Corynebacterium diphtheriae
A Microbial Biorealm page on the genus Corynebacterium diphtheriae
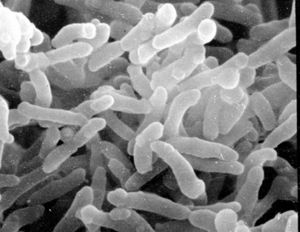
Classification
Higher order taxa
Bacteria; Actinobacteria; Actinomycetales; Corynebacteriaceae; Corynebacterium
Species
|
NCBI: Taxonomy |
Corynebacterium diphtheriae
Description and significance
C. diphtheriae is a Gram-positive, aerobic, nonmotile, toxin-producing, rod-shaped bacteria belonging to the order Actinomycetales, which are typically found in soil, but also have pathogenic members such as streptomyces and mycobacteria. C. diphtheriae is best known for causing the disease Diphtheria in human beings, which results from production of Diphtheria toxin in conjunction with infection by a bacteriophage which provides it with the toxin-producing gene. Because historically it has been a very deadly disease, especially for children where mortality rates before vaccines and antitoxin amounted to nearly 80%, it has been heavily studied. Oddly enough, mice and other rodents are naturally immune to the Diphtheria toxin so it has been difficult to study Diphtheria in the lab. [1]
Diseases resembling Diphtheria were described as early as the 4th Century BCE by Hippocrates. It was named in 1826 by French physician Pierre Bretonneau, who gave it the name Diphtheria from the Greek word which means "leather hide". It was named "leather hide" because of the leathery layer that grows on the tonsils, throat, and nasal passages. C. diphtheriae was first described in 1884 by Edwin Klebs and Friedrich Loffler, giving it the name Klebs-Loffler bacillus by which it used to be known. [4, 8]
In 1888, Pierre-Paul-Émile Roux discovered that a toxin produced by C. diphtheriae was the causative agent in the disease Diphtheria. Emil Von Behring (1854-1917) was one of the first pioneers in the race to fight Diphtheria. He studied C. diphtheriae from 1889 onwards, especially in animals. When he discovered that the serum of anthrax-resistant rats was able to kill the anthrax bacillus, he researched such possibilities for Diphtheria, eventually using the serum of Diphtheria toxin-resistant guinea pigs, and ultimately horses. Eventually he developed antiserum, based on his experiments and theories developed by a previous colleague Shibasaburo Kitasato. By 1892, the antiserum was ready for commercial production. In 1891, Behring was awarded the Nobel Prize in Medicine for his work. [5, 8]
In 1923, the work of Alexander Glenny, Barbara Hopkins, and Gaston Ramon produced a vaccine for Diphtheria. Glenny and Hopkins found that formaldehyde could prevent the Diphtheria toxin from being toxic, while Ramon discovered that the nontoxic form still retained its antigenic qualities. The form of the toxin which results from denaturing with formaldehyde is called an anatoxin, "toxoids", or "formalin-toxoids", and is used in vaccinations worldwide. [8]
Outbreaks of Diptheria were common earlier in the 18th and 19th century, but the introduction of the vaccine caused a decline in the number of cases. One of the more well-known outbreaks of Diphtheria was commemorated in the animated film "Balto", a story about the dog sled team (whose lead dog was named Balto) which brought the Diphtheria antiserum from Nenana to Nome, which was geographically isolated during the epidemic. They traveled 700 miles in 6 days through temperatures of minus 40 degrees Celsius and blizzard-speed winds. The dog sled team saved hundreds of lives, the story was sensationalized worldwide, and a statue of Balto, the lead dog, was erected in New York's Central Park. [7]
Despite having a vaccine and antiserum, there is currently an increase in the number of outbreaks in undeveloped or underdeveloped countries, and especially Eastern European countries where loss of infrastructure has led to decreases in vaccinations. Research is being done to determine why new epidemics are being seen - whether it is a result of a mutation in C. diphtheriae, or if it is from environmental and social factors such as poor immunization rates and unsanitary conditions.
Genome structure
The genome of C. diphtheriae contains 2,488,635 nucleotides, 87% of which are coding. This single circular chromosome contains 2,389 genes from which 2,272 proteins are coded. It contains no plasmids. There are 69 structural RNA genes, 15 coding for 5S, 23S, and 16S Ribosomal RNA, and 54 coding for the 20 amino acid tRNA molecules. The GC content is 53%, though it is not evenly distributed - it has higher proportion of GC pairs near the origin of the plasmid (around 55%), compared to the region near the terminus (49%) [1]
The genome contains what are termed Pathogenicity Islands (PAIs). These PAIs, of which C. diphtheriae has 13, contain local genetic anomalies in nucleotide composition or frequency, and many are closely associated with patches that code for tRNA. These PAIs code for the majority of fimbrial and fimbria-related genes, and iron-uptake systems which produce siderophores and lantibiotics. The ability to acquire iron allows C. diphtheriae to produce the Diphtheria toxin, which gives the bacteria its pathogenicity. [1]
Various methods were used to sequence the genome. The original strain of C. diphtheriae was obtained from the pharyngeal membrane of a 72-year-old female in 1997. The organism was then cultured and the DNA chromosome extracted. The genome sequence was obtained from 60,750 end sequences (with 9.2x coverage) derived from two pUC18 genomic shotgun libraries. All repeats were bridged using read-pairs or PCR, after which the genome was assembled. Computer analysis was used to identify possible genes and coding segments. [1]
Cell structure and metabolism
Cell Exterior
Being Gram-positive, C. diphtheriae has a cell membrane and a lipid-rich murein layer outside the membrane. Like M. tuberculosis, another Actinomycete, C. diphtheriae has an arabinogalactan polymer which anchors a lipid rich domain to the murein layer. Corynomycolic acids, alpha-alkyl beta-hydroxy fatty acids, are produced through a Claisen-like condensation reaction between two fatty acyl chains. [1]
Also in the exterior are the fimbriae (pili). Not much structural analysis has been done, however it is very likely that sortase-like proteins are produced to aid in anchoring the fimbriae and possibly polymerizing them. C. diphtheriae has six genes which code for putative sortases - five are localized on PAIs, none of which are found in closely related non-pathogenic corynebacteria (C. glutamicum and C. efficens). The absence of these genes in the non-pathogenic relatives suggests that those genes increase the pathogenicity of C. diphtheriae through the fimbriae, most likely by aiding the bacterium invade and adhere to host cells. The similarity to the sortase-related fimbrial system of Actinomyces, and the newness of the genes in the C. diphtheria genome, suggest that these genes were newly acquired, possibly the cause of the recent upsurge in outbreaks of Diphtheria. [1]
Metabolic processes
Pathways discovered for metabolism include the complete pentose phosphate pathway, glycolysis, and gluconeogenesis, based on the enzymes present in the cell. The citric acid cycle is present, however C. diphtheriae lacks succinyl-CoA synthetase and uses an enzyme transcribed by a homologue of the Clostridium kluyveri cat1 gene, which acts as a coenzyme A transferase rather than a synthetase. De novo amino acid biosynthesis is also present in C. diphtheriae, as well as both anaerobic and aerobic respiration pathways, though C. diphtheriae is primarily aerobic. Production of purines is complete, however of the pyrimidines, the cytidine production pathway lacks the final cytidine triphosphate synthetase, which is present in other related organisms such as C. efficens and M. tuberculosis. [1]
Specialized Iron Intake and Diphtheria Toxin
Diphtheria toxin is the most significant molecule produced by C. diphtheriae. The tox gene which codes for the Diphtheria toxin is located on the right-handed end of an integrated corynephage. C. diptheriae contains the protein DtxR, an iron-dependent negative-regulatory protein which prevents the tox gene from being transcribed under high iron conditions, as well as the high-energy iron uptake system of the bacterium. Because a host often reacts to infection by lowering iron levels, the bacteria have developed a mechanism which couples Diphtheria toxin gene expression with low iron levels. [1]
To import iron, C. diphtheriae has seven different systems, two of which have been identified. The siderophore system creates ferrisiderophores, molecules specialized in binding to iron and bringing it into the cell for usage. [1]
Ecology
C. diphtheriae is only found in the mouth, throat, nose, skin, bodily secretions, and wounds of infected persons. Animals do not easily contract Diphtheria from human beings, and naturally have immunity.
Pathology
C. diphtheriae primarily causes the disease diphtheria in human beings, and is prevalent in Eastern European countries, Latin America, the Caribbean, and undeveloped countries. Widespread vaccinations have nearly eradicated the disease in the United States. Diphtheria is very contagious and is spread through direct contact and aerosolized expulsions. C. diphtheriae appears in three substrains of differing generation times - gravis (60 minutes), intermedius (100 minutes), and mitis (180 minutes). The faster the strain grows, then the faster the iron is depleted, and the sooner the toxic effects of Diphtheria occur. Diptheria as a disease presents itself in two forms: respiratory and cutaneous.[3,11]
Cutaneous diphtheria can be caused by both the toxigenic and the nontoxigenic strains of C. diphtheriae. Cutaneous diphtheria presents as nondescript sores or shallow ulcers on the skin, and is usually mild and treatable. Only 1-2% of cutaneous cases become toxigenic.[3]
Respiratory diphtheria is caused by C. diphtheriae bacteria adhering to and colonizing the tonsils, nasal cavity, and throat. Mechanisms of adherence are unknown, but recent research suggests that proteins SpaB and SpaC found on two minor pili play an important role in specifically binding to pharyngeal cells.[6] A visible, leathery pseudomembrane forms over the surface of the cells, especially the tonsils and throat. A lesion develops and plasma from injured epithelial cells leaks into the lesion, producing a fibrin network infused with C. diphtheriae.[11] Symptoms present themselves 2-5 days after infection with C. diphtheriae. Initially, symptoms include a sore throat and low fever. More severe symptoms include inflammation of the neck and difficulty breathing or asphyxiation.[3]
Diphtheria infections are further complicated by the toxigenicity factor. If one is infected with a toxigenic strain of C. diphtheriae, two factors must be present to cause the production of the Diphtheria toxin:
1. Low extracellular concentrations of inorganic iron. In the presence of iron, a repressor molecule, coded for by the diphtheria toxin repressor (DtxR) gene, is activated by iron and prevents transcription of the tox gene. When concentrations of iron drop, the repressor molecule is inactivated and transcription of the tox gene proceeds, resulting in production of diphtheria toxin.[11]
2. Presence in the bacterial chromosome of a lysogenic prophage which contains the tox gene. The bacterium originally receives the tox gene from a specific Beta-phage.[11]
Diphtheria Toxin is a bacterial exotoxin which consists of an active (A) domain, and a binding (B) domain. The toxin acts by binding to the HB-EGF receptor on cells and enters by receptor-mediated endocytosis. The acidic environment of the endosome causes the toxin to undergo a conformational change which causes part of the A domain to enter the endosomal membrane. The A domain is then cleaved from the B domain and passes through the membrane into the cell cytoplasm. The A domain then catalyzes the transfer of ADP-ribose from NAD to Elongation Factor II (EF2), a key protein in protein synthesis during translation. With the addition of ADP-ribose to EF2, EF2 is deactivated. Once a significant amount of EF2 proteins have been deactivated, the cell death occurs.[11]
Diagnosis and Treatment
Clinically, the disease is described as: "An upper-respiratory tract illness characterized by sore throat, low-grade fever, and an adherent membrane of the tonsils, pharynx, and/or nose" [3]
If diphtheria is suspected, then a sample of the bacteria is isolated from the patient and cultured, and toxigenicity testing is performed. If the bacteria is C. diphtheriae, the substrain is further identified - intermedius, mitis, and gravis. To test for toxigenicity, the Elek test is performed. The CDC can also perform a PCR test, but it is not typically used when forming a diagnosis. Another test is serologic testing, which determines the level of antibodies to the diphtheria toxin. [3]
Diphtheria is treated with diphtheria antitoxin, and a 14-day course of antibiotics, preferably Erythromycin or Penicillin.
Application to Biotechnology
One major compound produced is diphtheria toxin, which has been shown to be an anti-tumor agent and has promise as a treatment of various cancers, depending on the diphtheria toxin immunity of the patient. Effects of the toxicity on patients who received treatment include "transient peripheral neuropathy and systematic delayed reaction to diphtheria toxin." [13]
C. diphtheriae is useful for studying iron-uptake systems, which is a key metabolic process in many pathogenic bacteria. Also, it is an excellent example of the relationship between bacteria and bacteriophages, and was one of the first bacteria for which an antitoxin treatment was developed.
Current Research
Corynebacterium diphtheriae employs specific minor pilins to target human pharyngeal epithelial cells, April 2007
Research conducted by Anjali Mandlik, Arlene Swierczynski, Asis Das and Hung Ton-That at the University of Connecticut Health Center has shed light on how C. diphtheriae attaches to pharyngeal epithelial cells in a host. They studied the three pili on the surface of C. diphtheriae: SpaA (major), SpaB (minor), and SpaC (minor). Deletion of the spaA gene, which codes for the major pilin SpaA, had little effect on the attachment ability of C. diphtheriae. Deletion of the spaB or spaC genes, which code for the minor pili SpaB and SpaC respectively, significantly hinders attachment ability. To further test how vital SpaB and SpaC were for attachment, they used antibodies specific to SpaB and SpaC which subsequently prevented attachment. They also coated latex beads with SpaB and SpaC protein, and found that the coated beads bound specifically to the pharyngeal cells. Immunoelectron microscopy and immunofluorescence studies also revealed that the minor pili remain anchored to the cell surface even in the absence of the shaft, suggesting they are tightly anchored in the cell wall. Such research could lead to better methods of treating people who contract C. diphtheriae or bacteria with similar attachment mechanisms. [6]
Transcription of the Contiguous sigB, dtxR, and galE Genes in Corynebacterium diphtheriae: Evidence for Multiple Transcripts and Regulation by Environmental Factors, November 4, 2005
Research conducted by Diana Marra Oram, Andrew D. Jacobson, and Randall K. Holmes at the Department of Microbiology, University of Colorado School of Medicine, gives the first quantitative data about the transcription of a gene which codes for a member of the diphtheria toxin repressor (DtxR) family and information about transcription regulation in C. diphtheriae. It was discovered that the gene dtxR has two promoters, both upstream from dtxR, and both are influenced by environmental factors, specifically iron concentration. By knowing more about transcription regulation, especially in pathogenic bacteria such as C. diphtheriae, novel methods of controlling C. diphtheriae and its close relatives could be developed. [9]
Mechanism of metal ion activation of the diphtheria toxin repressor DtxR, February 25, 2005
J. Alejandro D'Aquino, Jaclyn Tetenbaum-Novatt, Andre White, Fred Berkovitch, and Dagmar Ringe at Brandeis University studied the two metal ion binding sites on the monomers of diphtheria toxin repressor (DtxR) to identify the mechanisms behind metal ion activation. DtxR is particularly important because it is thought that it is key to the virulence of C. diphtheriae. They determined that binding site 1 (ancillary), which has a low metal ion affinity, is the first place that the ion binds before it is transferred to binding site 2 (primary), which has a high metal ion affinity. By being able to formulate a possible mechanism for metal ion activation, the factors which cause the deadliness of C. diphtheriae can be better understood, as well as the function of similar repressors in other organisms. [10]
References
2 Chima J. Ohuabunwo, et al. "Chapter 1: Diphtheria" "VPD Surveillance Manual, 3rd Edition" 2002
4 Medindia Health Network Pvt Ltd. "Diphtheria" 3 May 2007
5 Chung, King-Thom "Emil Von Behring (1857-1917)" , Department of Biology, University of Memphis.
7 Iditarod Trail Committee "Iditarod History" 2007
8 Encyclopedia Britannica, Inc. "Diphtheriae" 2007
11 Todar, Kenneth, University of Wisconsin-Madison Department of Bacteriology, "Diphtheria" 2002
12 American Academy of Pediatrics 2007
14 National Institute of Health 2007
15 WHO Regional Office for South-East Asia "Chapter 11: Diphtheria" 2007
16 Hutchinson, Woods "The Project Gutenberg eBook, A Handbook of Health" 5 January 2007
17 Uris(English Wikipedia), "Image:Balto.jpg" 27 April 2007
Edited by student Rachael Spradley of Rachel Larsen





