Deinococcus radiodurans
A Microbial Biorealm page on the genus Deinococcus radiodurans
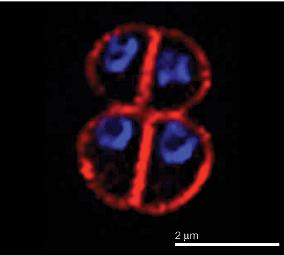
Classification
Higher order taxa
Bacteria; Deinococcus-Thermus; Deinococci; Deinococcales; Deinococcaceae; Deinococcus
Species
D. radiodurans, D. radiodurans R1
Description and significance
Deinococcus radiodurans was first discovered in 1956 in a can of ground meat that had been treated with large doses of radiation to remove all hazardous bacteria from the product. Since then this species has been intensely studied for its radiation resistant properties. It has been known to withstand radiation levels of up to 1,000 times that which would kill a normal human, living up to its latin name, "strange little berry that withstands radiation." D. radiodurans has since been isolated from a variety of habitats, mostly soil and feces based. Being a mesophile, this species grows relatively well between 30-37°C, normally in tetrads.<super>[2]</super>
Genome structure
The genome of D. radiodurans consists of four major parts. The complete sequence of the R1 strain has 3,284,156 base pairs made up of two circular chromosomes (2,648,638 and 412,348 base pairs), a major plasmid (177,466 base pairs), and a small plasmid (45,704 base pairs). No current research shows whether or not these plasmids contribute specifically to functionality or virality. However, it is known that multiple copies of each gene are found on all the chromosomes and plasmids, which most likely contributes to its amazing repair capabilities associated with its radiation resistance.
Cell structure and metabolism
The most interesting aspect about the cell structure of D. radiodurans is that it keeps 4-10 copies of all its genes at any given time depending on its current stage of growth. Many researchers believe this relates to the reason why it can withstand so much radiation. This ability does not rely on some "magic" gene that protects it from radiation, rather, it seems that D. radiodurans is able to more efficiently repair double strand breaks in its DNA that result from radiation damage thanks to these extra copies and a few other special proteins.
Ecology
Application to Biotechnology
Current Research
References
Edited by Edwin Cook, student of Rachel Larsen and Kit Pogliano
