for the bacteria to transfer its DNA into the host organism has made Agrobacterium a multifaceted tool in biotechnology as well as genetic engineering [11].
Plant Infection
The most studied Agrobacterium disease is crown gall disease caused by A. tumefaciens. Gall formation of this disease is initiated at the flesh wound site of the host plant [4]. From there the bacteria transfers its DNA into the host cell where it incorporates itself into the host genome. The insertion of the bacterial DNA causes formation of tumors in the host organism [5].
Chemotaxis
The bacteria is able to find wound sites on plants through sensing of metabolites and other molecules found at plant wounds. Phenolic metabolites such as acetosyringone, found to be involved in strengthening cell wall complexes, have been shown to induce infection of plant hosts. This is done through the triggering of virulence genes in the Agrobacterium [4]. Simple sugars and acidic pH, both signs of plant cell damage, also attract the bacterium to the wound site and induce infections [4].
Ti Plasmid
It has been found that virulence in Agrobacterium is dependent on tumor inducing plasmids or Ti plasmids found in the bacteria [5, 12]. Without this plasmid, the bacteria is unable to induce infection on its host. Many genes are found on this plasmid, including the tumor inducing oncogenes, the virulence genes needed for DNA transfer, as well as opine synthesis genes that provides a carbon source for the bacteria [5, 9]. However, not all of the genetic information on the plasmid is transferred into the host genome. Only a segment of the plasmid called transfer DNA or T-DNA is incorporated into the host during infection. T-DNA segments usually contain the oncogenes and opine synthesis genes, while the virulence genes are left out and not transferred [7]. This is because the proteins coded by the virulence genes are required for excision of the T-DNA from the plasmid within the bacteria as well as secreting the T-DNA into the host cell.
Vir Genes and T-DNA Secretion
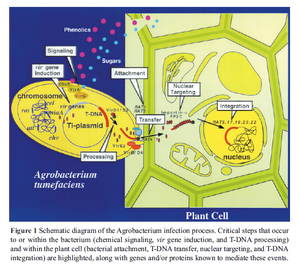
There are several virulence genes, named virA, virB, virC etc., all producing different proteins that have different roles in the transfer of T-DNA from Agrobacterium to its host. The VirA and VirG proteins are always expressed in the bacteria and make up a two component signal pathway where the binding of phenolic metabolites such as acetosyringone activates the kinase activity of VirA, which in turns phosphorylates and activates VirG [5]. Phospho-VirG is a transcription activator for the other vir genes, so the bacteria is only expresses the other vir gene products in the presence of phenolic compounds released from the wound site of the host. Once transcription of the other vir genes is activated by VirG, virD gene products bind to the right border of the T-DNA and cut a single strand of the T-DNA from the plasmid [1, 13]. Once the T-DNA segment is cut from the plasmid VirE2 proteins bind to and coat the single strand DNA [5]. The single strand T-DNA is secreted into the host cell by a type IV secretion system made up of various virulence gene proteins where it then sheds the VirE2 coating proteins and inserts itself randomly into the host genome [3, 14]. Once inside the host genome, the T-DNA is transcribed like any other gene in the host cell and the transcription products cause tumor formation in the host. Tumor formation does not kill the host and is used to create a hospitable environment for the bacteria. The transformed cells produce opine as a carbon source for the bacteria to strive on [5].
Human Infection
While Agrobacterium infection is more prevalent in plants, they are also able to establish infection in humans as well. They are not known for being a primary pathogen in healthy individuals, but there are many cases reported showing Agrobacterium to be opportunistic pathogens. In individuals with a weakened immune system, Agrobacterium have been known to establish opportunistic infections [2, 6]. However, there has not been any reported cases of DNA transfer by Agrobacterium into infected individuals. Nonetheless, scientists have been able to transfer DNA into cultivated human HeLa cells with the use of Agrobacterium [10]. But nothing can be concluded from this experiment about Agrobacterium infection though gene transfer of humans in nature.
Uses of Agrobacterium
Besides plant infection, Agrobacterium have been used in a variety of ways by humans, mainly for the bacteria’s ability to transfer DNA. This ability can be used to create mutant or transformed organisms. Because the insertion of DNA is random in the host cell, the T-DNA can disrupt and silence a certain gene, creating a mutant strain of the organism that can be used for research [12]. The T-DNA can also be altered so transformed organisms express the product of the T-DNA inserted gene. These transformed organisms can be used for research as well as creating GMO’s for agricultural uses [11, 12].
References
1. Dafny-Yelin, M., Levy, A., & Tzfira, T. (2008). The ongoing saga of Agrobacterium–host interactions. Trends in Plant Science, 13(3), 102-105.
2. Dunne, W. M.,Jr, Tillman, J., & Murray, J. C. (1993). Recovery of a strain of agrobacterium radiobacter with a mucoid phenotype from an immunocompromised child with bacteremia. Journal of Clinical Microbiology,31(9), 2541-2543.
3. Francis, K. E., & Spiker, S. (2005). Identification of arabidopsis thaliana transformants without selection reveals a high occurrence of silenced T-DNA integrations. The Plant Journal : For Cell and Molecular Biology,41(3), 464.
4. Gartland, K. M. A., Davey, M. R., & ebrary, I. (1995). Agrobacterium protocols. Totowa, N.J: Humana Press.
5. Gelvin, S. (2000). Agrobacterium and plant genes involved in T-DNA transfer and integration. Annual Review of Plant Physiology and Plant Molecular Biology, 51, 223-256.
6. Hulse, M., Johnson, S., & Ferrieri, P. (1993). Agrobacterium infections in humans: Experience at one hospital and review. Clinical Infectious Diseases : An Official Publication of the Infectious Diseases Society of America,16(1), 112-117.
7. Lacroix, B., Li, J., Tzfira, T., & Citovsky, V. (2006). Will you let me use your nucleus? how agrobacterium gets its T-DNA expressed in the host plant cell. Canadian Journal of Physiology and Pharmacology, 84(3-4), 333-345.
8. Ondrej, M., Matousek, J., & Vlasak, J. (1987). Differentiation of transformed plants from tumors induced by agrobacterium rubi ATCC 13335. Journal of Plant Physiology, 126(4-5), 397-407.
9. Pitzschke, A. & Hirt, H. (2010). New insights into an old story: Agrobacterium -induced tumour formation in plants by plant transformation. The EMBO Journal, 29(6), 1021-1032.
10. Talya Kunik, Tzvi Tzfira, Yoram Kapulnik, Yedidya Gafni, Colin Dingwall, & Vitaly Citovsky. (2001). Genetic transformation of HeLa cells by agrobacterium. Proceedings of the National Academy of Sciences of the United States of America, 98(4), 1871-1876.
11. Tepfer, D. (1990). Genetic-transformation using agrobacterium-rhizogenes. Physiologia Plantarum, 79(1), 140-146.
12. Tzfira, T., Citovsky, V., & ebrary, I. (2008). Agrobacterium: From biology to biotechnology. New York: Springer.
13. Tzfira, T., Li, J., Lacroix, B., & Citovsky, V. (2004). Agrobacterium T-DNA integration: Molecules and models. Trends in Genetics, 20(8), 375-383.
14. Wallden, K., Rivera-Calzada, A., & Waksman, G. (2010). Type IV secretion systems: Versatility and diversity in function. Cellular Microbiology, 12(9), 1203-1212.