Clostridium cellulovorans: Difference between revisions
| Line 22: | Line 22: | ||
[[File:Cellulochromosome.jpg|220px|thumb|left|Chromosome map]] | [[File:Cellulochromosome.jpg|220px|thumb|left|Chromosome map]] | ||
<Br> <Br> | <Br> <Br> | ||
Genome sequencing of ''C. cellulovorans'' has been completed. ''C. Cellulovorans'' contains circular chromosome | Genome sequencing of ''C. cellulovorans'' has been completed. ''C. Cellulovorans'' contains a circular chromosome with a length of 5,262,222 base pairs which is about 1 Mbp larger than the genomes from other cellulosomal clostridia. 31% of the genome is GC and 69% is AT. 57 cellulosomal genes were reported in ''C. cellulovorans''. ''C. cellulovorans'' contains large number of genes encoding non-cellulosomal enzymes which are more associated with polysaccharide (such as hemicelluloses and pectins) degradations other than cellulose. Scientists have found two novel genes encoding scaffolding proteins in ''C. cellulovorans'' genome <Br> <Br> <Br> <Br> <Br><Br> <Br> | ||
==Cell Structure, Metabolism and Life Cycle== | ==Cell Structure, Metabolism and Life Cycle== | ||
Revision as of 18:54, 20 April 2011
Classification
• Kingdom - Bacteria
• Phylum - Firmicutes
• Class - Clostridia
• Order - Clostridiales
• Family - Clostridiaceae
• Genus - Clostridium
Species
Clostridium cellulovorans
Other name: Clostridium cellulovorans strain 743B
Description and Significance
Clostridium cellulovorans (ATCC 35296) is anaerobic, spore forming and stain gram negative non-motile rods originally isolated from a batch methanogenic fermentation of hybrid poplar wood. C. cellulovorans is a mesophilic bacterium with optimum growth temperature of 37°C, though it can grow in a temperature range of 20 to 40°C. Optimum pH is 7.0, and the pH range of growth is 6.4 to 7.8. This organism produces extracellular enzyme complex known as cellulosome which can degrade plant cell walls, notably cellulose. As most abundantly available potential source of fermentable sugars in the world are the cell walls in higher plants, utilization of such a vast resource for energy production would reduce the dependency on non-renewable fossil fuel. Hence, C. cellulovorans have potential industrial application for energy production.
Genome Structure
Genome sequencing of C. cellulovorans has been completed. C. Cellulovorans contains a circular chromosome with a length of 5,262,222 base pairs which is about 1 Mbp larger than the genomes from other cellulosomal clostridia. 31% of the genome is GC and 69% is AT. 57 cellulosomal genes were reported in C. cellulovorans. C. cellulovorans contains large number of genes encoding non-cellulosomal enzymes which are more associated with polysaccharide (such as hemicelluloses and pectins) degradations other than cellulose. Scientists have found two novel genes encoding scaffolding proteins in C. cellulovorans genome
Cell Structure, Metabolism and Life Cycle
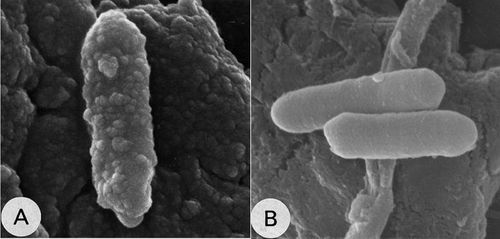
C. cellulovorans do not reduce sulfate and are obligate anaerobes. Cells are 0.7 to 0.9 by 2.5 to 3.5 µm in size and are non-motile rods, though peritrichous flagella were detected under electron microscopy. Both spores and vegetative colonies of C. cellulovorans are irregular, containing opaque edge and a center devoid. Spores are oblong that occur either centrally or subterminally within the mature sporangium. It produces plant cell wall degrading extracellular multienzyme complex called cellulosome. When grown in cellulose, C. cellulovorans forms ultrastructural protuberances, which may be aggregation of smaller cellulosome complexes, also known as polycellulosomes. These protuberances were detected only in cellulose-grown cells and disappeared rapidly when other soluble carbohydrates were added to the growth medium.
Cellulosomal components synergistically interact to catalyze the degradation of cellulose and hence, cellulosome acts as a macromolecular machine. Apart from cellulose, C. cellulovorans ferments various carbon sources, such as xylan, pectin, cellobiose, glucose, fructose, galactose, sucrose, lactose and mannose and the fermentation products are hydrogen, carbon dioxide, acetate, butyrate, formate and lactate.
Ecology and Pathogenesis
Clostridium cellulovorans is non pathogenic to human beings.
References
Author
Page authored by Umesh Adhikari and Joe Araiz, student of Prof. Jay Lennon at Michigan State University.