Clostridium thermocellum
A Microbial Biorealm page on the genus Clostridium thermocellum
Classification
Higher order taxa
Bacteria; Firmicutes; Clostridia; Clostridiales; Clostridiaceae; Clostridium
Species
Clostridium thermocellum
|
NCBI: Taxonomy |
Description and significance
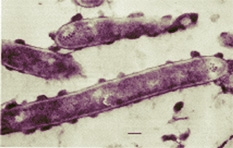
Like most species from the Clostridium genus, C. thermocellum is a bacteria that has a rod-like shape for its cell body. It is classified as a gram-positive bacteria which means that the cell body is only surrounded by a single bilayer lipid membrane. Because it is a gram-positive bacterium, the outside of the cell membrane also contains a thick cell wall known as murein, which is made of peptidoglycans. C. thermocellum is an anaerobic and thermophilic organism that produces spores. In addition, it has cellular structures and mechanism that give it motility in the environment it resides in. The organism was completely isolated and sequenced at the DOE Joint Genome Institute. The bacterium had its genome completely sequenced because it contains a unique extracellular enzyme system capable of breaking down insoluble cellulose into ethanol which is vital for biomass energy. Because of its ability to break down cellulose, it is highly likely that C. thermocellum can be isolated from many plant organisms. In addition, it can be isolated from cow and horse manure because it is required in the animals' digestive system to break down the cellulose of grass.
Genome structure
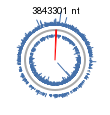
The number of nucleotides present in the genome of C. thermocellum has been discovered and reported to be at 3,843,301 base pairs which makes up 3307 genes [1].
The nucleotides make up one double-stranded circular DNA. The circularity of the chromosome is advantageous for C. thermocellum because exonucleases cannot digest the ends of its DNA. Another advantageous feature of the circularity of the chromosome is that the chromosome can be supercoiled which drastically lowers the energy barrier making it easier for DNA to be activated and separated into single strands for transcription into RNA. In regards to genomic organization, C. thermocellum like all other prokaryotic organisms lack a true nucleus for DNA storage and transcription. Instead, the entire genome is tightly packaged in a small region called the nucleoid and transcription takes place between the cytoplasm and nucleoid. In addition to the circular chromosome, C. thermocellum also has a plasmid which is an extrachromosomal genetic element that is not essential in affecting its lifestyle. Unlike the circular chromosome which is necessary for viability, the loss of the plasmid will not affect the bacteria's ability to reproduce. Because C. thermocellum is known as a degrader of cellulose, its DNA contains specific nucleotide sequences that make up the genes that encode for the system of enzymes that are necessary for cellulose degradation.
Cell structure and metabolism
The general interior cellular structure of C. thermocellum is similar to most rod-shaped bacteria. It contains a nucleoid region for packaging and transcription of the DNA into RNA. A nuclear envelope encompasses the nucleoid. In the cytoplasm, a chain or cluster of ribosomes form polyribosomes which function as a continuous protein factory that translates mRNA into polypeptide chains.
Aside from the general internal framework of the cell, C. thermocellum is a gram-positive bacteria. As a result, the exterior cellular structure of the bacteria consists of a thick cell wall that is composed of peptidoglycan, a complex polymer of sugars and amino acids. The type of peptidoglycan that forms the cell wall of C. thermocellum is known as murein which protects its single bilayer membrane from high turgor pressure and also provides the cell its shape and rigidity.
The most interesting and special external cellular feature for this bacteria however stems from its unique and trademark ability to degrade insoluble cellulose. The cellulosome, a complex system of cellulolytic enzymes, is located at the exterior of the C. thermocellum cell. The cellulosome contains nearly 20 catalytic enzymes that are encoded by over 100 genes. Some of the genes play a role in synthesizing the enzymes while others are responsible for regulating cellulosome activity with inducers and repressors. The cellulases are each responsible in the cellulose breakdown process. This also suggests that the cellulose degrading bacteria has extracellular appendages. In order to perform cellulolysis, C. thermocellum must adhere to the surfaces of plants [2]. To be able to stick onto the surfaces of plants suggests that the bacteria have pili, which are structures that provide it attachment and motility capabilities. The pili are like grappling hooks which retract and attach onto the surfaces of plants.
In terms of energy production, since C. thermocellum is an anaerobic organism, it obtains most of its energy through the metabolic process of fermentation. In fermentation, the bacterial cell does not consume oxygen to produce energy because there is a lack of good electron acceptors. Rather, the bacteria utilizes smaller sugar molecules such as glucose obtained from cellulose breakdown of plants to obtain energy during fermentation. During fermentation, waste products are generated such as hydrogen, carbon dioxide, acetate, and primarily ethanol [3].
Ecology
Describe any interactions with other organisms (included eukaryotes), contributions to the environment, effect on environment, etc.
Pathology
Most bacterial species in the Clostridium genus generate toxins that are harmful and malevolent to other host cells. For instance, one of the most deadly toxins is produced by Clostridium botulinum which causes paralysis by stopping all motor and neural activities. Paralysis usually occurs from the head, then affects the upper body, and then gradually affects the lower body of humans because the cranial nerves are affected first by the toxins. In the worst case scenario, the respiratory muscles are infected by the toxins of C. botulinum leading to the inability to breathe and finally death. Infection by C. botulinum usually occurs from overdose of Botox injection leading to toxification of the human body or from ingestion of bad, spoiled food products [4]. The other prominent Clostridium species that exhibits pathogenic effects is Clostridium tetani. C. tetani primarily infect young children by producing toxins in their opened, unclean wounds. The production of the neurotoxins leads to the disease called tetanus which is usually causes stiffening of the muscles, swelling, sweating, and muscle spasms [5]. Unlike its bacterial relatives, C. thermocellum does not produce or secrete any kind of toxins that will induce any pathological effects on human, animal, or plant hosts.
Application to Biotechnology
C. thermocellum is best known for its ability to degrade cellulose which is supposedly an extremely insoluble compound. However, in order for cellulose degradation to occur, C. thermocellum produces many enzymatic proteins that are particularly vital for cellulolysis. Biotechnological research has shown that the cellulose degrading bacteria produces a large, complex cellulase system known as the cellulosome which consists about 20 catalytic proteins that are involved in the bacteria's adherence to cellulose, breakdown and regulation of cellulose degradation, and the transport of sugar monomers [6]. One of the most important proteins of the cellulosome is the CipA which is a large, non-catalytic 250 kDa scaffold protein. As a scaffold protein, CipA is a large protein that has multiple specific binding sites which serves to recruit other smaller proteins together which share the same signaling pathway. When brought together by CipA, the proteins can interact to signal and ultimately trigger cellulose degradation. In this case, CipA has nine cohesin domains for protein binding and is mediated by the dockerin domains on the catalytic proteins of the cellulosome. In addition to activation of cellulosome degradation CipA also contains cellulose binding factors which are absolutely essential for cellulolysis to even occur in the first place. The cellulose binding factors allow C. thermocellum to adhere onto the surface of organisms containing cellulose so that the insoluble substrate can be degraded. Aside from the scaffold protein in the cellulosome, C. thermocellum also produces glycosyl hydrolases which function as the catalytic subunits that engage in the actual cleavage of cellulose [7].
Current Research
Because of its ability to degrade cellulose, C. thermocellum has been highly scrutinized and researched by the researchers in the field of microbiology. In a recent journal article on C. thermocellum by Williams, Combs, Lynn, and Strobel in, the research on the bacteria addressed the inhibitory effects of cellular growth and ethanol production through fermentation of C. thermocellum. Apparently in the ethanol adapted strain, ethanol production was low or inhibited when the concentration of ethanol was higher than one percent. This prompted the proteomic analysis which is basically the study of the proteins involved in cellulose degradation. According to the analysis, it showed that the membrane proteins were the cause of the low levels of ethanol production from fermentation. When ethanol concentration hit a particular level, the majority of the membrane proteins were down-regulated in the ethanol-adapted strain of C.thermocellum. By down-regulating the membrane associated proteins, synthesis of proteins involved in degradation and fermentation were slowed down. This negatively affects ethanol production because there are not enough membrane-associated proteins for carbohydrate transport across the membrane and metabolism. Transport of carbohydrates is necessary because smaller sugars monomers that are broken down from cellulose occur extracellularly and need to be imported into the baterial cell. Also, by down-regulating the level of membrane-associated proteins, metabolic processes such as fermentation cannot occur which is vital for breaking down cellulose. Research is being conducted specifically on the membrane-associated proteins of C. thermocellum to understand their functions and structures so that an enhanced cell can be designed to produce high levels of ethanol. With high levels of ethanol production, there will be a large source of biomass energy that can be obtained through C. thermocellum.
Recent research on C. thermocellum has also involved the analysis of the fermentative products generated from cellulose breakdown. In one particular research done by Sparling, Islam, and a number of other researchers, C. thermocellum 27405 is cultured in cellubiose and alpha cellulose. According to analysis of a certain strain of C. thermocellum, formate production was detected. H However, when C. thermocellum 27405 was analyzed in various conditions of growth, formate presence was not confirmed. Therefore, polymerase chain reaction and BLAST were done on C. thermocellum 27405 to confirm that there indeed is formate synthesis during fermentation. It was discovered that the pfl, fnr, and adhE genes were responsible for encoding pyruvate-formate lyase (PFL), PFL-activating enzyme, and alcohol dehydrogenase E enzyme which play a role in formate synthesis. However, the production of formate results in the lower production of other fermentative products such as hydrogen, carbon dioxide, and ethanol because it tries to outcompete their synthesis. To obtain more industrially applicable products such as hydrogen and ethanol which can be a source for fuel and energy, this research is conducted to learn how to counter and ultimately lower formate synthesis by targeting and modifying the genes or enzymes that contribute its formation.
Extensive scienfic work on C. thermocellum has also involved microbiologists doing research on the genes that control cellulose degradation. A current specific research on cellulose degradation involved the analysis of the cellulase system, cellulosome. According to the scientific studie and experiment, it is shown that over 100 genes are involved in encoding proteins involved in cellulose degradation. The research on C. thermocellum is taken a step further by determining how all the genes involved in the cellulose degradation are regulated by the bacteria. It has been determined that the gene glyR3 is cotranscibed on the same operon with the celC and licA genes and acts as a regulator for encoding cellulase [7]. These three genes make up the celC operon. When the protein repressor GlyR3 was bound to the promoter region of the celC operon, the expression of the genes to encode for cellulase was repressed. However, it was discovered in this experiment that when laminaribiose binds to the GlyR3 repressor, it resulted in the release of the repressor from the promoter thus allowing the glyR3, celC, and licA genes on the operon to be expressed. This research and others related to it is essential for the future development of conversion of biomass into energy because by understanding which genes encode for cellulose degrading proteins and how their expression are regulated and induced, cellulose degrading ability of C. thermocellum can be manipulated and amplified as a mass energy source.
References
1. National Center for Biotechnology Information.
2. Bayer, E., Kenig, R., and Lamed, R. "Adherence of Clostridium thermocellum to Cellulose". Journal of Bacteriology. 1983. Volume 156. p. 818-827.
3. Weimer, P. and Zeikus, J. "Fermentation of Cellulose and Cellobiose by Clostridium thermocellum in the Absence and Presence of Methanobacterium thermoautotrophicum". Applied and Environmental Microbiology. 1977. Volume 33. p. 289-297.
4. Nantel, Albert. "Clostridium thermocellum. International Programme on Chemical Safety Poisons Information Monograph 858 Bacteria". World Health Organization. 1999. p. 1-32.
5. "Clostridium tetani". World Health Organization. 2007.
6. JGI. Clostridium thermocellum ATCC 27405.
7. Newcomb, M., Chen, C., and Wu, J.H. "Induction of the celC operon of Clostridium thermocellum by laminaribiose". Proceedings of the National Academy of Sciences of the United States of America. 2007. Volume 104. p. 3747-3752.
8. Williams, T., Combs, J., Lynn, B., and Strobel, H. [ "Proteomic profile changes in membranes of ethanol-tolerant Clostridium thermocellum". Applied and Environmental Microbiology.] 2006. Volume 74. p. 422-432.
9. Sparling, R., Islam, R., Cicek, N., Carere, C., Chow, H., and Levin, D. ["Formate synthesis by Clostridium thermocellum during anaerobic fermentation". National Research Council Canada.] 2006. Volume 52. p. 681-688.
Schaechter, M., Ingraham, J., and Neidhardt, F. Microbe. American Society for Microbiology. 2006. Chapter 2. p. 23-24, Chapter 3. p. 39-42.
Edited by Kenny Lam