Bacterial nucleation in pseudomonas syringae
From MicrobeWiki, the student-edited microbiology resource
Introduction
Pseudomonas syringae
[Image 1]
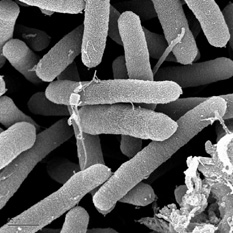
Pseudomonas syringae shown using SEM. Source: Gordon Vrdoljak, Electron Microscopy Laboratory, U.C. Berkeley [1]
Pseudomonas syringae shown using SEM. Source: Gordon Vrdoljak, Electron Microscopy Laboratory, U.C. Berkeley [1]
Ice Nucleation Active (INA) Proteins
Current Research
Conclusion
References
[1] http://genome.jgi-psf.org/psesy/psesy.home.html [2]
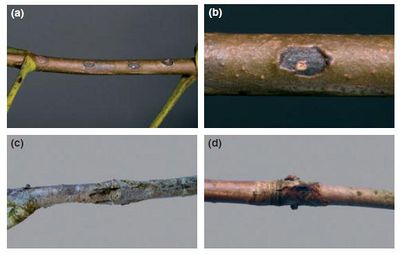
[2] Steele, H., B.E. Laue, G.A. MacAskill, A.J. Hendry, and S. Green. “Analysis of the natural infection of European horse chestnut (Aesculus hippocastanum) by Pseudomonas syringae pv. Aesculi.” Plant Pathology 59: 1005-1013.
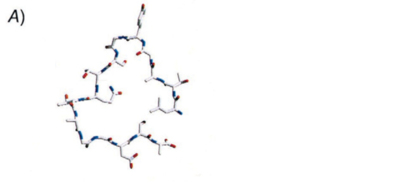
[3] Graether, S.P., and Z. Jia. 2001. Modeling Pseudomonas syringae ice-nucleation protein as a B-helical protein. Biophysical Journal 80: 1169-1173.
Edited by Ryan O'Connor,student of Joan Slonczewski for BIOL 238 Microbiology, 2011, Kenyon College.