Chimpanzee Evolution: Difference between revisions
Schwingel1 (talk | contribs) |
Schwingel1 (talk | contribs) |
||
| Line 1: | Line 1: | ||
==Introduction and Early Evolution== | ==Introduction and Early Evolution== | ||
[[Image:Pan troglodytes .jpg|thumb|250px|right|<b>Figure 1</b>: Adult male chimpanzee (<i>Pan troglodytes</i>); photo taken 26 October 2006 in Shanghai Wildlife Park, Shanghai, China. Photographer: David Blank[https://animaldiversity.org/collections/contributors/david_blank/chimpmale/large.jpg].]] | [[Image:Pan troglodytes .jpg|thumb|250px|right|<b>Figure 1</b>: Adult male chimpanzee (<i>Pan troglodytes</i>); photo taken 26 October 2006 in Shanghai Wildlife Park, Shanghai, China. Photographer: David Blank[https://animaldiversity.org/collections/contributors/david_blank/chimpmale/large.jpg].]] | ||
<i>Pan troglodytes</i>, better known as chimpanzees, are a species of great ape widely regarded as the closest living relatives to bonobos (<i>Pan paniscus</i>) and one of the closest living relatives to humans (<i>Homo sapiens</i>; Fig. 1).<ref name = Bjork/>< [https://academic.oup.com/mbe/article/28/1/615/984161 Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.]</ref><ref>[https://www.sciencedirect.com/science/article/pii/S0047248404001666?casa_token=uz3pgX8ckB8AAAAA:LcGJiaqHBe-Xl4i_dq39AsQ6j236VPkcj_mBQLjJRjvPPJKamjZDkAne7Zzpl0vZvwhWhLsOrg Raaum RL, Sterner KN, Noviello CM, Stewart CB, Disotell TR. Catarrhine primate divergence dates estimated from complete mitochondrial genomes: Concordance with fossil and nuclear DNA evidence. J Hum Evol. 2005;48(3):237–57.]</ref> | <i>Pan troglodytes</i>, better known as chimpanzees, are a species of great ape widely regarded as the closest living relatives to bonobos (<i>Pan paniscus</i>) and one of the closest living relatives to humans (<i>Homo sapiens</i>; Fig. 1).<ref name = Bjork/><[https://academic.oup.com/mbe/article/28/1/615/984161 Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.]</ref><ref>[https://www.sciencedirect.com/science/article/pii/S0047248404001666?casa_token=uz3pgX8ckB8AAAAA:LcGJiaqHBe-Xl4i_dq39AsQ6j236VPkcj_mBQLjJRjvPPJKamjZDkAne7Zzpl0vZvwhWhLsOrg Raaum RL, Sterner KN, Noviello CM, Stewart CB, Disotell TR. Catarrhine primate divergence dates estimated from complete mitochondrial genomes: Concordance with fossil and nuclear DNA evidence. J Hum Evol. 2005;48(3):237–57.]</ref> | ||
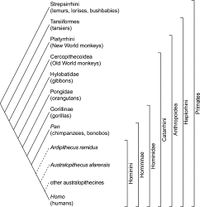
The evolution of chimpanzees is thought to have begun, like that of many other primates, with the divergence of platyrrhines (also known as New World monkeys) and catarrhines (a group containing Old World monkeys and apes) 25-40 million years ago <ref>[https://www.sciencedirect.com/science/article/pii/S0960982207003673 Stewart CB, Disotell TR. Primate evolution - In and out of Africa. Curr Biol. 1998;8(16):582–8.]</ref> This was followed by a divergence in the catarrhines, forming the Hominoidea (ape) and Cercopithecoidea (Old World monkey) groups 23 million years ago. <ref>[https://www.sciencedirect.com/science/article/pii/S0047248404001666?casa_token=uz3pgX8ckB8AAAAA:LcGJiaqHBe-Xl4i_dq39AsQ6j236VPkcj_mBQLjJRjvPPJKamjZDkAne7Zzpl0vZvwhWhLsOrg Raaum RL, Sterner KN, Noviello CM, Stewart CB, Disotell TR. Catarrhine primate divergence dates estimated from complete mitochondrial genomes: Concordance with fossil and nuclear DNA evidence. J Hum Evol. 2005;48(3):237–57.]</ref> It is estimated that the genera <i>Homo</i> and <i>Pan</i> diverged 5-6 million years ago, followed by a split in the genus <i>Pan</i> roughly 2.4 million years ago, eventually culminating into the chimpanzees and bonobos present today (Fig. 2). | The evolution of chimpanzees is thought to have begun, like that of many other primates, with the divergence of platyrrhines (also known as New World monkeys) and catarrhines (a group containing Old World monkeys and apes) 25-40 million years ago <ref>[https://www.sciencedirect.com/science/article/pii/S0960982207003673 Stewart CB, Disotell TR. Primate evolution - In and out of Africa. Curr Biol. 1998;8(16):582–8.]</ref> This was followed by a divergence in the catarrhines, forming the Hominoidea (ape) and Cercopithecoidea (Old World monkey) groups 23 million years ago. <ref>[https://www.sciencedirect.com/science/article/pii/S0047248404001666?casa_token=uz3pgX8ckB8AAAAA:LcGJiaqHBe-Xl4i_dq39AsQ6j236VPkcj_mBQLjJRjvPPJKamjZDkAne7Zzpl0vZvwhWhLsOrg Raaum RL, Sterner KN, Noviello CM, Stewart CB, Disotell TR. Catarrhine primate divergence dates estimated from complete mitochondrial genomes: Concordance with fossil and nuclear DNA evidence. J Hum Evol. 2005;48(3):237–57.]</ref> It is estimated that the genera <i>Homo</i> and <i>Pan</i> diverged 5-6 million years ago, followed by a split in the genus <i>Pan</i> roughly 2.4 million years ago, eventually culminating into the chimpanzees and bonobos present today (Fig. 2). | ||
Revision as of 02:40, 9 December 2020
Introduction and Early Evolution

Pan troglodytes, better known as chimpanzees, are a species of great ape widely regarded as the closest living relatives to bonobos (Pan paniscus) and one of the closest living relatives to humans (Homo sapiens; Fig. 1).[1]<Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.</ref>[2]
The evolution of chimpanzees is thought to have begun, like that of many other primates, with the divergence of platyrrhines (also known as New World monkeys) and catarrhines (a group containing Old World monkeys and apes) 25-40 million years ago [3] This was followed by a divergence in the catarrhines, forming the Hominoidea (ape) and Cercopithecoidea (Old World monkey) groups 23 million years ago. [4] It is estimated that the genera Homo and Pan diverged 5-6 million years ago, followed by a split in the genus Pan roughly 2.4 million years ago, eventually culminating into the chimpanzees and bonobos present today (Fig. 2). [1][5] [6][7]
Continuing Evolution and Speciation
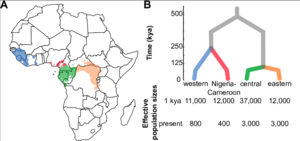
Evolution has not stopped with the modern chimpanzee. There are currently four recognized subspecies: the central chimpanzee (P. t. troglodytes), the western chimpanzee (P. t. verus), the eastern chimpanzee (P. t. schweinfurthii), and the Nigeria-Cameroon chimpanzee (P. t. ellioti) (Fig. 3). [8][9][10][11]
The subspecies are geographically isolated by the Sanaga River, which separates the western and Nigeria-Cameroon subspecies from the central and eastern chimpanzees, and the Ubangi River, which is the primary barrier between the latter two subspecies. [12] The Niger River and the Dahomey Gap (a stretch of dry forest across Western Africa) have both been hypothesized as barriers that have separated the western and Nigeria-Cameroon chimpanzees.[13] However, not all of these geographic barriers are complete; researchers have recently recorded migration between eastern and central chimpanzee subspecies.[14][15] Sequencing of mitochondrial DNA has revealed that there are two major chimpanzee lineages; one clade is formed by the central and eastern chimpanzees, while western and Nigeria-Cameroon chimpanzees form the other clade.[16] Subspeciation is further enforced by variable susceptibility to specific viral infections.[17][18]Simian immunodeficiency virus (SIV), the precursor to the human immunodeficiency virus (HIV), is known to infect only P. t. troglodytes and P. t. schweinfurthii.[19][20] Chimpanzee subspecies have also shown specific behavioral differences, most notably that nut-cracking behavior only occurs in the west.[21][22]
In addition to continuous speciation, chimpanzees face extinction.[23][24] [25] Each subspecies is classified as endangered, the western chimpanzee critically so, having declined by about 80% in 24 years.[26] This population decrease has been attributed to factors including illegal hunting, habitat loss, and infectious diseases; regional extinctions in Burkina Faso and Côte d’Ivoire have reduced the western chimpanzee’s geographic range by 20%.[27]
Distinct Anatomical Adaptations
Physiology
Chimpanzees, living in a variety of environments, are adapted in a variety of ways. The effect of environment is evident in locomotion, suitable for both terrestrial and arboreal settings.[28] Scientists have recorded instances of quadrupedalism and bipedalism in chimpanzee populations; neither type of locomotion is favored over the other in terms of energy efficiency, but locomotion in general has a relatively high metabolic cost in chimpanzees (41% more energy than expected for a mammal of similar body mass)[29] In addition, when bipedal, chimpanzees tend to take relatively small, frequent steps and require more side-to-side motion than habitual bipeds, typical of non-human primates.[30] This may be indicative of the lack of specialized anatomical structures for bipedalism; with a necessity to move effectively in a variety of environments, chimpanzees benefit from different possible types of locomotion.[31][32]
Many of the chimpanzee’s features are studied in relation to its human relatives, particularly concerning a chimpanzee-human last common ancestor and the rise of habitual bipedality.[33][34][35][36] For example, chimpanzees and humans are depicted similarly in the effect of aging on the thoracolumbar region, for which there is a difference seen in the region based on age (as opposed to the thoracolumbar region of gorillas).[37] In further comparison to Homo sapiens, alternative specialization is often studied in chimpanzees as means of characterizing the last common Pan-Homo common ancestor. This is evident in a comparison of muscle strength in the two primates; chimpanzees, on average, display a mass-specific muscular performance 1.5 times greater than humans, which may be similar to that of the last common ancestor between humans and chimpanzees. [38]
Gut Microbiome
Chimpanzees have also developed a distinct microbiome, most notably in the gut, which fluctuates socially and seasonally. [39][40] Chimpanzee gut microbiomes are mostly similar to those of other great apes, with a few key differences. [41] The gut microbiome is largely regulated by diet; because chimpanzees are primarily (though not exclusively) frugivores, their gut microbiome does not fluctuate nearly as much as those of other great apes who might be more seasonal eaters.[42] However, this does not mean that the chimpanzee gut microbiome does not fluctuate seasonally; in a 2018 study, it was noted that chimpanzees were distinguished by the relative abundance of Erysipelotrichaceae taxa, Prevotella, RFN20, and Sphaerochaeta. [43] ). During dry months (May-October), chimpanzees had a considerable amount of Prevotella, while wet months (November-April) were characterized by high levels of Sphaerochaeta. [44][45] This seasonal fluctuation is coupled with social fluctuation.[46] Socially-based microbial development, unlike seasonal fluctuations, affect species richness within individuals, rather than specific bacterial abundance. [47] Chimpanzees display different social patterns in accordance with the season; while dry seasons are usually associated with relative solitude, wet seasons correspond with high sociability.[48] Furthermore, inheritance of gut microbial communities appears to be primarily horizontal, rather than vertically from parent to offspring, suggesting that sociability is necessary to maintain and develop microbial communities in chimpanzees.[49] In this way, chimpanzee populations cultivate microbial communities, in which microbiomes are connected by the interaction of their hosts; microbes have even been known to transfer between the guts of chimpanzees and gorillas.[50][51] ). The combination of seasonal and social fluctuations in the gut microbiome has granted chimpanzees resilience to changes in nutrient availability and disease.[52][53]
Behavioral Adaptations
Behavior in chimpanzees, much like that of other primates, is thought to be the variable result of a combination of genetics and the environment.[54] Chimpanzee behavior, therefore, varies, often in accordance with population; each population exhibits broad behavioral patterns, ranging from rain dances to use of levers (Fig. 4).[55]
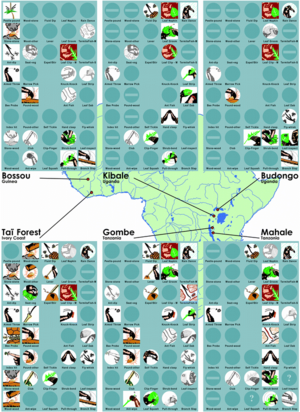
Tool use, in particular, has been studied in chimpanzees.[56][57][58] The use of percussive tools, most commonly used to crack nuts, is a common example of tool use in chimpanzees and currently the only significant instance of behavioral differentiation between subspecies, as opposed to populations.[59][60] In foraging and hunting, chimpanzees often use probing sticks to collect insects or honey, though specific use varies by population.[61][62] In addition, chimpanzees have been known to fashion spears and lances, made of wood and sharpened with their teeth, to puncture prey (Fig. 5).[63]

In hunting, chimpanzees and their demeanor can be more readily compared to members of the genus Homo than to other primates using similar methods, suggesting that genetics have played a role in tool use, despite common techniques among other primates.[64]
Contributions to Medicine and a Collective Understanding of Homo sapiens
Disease
Attempting to understand chimpanzees and how they have evolved has resulted in crucial insight into issues that plague Homo sapiens, providing an expanded understanding of human disease and medicine. [65][66][67][68][69] By studying chimpanzees, scientists have come closer to understanding malaria (Plasmodium falciparum), a mosquito-borne illness that continues to plague certain human populations today, and how it evolved (Fig. 6). [70]
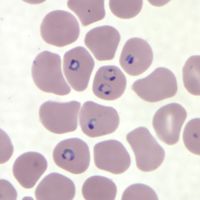
Chimpanzees are affected by similar strains of malaria (P. reichenowi and P. gaboni), which likely gave rise to the common human strain through horizontal transfer and subsequent speciation. [71]
Trisomy 22
In addition to pathogenic diseases, the study of chimpanzees allows scientists to gather information about genetic diseases.[72] Human trisomy 21, commonly known as Down syndrome, has been mirrored in chimpanzees in trisomy of the 22nd chromosome.[73] In a study published in 2017, a chimpanzee affected with trisomy 22 was reported to show similar symptoms to humans affected with trisomy 21, including early-onset vision impediments, congenital heart defects, hypodontia, and considerable social difficulties, contributed to by blindnes (Fig. 7).[74]

Evidence of somewhat shared disabilities between humans and chimpanzees can contribute both to a more developed understanding of relatedness between the species and potentially grant insight into treating such illnesses.
Vaccines
While chimpanzees can give insight into common illnesses, they can also be integral in the development of vaccines. Adenoviruses (Ads), a type of double-stranded DNA virus that broadly infects vertebrates, have historically been used as vaccine vectors.[75] Although human serotypes of adenovirus (AdHus) have frequently been used, studies of chimpanzee adenoviruses (AdCs) are rising in prevalence due to a number of apparent benefits, including increased immunogenicity and a decrease in neutralizing cell responses in comparison to human adenoviral vectors.[76] [77] Chimpanzee adenovirus vectors have been developed in response to a number of pervasive and otherwise common diseases; in response to the 2016 Ebola outbreak, for example, researchers developed AdCs that successfully activated memory T-cell responses, distributed equally between CD4 and CD8 T-cells.[78][79]
In January 2017, the Coalition for Epidemic Preparedness Innovations (CEPI) was formed with the purpose of creating and stockpiling vaccines for a number of pathogens with pandemic potential, including Ebola virus disease, Nipah Virus, Lassa virus, and novel coronaviruses.[80] In this case, scientists based the vaccines for outbreak pathogens on chimpanzee adenovirus vectors because of their relative safety and thus likeliness to be approved for temporary emergency use, if necessary.[81] In 2017, CEPI attempted to combat novel coronaviruses with a focus on preventing expression of spike proteins, nucleocapsid proteins, or both, with significant resource limitations in the laboratory.[82] As of July 2020, however, a chimpanzee-adenovirus-vectored vaccine (ChAd-SARS-CoV-2-S) has shown promising results in testing, reporting complete prevention of SARS-CoV-2 infection in the upper and lower respiratory tracts when administered intranasally.[83] In the development of vaccines, chimpanzees continue to provide important insight and treatment for a number of human diseases.[84][85][86]
AdCs have also been altered for use in cancer treatment, bypassing pre-existing neutralizing antibodies that posed a problem for human adenoviral vectors; a particular AdC vector (AdC7-SP/E1A-E3) has been shows to have significant anti-tumor effects in certain cancer cells and in mice.[87]
Conclusion
Over the past 40 million years, Pan troglodytes have evolved into a distinct species that is often studied for its similarity to other great apes, particularly Homo sapiens. Specific genetic and behavioral adaptations, particularly those that relate to tool use and potential culture, highlight not only how chimpanzees are a unique species, but how they are similar to both human and other non-human primates. A growing understanding of chimpanzee genetics and evolutionary relationships allows humans to further develop an understanding of shared genetics and therefore lends great insight into disease origin and treatment; genetic similarity between chimpanzees and humans also allows for significant advancements in the field of medicine and pathology. Continuing evolution of Pan troglodytes has given rise to four clear subspecies, each demonstrating the effect of the environment and incomplete geographic isolation on primate populations. However, with growing threats to chimpanzee habitats and rapidly dwindling population sizes, the continuing evolution and survival of chimpanzees is at risk.
References
- ↑ 1.0 1.1 Cite error: Invalid
<ref>tag; no text was provided for refs namedBjork - ↑ Raaum RL, Sterner KN, Noviello CM, Stewart CB, Disotell TR. Catarrhine primate divergence dates estimated from complete mitochondrial genomes: Concordance with fossil and nuclear DNA evidence. J Hum Evol. 2005;48(3):237–57.
- ↑ Stewart CB, Disotell TR. Primate evolution - In and out of Africa. Curr Biol. 1998;8(16):582–8.
- ↑ Raaum RL, Sterner KN, Noviello CM, Stewart CB, Disotell TR. Catarrhine primate divergence dates estimated from complete mitochondrial genomes: Concordance with fossil and nuclear DNA evidence. J Hum Evol. 2005;48(3):237–57.
- ↑ Raaum RL, Sterner KN, Noviello CM, Stewart CB, Disotell TR. Catarrhine primate divergence dates estimated from complete mitochondrial genomes: Concordance with fossil and nuclear DNA evidence. J Hum Evol. 2005;48(3):237–57.
- ↑ Stewart CB, Disotell TR. Primate evolution - In and out of Africa. Curr Biol. 1998;8(16):582–8.
- ↑ S, Hedges SB. A molecular timescale for vertebrate development. Nature [Internet. 1998;392:917–20.]
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Palaeontographica VA, Das P, Whiten a, Goodall J, Mcgrew WC, Nishida T, et al. Cultures in chimpanzees troglodytes ) have achieved long-term status across Africa , differ- lated 151 years of chimpanzee observation . This comprehensive analysis reveals patterns of variation that are far more extensive than have previously been docume. Nature. 1999;399(JUNE):15–8.
- ↑ Schmidt JM, De Manuel M, Marques-Bonet T, Castellano S, Andrés AM. The impact of genetic adaptation on chimpanzee subspecies differentiation. Vol. 15, PLoS Genetics. 2019. 1–32 p.
- ↑ Bowden R, MacFie TS, Myers S, Hellenthal G, Nerrienet E, Bontrop RE, et al. Genomic tools for evolution and conservation in the chimpanzee: Pan troglodytes ellioti is a genetically distinct population. PLoS Genet. 2012;8(3):1–10.
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Schmidt JM, De Manuel M, Marques-Bonet T, Castellano S, Andrés AM. The impact of genetic adaptation on chimpanzee subspecies differentiation. Vol. 15, PLoS Genetics. 2019. 1–32 p.
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Schmidt JM, De Manuel M, Marques-Bonet T, Castellano S, Andrés AM. The impact of genetic adaptation on chimpanzee subspecies differentiation. Vol. 15, PLoS Genetics. 2019. 1–32 p.
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Schmidt JM, De Manuel M, Marques-Bonet T, Castellano S, Andrés AM. The impact of genetic adaptation on chimpanzee subspecies differentiation. Vol. 15, PLoS Genetics. 2019. 1–32 p.
- ↑ Palaeontographica VA, Das P, Whiten a, Goodall J, Mcgrew WC, Nishida T, et al. Cultures in chimpanzees troglodytes ) have achieved long-term status across Africa , differ- lated 151 years of chimpanzee observation . This comprehensive analysis reveals patterns of variation that are far more extensive than have previously been docume. Nature. 1999;399(JUNE):15–8.
- ↑ Proffitt T, Haslam M, Mercader JF, Boesch C, Luncz L V. Revisiting Panda 100, the first archaeological chimpanzee nut-cracking site. J Hum Evol. 2018;124:117–39.
- ↑ Schmidt JM, De Manuel M, Marques-Bonet T, Castellano S, Andrés AM. The impact of genetic adaptation on chimpanzee subspecies differentiation. Vol. 15, PLoS Genetics. 2019. 1–32 p.
- ↑ Kühl HS, Sop T, Williamson EA, Mundry R, Brugière D, Campbell G, et al. The Critically Endangered western chimpanzee declines by 80%. Am J Primatol. 2017;79(9).
- ↑ Kalan AK, Kulik L, Arandjelovic M, Boesch C, Haas F, Dieguez P, et al. Environmental variability supports chimpanzee behavioural diversity. Nat Commun. 2020;11(1).
- ↑ Kühl HS, Sop T, Williamson EA, Mundry R, Brugière D, Campbell G, et al. The Critically Endangered western chimpanzee declines by 80%. Am J Primatol. 2017;79(9).
- ↑ Kühl HS, Sop T, Williamson EA, Mundry R, Brugière D, Campbell G, et al. The Critically Endangered western chimpanzee declines by 80%. Am J Primatol. 2017;79(9).
- ↑ Pontzer H, Raichlen DA, Rodman PS. Bipedal and quadrupedal locomotion in chimpanzees. J Hum Evol [Internet. 2014;66(1):64–82. Available from: http://dx.doi.org/10.1016/j.jhevol.2013.10.002]
- ↑ Pontzer H, Raichlen DA, Rodman PS. Bipedal and quadrupedal locomotion in chimpanzees. J Hum Evol [Internet. 2014;66(1):64–82. Available from: http://dx.doi.org/10.1016/j.jhevol.2013.10.002]
- ↑ Pontzer H, Raichlen DA, Rodman PS. Bipedal and quadrupedal locomotion in chimpanzees. J Hum Evol [Internet. 2014;66(1):64–82. Available from: http://dx.doi.org/10.1016/j.jhevol.2013.10.002]
- ↑ Pontzer H, Raichlen DA, Rodman PS. Bipedal and quadrupedal locomotion in chimpanzees. J Hum Evol [Internet. 2014;66(1):64–82. Available from: http://dx.doi.org/10.1016/j.jhevol.2013.10.002]
- ↑ Nalley TK, Scott JE, Ward C V., Alemseged Z. Comparative morphology and ontogeny of the thoracolumbar transition in great apes, humans, and fossil hominins. J Hum Evol [Internet. 2019;134:102632. Available from: https://doi.org/10.1016/j.jhevol.2019.06.003]
- ↑ Nalley TK, Scott JE, Ward C V., Alemseged Z. Comparative morphology and ontogeny of the thoracolumbar transition in great apes, humans, and fossil hominins. J Hum Evol [Internet. 2019;134:102632. Available from: https://doi.org/10.1016/j.jhevol.2019.06.003]
- ↑ Arlegi M, Gómez-Robles A, Gómez-Olivencia A. Morphological integration in the gorilla, chimpanzee, and human neck. Am J Phys Anthropol. 2018;166(2):408–16.
- ↑ O’Neill MC, Umberger BR, Holowka NB, Larson SG, Reiser PJ, Slade JM. Chimpanzee super strength and human skeletal muscle evolution. Proc Natl Acad Sci U S A. 2017;114(28):7343–8.
- ↑ Bründl AC, Tkaczynski PJ, Nohon Kohou G, Boesch C, Wittig RM, Crockford C. Systematic mapping of developmental milestones in wild chimpanzees. Dev Sci. 2020;(December 2019):1–13.
- ↑ Nalley TK, Scott JE, Ward C V., Alemseged Z. Comparative morphology and ontogeny of the thoracolumbar transition in great apes, humans, and fossil hominins. J Hum Evol [Internet. 2019;134:102632. Available from: https://doi.org/10.1016/j.jhevol.2019.06.003]
- ↑ O’Neill MC, Umberger BR, Holowka NB, Larson SG, Reiser PJ, Slade JM. Chimpanzee super strength and human skeletal muscle evolution. Proc Natl Acad Sci U S A. 2017;114(28):7343–8.
- ↑ Hicks AL, Lee KJ, Couto-Rodriguez M, Patel J, Sinha R, Guo C, et al. Gut microbiomes of wild great apes fluctuate seasonally in response to diet. Nat Commun [Internet. 2018;9(1). Available from: http://dx.doi.org/10.1038/s41467-018-04204-w]
- ↑ Moeller AH, Foerster S, Wilson ML, Pusey AE, Hahn BH, Ochman H. Social behavior shapes the chimpanzee pan-microbiome. Sci Adv. 2016;2(1).
- ↑ Hicks AL, Lee KJ, Couto-Rodriguez M, Patel J, Sinha R, Guo C, et al. Gut microbiomes of wild great apes fluctuate seasonally in response to diet. Nat Commun [Internet. 2018;9(1). Available from: http://dx.doi.org/10.1038/s41467-018-04204-w]
- ↑ Hicks AL, Lee KJ, Couto-Rodriguez M, Patel J, Sinha R, Guo C, et al. Gut microbiomes of wild great apes fluctuate seasonally in response to diet. Nat Commun [Internet. 2018;9(1). Available from: http://dx.doi.org/10.1038/s41467-018-04204-w]
- ↑ Hicks AL, Lee KJ, Couto-Rodriguez M, Patel J, Sinha R, Guo C, et al. Gut microbiomes of wild great apes fluctuate seasonally in response to diet. Nat Commun [Internet. 2018;9(1). Available from: http://dx.doi.org/10.1038/s41467-018-04204-w]
- ↑ Hicks AL, Lee KJ, Couto-Rodriguez M, Patel J, Sinha R, Guo C, et al. Gut microbiomes of wild great apes fluctuate seasonally in response to diet. Nat Commun [Internet. 2018;9(1). Available from: http://dx.doi.org/10.1038/s41467-018-04204-w]
- ↑ Moeller AH, Foerster S, Wilson ML, Pusey AE, Hahn BH, Ochman H. Social behavior shapes the chimpanzee pan-microbiome. Sci Adv. 2016;2(1).
- ↑ Moeller AH, Foerster S, Wilson ML, Pusey AE, Hahn BH, Ochman H. Social behavior shapes the chimpanzee pan-microbiome. Sci Adv. 2016;2(1).
- ↑ Moeller AH, Foerster S, Wilson ML, Pusey AE, Hahn BH, Ochman H. Social behavior shapes the chimpanzee pan-microbiome. Sci Adv. 2016;2(1).
- ↑ Moeller AH, Foerster S, Wilson ML, Pusey AE, Hahn BH, Ochman H. Social behavior shapes the chimpanzee pan-microbiome. Sci Adv. 2016;2(1).
- ↑ Moeller AH, Foerster S, Wilson ML, Pusey AE, Hahn BH, Ochman H. Social behavior shapes the chimpanzee pan-microbiome. Sci Adv. 2016;2(1).
- ↑ Moeller AH, Foerster S, Wilson ML, Pusey AE, Hahn BH, Ochman H. Social behavior shapes the chimpanzee pan-microbiome. Sci Adv. 2016;2(1).
- ↑ Moeller AH, Peeters M, Ndjango JB, Li Y, Hahn BH, Ochman H. Sympatric chimpanzees and gorillas harbor convergent gut microbial communities. Genome Res. 2013;23(10):1715–20.
- ↑ Hicks AL, Lee KJ, Couto-Rodriguez M, Patel J, Sinha R, Guo C, et al. Gut microbiomes of wild great apes fluctuate seasonally in response to diet. Nat Commun [Internet. 2018;9(1). Available from: http://dx.doi.org/10.1038/s41467-018-04204-w]
- ↑ Moeller AH, Foerster S, Wilson ML, Pusey AE, Hahn BH, Ochman H. Social behavior shapes the chimpanzee pan-microbiome. Sci Adv. 2016;2(1).
- ↑ Greene HW. Evolutionary scenarios and primate natural history. Am Nat. 2017;190(august):S69–86.
- ↑ Palaeontographica VA, Das P, Whiten a, Goodall J, Mcgrew WC, Nishida T, et al. Cultures in chimpanzees troglodytes ) have achieved long-term status across Africa , differ- lated 151 years of chimpanzee observation . This comprehensive analysis reveals patterns of variation that are far more extensive than have previously been docume. Nature. 1999;399(JUNE):15–8.
- ↑ Palaeontographica VA, Das P, Whiten a, Goodall J, Mcgrew WC, Nishida T, et al. Cultures in chimpanzees troglodytes ) have achieved long-term status across Africa , differ- lated 151 years of chimpanzee observation . This comprehensive analysis reveals patterns of variation that are far more extensive than have previously been docume. Nature. 1999;399(JUNE):15–8.
- ↑ Proffitt T, Haslam M, Mercader JF, Boesch C, Luncz L V. Revisiting Panda 100, the first archaeological chimpanzee nut-cracking site. J Hum Evol. 2018;124:117–39.
- ↑ Greene HW. Evolutionary scenarios and primate natural history. Am Nat. 2017;190(august):S69–86.
- ↑ Palaeontographica VA, Das P, Whiten a, Goodall J, Mcgrew WC, Nishida T, et al. Cultures in chimpanzees troglodytes ) have achieved long-term status across Africa , differ- lated 151 years of chimpanzee observation . This comprehensive analysis reveals patterns of variation that are far more extensive than have previously been docume. Nature. 1999;399(JUNE):15–8.
- ↑ Proffitt T, Haslam M, Mercader JF, Boesch C, Luncz L V. Revisiting Panda 100, the first archaeological chimpanzee nut-cracking site. J Hum Evol. 2018;124:117–39.
- ↑ Palaeontographica VA, Das P, Whiten a, Goodall J, Mcgrew WC, Nishida T, et al. Cultures in chimpanzees troglodytes ) have achieved long-term status across Africa , differ- lated 151 years of chimpanzee observation . This comprehensive analysis reveals patterns of variation that are far more extensive than have previously been docume. Nature. 1999;399(JUNE):15–8.
- ↑ Greene HW. Evolutionary scenarios and primate natural history. Am Nat. 2017;190(august):S69–86.
- ↑ Greene HW. Evolutionary scenarios and primate natural history. Am Nat. 2017;190(august):S69–86.
- ↑ Greene HW. Evolutionary scenarios and primate natural history. Am Nat. 2017;190(august):S69–86.
- ↑ Sundararaman SA, Plenderleith LJ, Liu W, Loy DE, Learn GH, Li Y, et al. Genomes of cryptic chimpanzee Plasmodium species reveal key evolutionary events leading to human malaria. Nat Commun. 2016;7.
- ↑ Guo J, Mondal M, Zhou D. Development of novel vaccine vectors: Chimpanzee adenoviral vectors. Hum Vaccines Immunother [Internet. 2018;14(7):1679–85. Available from: https://doi.org/10.1080/21645515.2017.1419108]
- ↑ Ewer K, Sebastian S, Spencer AJ, Gilbert S, Hill AVS, Lambe T. Chimpanzee adenoviral vectors as vaccines for outbreak pathogens. Hum Vaccines Immunother [Internet. 2017;13(12):3020–32. Available from: https://doi.org/10.1080/21645515.2017.1383575]
- ↑ Hassan A, Kafai N, Dmitriev I, Fox J, Smith B, Harvey I, et al. A single intranasal dose of chimpanzee adenovirus-vectored vaccine confers sterilizing immunity against SARS-CoV-2 infection. 2020;
- ↑ Hirata S, Hirai H, Nogami E, Morimura N, Udono T. Chimpanzee Down syndrome: a case study of trisomy 22 in a captive chimpanzee. Primates. 2017;58(2):267–73.
- ↑ Sundararaman SA, Plenderleith LJ, Liu W, Loy DE, Learn GH, Li Y, et al. Genomes of cryptic chimpanzee Plasmodium species reveal key evolutionary events leading to human malaria. Nat Commun. 2016;7.
- ↑ Sundararaman SA, Plenderleith LJ, Liu W, Loy DE, Learn GH, Li Y, et al. Genomes of cryptic chimpanzee Plasmodium species reveal key evolutionary events leading to human malaria. Nat Commun. 2016;7.
- ↑ Hirata S, Hirai H, Nogami E, Morimura N, Udono T. Chimpanzee Down syndrome: a case study of trisomy 22 in a captive chimpanzee. Primates. 2017;58(2):267–73.
- ↑ Hirata S, Hirai H, Nogami E, Morimura N, Udono T. Chimpanzee Down syndrome: a case study of trisomy 22 in a captive chimpanzee. Primates. 2017;58(2):267–73.
- ↑ Hirata S, Hirai H, Nogami E, Morimura N, Udono T. Chimpanzee Down syndrome: a case study of trisomy 22 in a captive chimpanzee. Primates. 2017;58(2):267–73.
- ↑ Guo J, Mondal M, Zhou D. Development of novel vaccine vectors: Chimpanzee adenoviral vectors. Hum Vaccines Immunother [Internet. 2018;14(7):1679–85. Available from: https://doi.org/10.1080/21645515.2017.1419108]
- ↑ Guo J, Mondal M, Zhou D. Development of novel vaccine vectors: Chimpanzee adenoviral vectors. Hum Vaccines Immunother [Internet. 2018;14(7):1679–85. Available from: https://doi.org/10.1080/21645515.2017.1419108]
- ↑ Ewer K, Sebastian S, Spencer AJ, Gilbert S, Hill AVS, Lambe T. Chimpanzee adenoviral vectors as vaccines for outbreak pathogens. Hum Vaccines Immunother [Internet. 2017;13(12):3020–32. Available from: https://doi.org/10.1080/21645515.2017.1383575]
- ↑ Guo J, Mondal M, Zhou D. Development of novel vaccine vectors: Chimpanzee adenoviral vectors. Hum Vaccines Immunother [Internet. 2018;14(7):1679–85. Available from: https://doi.org/10.1080/21645515.2017.1419108]
- ↑ Ewer K, Sebastian S, Spencer AJ, Gilbert S, Hill AVS, Lambe T. Chimpanzee adenoviral vectors as vaccines for outbreak pathogens. Hum Vaccines Immunother [Internet. 2017;13(12):3020–32. Available from: https://doi.org/10.1080/21645515.2017.1383575]
- ↑ Ewer K, Sebastian S, Spencer AJ, Gilbert S, Hill AVS, Lambe T. Chimpanzee adenoviral vectors as vaccines for outbreak pathogens. Hum Vaccines Immunother [Internet. 2017;13(12):3020–32. Available from: https://doi.org/10.1080/21645515.2017.1383575]
- ↑ Ewer K, Sebastian S, Spencer AJ, Gilbert S, Hill AVS, Lambe T. Chimpanzee adenoviral vectors as vaccines for outbreak pathogens. Hum Vaccines Immunother [Internet. 2017;13(12):3020–32. Available from: https://doi.org/10.1080/21645515.2017.1383575]
- ↑ Ewer K, Sebastian S, Spencer AJ, Gilbert S, Hill AVS, Lambe T. Chimpanzee adenoviral vectors as vaccines for outbreak pathogens. Hum Vaccines Immunother [Internet. 2017;13(12):3020–32. Available from: https://doi.org/10.1080/21645515.2017.1383575]
- ↑ Hassan A, Kafai N, Dmitriev I, Fox J, Smith B, Harvey I, et al. A single intranasal dose of chimpanzee adenovirus-vectored vaccine confers sterilizing immunity against SARS-CoV-2 infection. 2020;
- ↑ Guo J, Mondal M, Zhou D. Development of novel vaccine vectors: Chimpanzee adenoviral vectors. Hum Vaccines Immunother [Internet. 2018;14(7):1679–85. Available from: https://doi.org/10.1080/21645515.2017.1419108]
- ↑ Ewer K, Sebastian S, Spencer AJ, Gilbert S, Hill AVS, Lambe T. Chimpanzee adenoviral vectors as vaccines for outbreak pathogens. Hum Vaccines Immunother [Internet. 2017;13(12):3020–32. Available from: https://doi.org/10.1080/21645515.2017.1383575]
- ↑ Hassan A, Kafai N, Dmitriev I, Fox J, Smith B, Harvey I, et al. A single intranasal dose of chimpanzee adenovirus-vectored vaccine confers sterilizing immunity against SARS-CoV-2 infection. 2020;
- ↑ Guo J, Mondal M, Zhou D. Development of novel vaccine vectors: Chimpanzee adenoviral vectors. Hum Vaccines Immunother [Internet. 2018;14(7):1679–85. Available from: https://doi.org/10.1080/21645515.2017.1419108]
Edited by [Maya Schwingel], student of Joan Slonczewski for BIOL 116 Information in Living Systems, 2020, Kenyon College.
