Clostridium perfringens
A Microbial Biorealm page on the genus Clostridium perfringens
Classification
Higher order taxa
Bacteria (Domain), Firmicutes (Phylum), Clostridia (Class), Clostridiales (Order), Clostridiaceae (Family), Clostridium (Genus), C. perfringens (Species) [14]
Species
|
NCBI: Taxonomy |
Clostridium perfringens
Description and significance
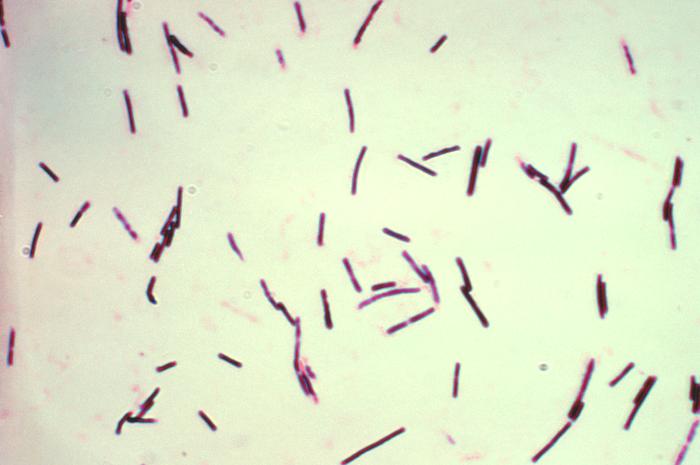
Clostridium perfringens is a rod-shaped Gram-positive bacterium, which is a mesophile that has an optimal growing temperature of 37 C [2]. It is a non-motile pathogen that produces endospores. This bacterium produces energy via anaerobic respiration using compounds other than oxygen, such as Nitrate, as its final electron acceptor [12]. Although C. perfringens is an inhabitant of human normal intestinal flora, it is a pathogen responsible for many gastrointestinal illnesses with severity ranging from mild enterotoxaemia to fatal gas gangrene [3]. This bacterium is important, because it was the primary pathogenic agent that caused many injured soldiers to die from gas gangrene during World War I. When the soldiers were injured, their first line of defense—skin—could no longer prevent the vegetative spores of C. perfringens from entering into the body through an injury and destroying the host tissues with the toxins that they made [4]. C. perfringens was isolated from fecal specimens from human and animals. Samples of the specimens were then classified accordingly by the type of enterotoxin (CPE) they produced with the help of a multiplex PCR assay [13]. After classifying them into specific serotypes, the fecal isolates were further tested using pulsed-field gel electrophoresis and restriction fragment length polymorphism to determine which CPE gene, plasmid CPE gene or chromosomal CPE gene, they contained [13]. C. perfringens is similar to another gastrointestinal-disease-causing pathogen, C. difficile. Therefore, it is very beneficial to have its genome sequenced, so that fast detection and quantification of these two species can be achieved.
Genome structure
Clostridium perfringens has a single circular chromosome made up of approximately 3.6 million base pairs, with a GC content ranging from 24 to 55% [3]. C. perfringens has a relative low GC content comparing to that of the majority of Gram-positive bacteria. The chromosome contains 10 rRNA genes and 96 tRNA genes [4]. The genome contains genes that encode a variety of transporters that transport “amino acids, cations/anions, carbohydrates, and nucleosides/nucleotides” [4]. Like many Mycoplasma bacteria and Bacillus subtilis, C. perfringens arrange their genes in a way such that their transcriptional process orients in the same direction as their replication direction[4].
Cell structure and metabolism
Clostridium perfringens are non-motile rod-shaped Gram-positive bacteria. It possesses the typical characteristics of Gram-positive bacteria, such as a protective thick cell wall, which is made up of peptidoglycan, surrounding an inner membrane. C. perfringens is an anaerobic bacterium, who acquires energy by performing anaerobic respiration using Nitrate as its electron acceptor. There is an increase in growth when this bacterium is grown in the presence of Nitrate, because this inorganic acceptor allows more metabolites molecules to undergo substrate-level phosphorylation reactions, leading to an increase yield in energy production [12]. C. perfringens can also undergo anaerobic fermentation to produce gases, such as carbon dioxide, that may increase its survival by creating a preferred anaerobic habitat in host tissues [4]. C. perfringens also has all the enzymes necessary to carry out glycolysis and glycogen metabolism [4]. Sugar compounds are broken down into simpler forms using various glycolytic enzymes encoded by genes located in the genome [4]. However, C. perfringens does not have a complete set of genes necessary for amino acid biosynthesis; in fact, only 45 enzyme-encoding genes were discovered [4]. Therefore, C. perfringens cannot survive on media that are lacking an essential amino acid supply [4].
Ecology and Pathology
C. perfringens is categorized into five serotypes—A, B, C, D, and E—depending on the types of extracellular toxins (alpha-, beta, epsilon-, and iota-toxins) they make and their forms of tropisms [8]. C. perfringens is a pathogen whose primary targets are human and animals. The bacterium can be found in many different habitats, such as the normal flora of human gastrointestinal (GI) tract, and environment, such as sewage and soil [4]. Several common diseases associated with C. perfringens are food-poisoning, gas gangrene, and many veterinary diseases. C. perfringens enterotoxin (CPE) is the main virulent factor that initiates many critical GI diseases. When food contaminated with C. perfringens is consumed, CPE begins its membrane action in a unique four-step mechanism. The first step of the mechanism is the binding CPE to the target receptor on plasma membrane protein or claudin proteins, which leads to the formation of a small complex [5]. The complex then undergoes physical change when it binds to other membrane proteins and forms a larger complex in the membranes, which results in the disruption of the membrane’s permeability [5]. This usually leads to cell death, because the osmotic equilibrium is not maintained due to the breakdown of the membrane’s permeability [5]. CPE is capable of forming a larger complex in the membrane and its toxic level is greatly enhanced when the first 45 N-terminal amino acids are eliminated [5]. On the other hand, eliminating amino acids outside of residue 45 prohibits CPE from forming large complex, and thus loses its toxicity [5].
Using pulsed-field gel electrophoresis, researchers found that GI diseases can also be non-food borne when the chromosomal CPE is carried by an episome. Antibiotic-associated diarrhea (AAD) is an example of non-food borne GI illness caused by plasmid CPE [13].
Application to Biotechnology
C. perfringens is a mesophile who has a fastest growth rate at the temperature of 37 C [2]. When the spores of C. perfringens are exposed to a hyperthermal condition with a temperature of 105C for five minutes, their sensitivity to polymyxin and neomycin is increased significantly [6]. Polymyxin is an antibiotic that damages the structure, the permeability and the rigidity of the bacterial plasma membrane, making the cell membrane too fragile to maintain the osmotic equilibrium [6]. Neomycin is a metabolic inhibitor that stops protein synthesis [6]. Therefore, polymyxin and neomycin can be used together to kill C. perfringens cells after exposing them to super high temperature for a short amount of time. This discovery is beneficial to human and animals, because the combination of polymyxin and neomycin can be used to control C. perfringens, which presents a threat as a toxic causative agent of many food-borne diseases. The results also implied that refrigerated food should be heated properly before eating them, because it can reduce the risk of getting C. perfringens-related illnesses.
Another beneficial biotechnological application discovered by researchers focused on the relationship between C. perfringens’ penicillin-binding proteins and the beta-lactam antibiotics. A physical map of C. perfringens was assembled using pulsed-field gel electrophoresis (PFGE), which made the genetic study of C. perfringens and its virulence factors more feasible. Using the physical map, researchers have identified 6 penicillin-binding proteins (PBPs) in the C. perfringens’ membranes [7]. Like many aerobic bacteria, C. perfringens have many PBPs in their cytoplasmic membrane [7]. Some of these proteins play important roles in murein synthesis, cell elongation, and maintenance of cell rigidity [7]. When beta-lactam antibiotics are induced to the cells, “PBP 3 and 4 were saturated by beta-lactam at their MICs…and caused cell death at concentrations which resulted in no detectable binding to PBP 5 and 6” [7]. PBP 3 and 4 are higher-molecular-weight PBPs comparing to the molecular weight of the other PBP 5 and 6. Due to their important roles in the growth of the bacterial cells, many beta-lactam antibiotics were designed to recognize PBPs as their targets.
Current Research
1) A current research conducted by International Association for Food Protection examined the inhibitory effects green tea extracts had on the germination of C. perfringens spores in abusive-chilled retail cooked meat [9]. The experiment was carried out by treating samples of thawed pork, beef and chicken with different concentration levels of green tea extracts along with “heat activated (75 degrees C for 20 min) three-strain spore cocktail” [9]. Then these samples were incubated in a 71 C water bath for an hour [9]. Then the temperature of the samples was allowed to drop from 54.4 to 7.2 C [9]. The first round of experiment was tested without adding any green tea extracts, and germination of C. perfringens population thrived. The second round of experiment was carried out with the addition of GTL, which was a green tea extract with low level of catechin, but inhibition of C. perfringens growth was not observed [9]. However, when GTE, which was a green tea extract containing high level of catechin, was added to the meat samples, germination of C. perfringens population was inhibited [9]. The results of this research were significant, because the uptake of catechin could decrease the chance of C. perfringens spores from germinating during abusive cooling. Thus the risk of human or animals being affected by ingested C. perfringens spores is reduced.
2) A recent study conducted by Departments of Medicine, Microbiology and Immunology at Vanderbilt University School of Medicine focused on the inhibiting activities of two monoclonal neutralizing antibodies against the epsilon-toxin produced by C. perfringens, which could cause human to suffer from edema of heart, kidney and brain [10]. According to the study, when cultured MDCK cells were stained with membrane-impermeant dye 7-aminoactinomycin D, it showed that both antibodies were effective in inhibiting the epsilon-toxin from forming spores in the MDCK cell membranes [10]. Both antibodies were capable of recognizing epitopes in the region containing amino acids 134 to 145, which was a region that “overlaps an amphipathic loop corresponding to the putative membrane insertion domain of the toxin” [10]. This is an important finding, because once the epitopes recognized by these antibodies are identified, it will allow scientists to develop therapeutic means to act against the detrimental effects of the toxin.
3) Although C. perfringens is a pathogen responsible for many cases of avian enteritis, there was insufficient knowledge on which toxins are the primary causative agent of these avian diseases. A current study done by California Animal Health and Food Safety Laboratory System at University of California Davis compared the toxinotypes of C. perfringens isolates obtained from infected birds, including quail, chickens, turkey and psittacines, against the toxinotypes of 19 isolates obtained from birds that were not infected by C. perfringens [11]. The results showed that the toxinotype of all C. perfringens isolates obtained from both the diseased and the healthy birds belonged to type A, which is a type of C. perfringens that are capable of producing alpha-toxins [11]. Many of the isolates contained the gene that encodes the beta2 toxin, but about half of the isolates failed to make the beta 2 toxin [11]. The significance of this result is that avian enteritis is not associated with the specific C. perfringens toxinotypes [11].
References
(2) “Clostridium perfringens CPE str. F4969 project at J. Craig Venter Institute.
(14) National Center for Biotechnology Information.
Edited by Kuang-Hsin Lee, student of Rachel Larsen at UCSD.
KMG
