Dermabacter hominis: Difference between revisions
No edit summary |
No edit summary |
||
| Line 1: | Line 1: | ||
{{ | <!--Do not edit or remove this line!-->{{MooreL}} | ||
{{Biorealm Genus}} | {{Biorealm Genus}} | ||
==Classification== | ==Classification== | ||
| Line 62: | Line 62: | ||
Edited by W. David Nesher of Dr. Lisa R. Moore, University of Southern Maine, Department of Biological Sciences, http://www.usm.maine.edu/bio | Edited by W. David Nesher of Dr. Lisa R. Moore, University of Southern Maine, Department of Biological Sciences, http://www.usm.maine.edu/bio | ||
Revision as of 14:35, 22 January 2016
A Microbial Biorealm page on the genus Dermabacter hominis
Classification
Higher order taxa
Bacteria; Actinobacteria; Actinobacteria (class); Microccocales; Dermabacteraceae
Species
|
NCBI: Taxonomy |
Dermabacter hominis
Description and significance
Dermabacter hominis can be found most commonly on human skin. It has also been isolated in a variety of other clinical specimens, including abscesses, infections of bone, a wound, or an eye, and blood cultures. Though considered an opportunistic human pathogen when first isolated in 1988, there has not been any observed direct mortality in relation to D. hominis (as of January 2014).
Genome structure
A draft genome sequence of D. hominis 1368, a multidrug resistant strain of the species D. hominis, revealed 2,507,630 base pairs. Also included in the draft genome of D. hominis 1368 are one non-coding RNA gene, 21 pseudogenes, 48 tRNA genes, and 2,227 protein-coding genes.
Cell and colony structure
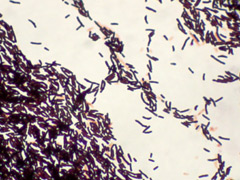
D. hominis are non-motile, mesophilic, and rod shaped. Before assignment to its current genus, D. hominis was grouped with coryneform groups 3 and 5 of the Center for Disease Control. It forms white, convex, creamy or dry colonies on nutrient and blood agar. The colonies grow to 1-1.5 millimeters in diameter after 48 hours.
Metabolism
A facultative anaerobe, D. hominis is able to ferment glucose, maltose, sucrose and lactose. D. hominis is irregularly shaped (some sources list it as being coccus-shaped while others list it as rod-shaped) and is catalase positive and able to decarboxylate ornithine and lysine. Additionally, the microbe is not able to reduce urease, oxidase, or nitrate, which is helpful for identification.
Ecology
D. hominis is a mesophilic microbe, preferring temperatures between 20°C and 45°C. This makes sense since D. Hominis thrives in the human microbiome, as noted above.
In an analysis of Genera Brevibacterium and Dermabacter, it was discovered that D. Hominis seems to be more closely related to the Anthrobacter and Micrococcus subline (roughly 93% 16S rRNA similarity), than to Brevibacterium. This is of interest because both Dermabacter and Brevibacterium have cell walls that contain meso-diaminopimelic acid as a component, a opposed to Anthrobacter and Micrococcus which have lysine as their dibasic amino acid the their cell wall.
Also of note: D. hominis is resistant to daptomycin, which is rare for gram-positive organisms. As of 2014, no definitive reason for this resistance was known, though it was not thought that the resistance was due to use of daptomycin (which is a common cause of resistance).
Pathology
Though there are no recorded cases of mortality as a direct result of the presence of D. hominis, it has caused peritonitis (an inflammation of the peritoneum) in a patient receiving renal replacement therapy with peritoneal dialysis. Even though there have not been any recorded fatalities directly linked to D. hominis, there are several recorded cases where D. hominis is thought to have played a role in the death of an already compromised patient. In addition the the peritonitis case, there are two documented cases of patients with underlying pathologies (one patient had AIDS and the other suffered from chronic cardiorespiratory distress) who developed bacteremias by D. hominis.
References
[http://doi.org/10.1128/genomeA.00728-14; Albersmeier, A., Bomholt, C., Glaub, A., Rückert, C., Soriano, F., Fernández-Natal, I., & Tauch, A. (2014). Draft Genome Sequence of the Multidrug-Resistant Clinical Isolate Dermabacter hominis 1368. Genome Announcements, 2(4), e00728–14.}
[http://doi.org/10.1002/2052-2975.31; Fernández-Natal, I., Sáez-Nieto, J. A., Medina-Pascual, M. J., Albersmeier, A., Valdezate, S., Guerra-Laso, J. M., … Soriano, F. (2013). Dermabacter hominis: a usually daptomycin-resistant gram-positive organism infrequently isolated from human clinical samples. New Microbes and New Infections, 1(3), 35–40.}
[http://doi.org/10.1128/JCM.39.6.2356-2357.2001; Gómez-Garcés, J. L., Oteo, J., García, G., Aracil, B., Alós, J. I., & Funke, G. (2001). Bacteremia by Dermabacter hominis, a Rare Pathogen. Journal of Clinical Microbiology, 39(6), 2356–2357.}
[http://www.ncbi.nlm.nih.gov/pmc/articles/PMC263903/pdf/jcm00008-0096.pdf; Gruner, E., Steigerwalt, A. G., Hollis, D. G., Weyant, R. S., Weaver, R. E., Moss, C. W., … Brenner, D. J. (1994). Recognition of Dermabacter hominis, formerly CDC fermentative coryneform group 3 and group 5, as a potential human pathogen. Journal of Clinical Microbiology, 32(8), 1918–1922.} [doi:10.1099/ijs.0.028217-0.]
[http://doi.org/10.1128/JCM.39.9.3420-3421.2001; Radtke, A., Bergh, K., Øien, C. M., & Bevanger, L. S. (2001). Peritoneal Dialysis-Associated Peritonitis Caused by Dermabacter hominis. Journal of Clinical Microbiology, 39(9), 3420–3421.}
[http://www.microbiologyresearch.org/docserver/fulltext/ijsem/44/3/ijs-44-3-583.pdf?expires=1449940194&id=id&accname=guest&checksum=69677DA902D9116428BE8200DC74729C; Ca J., Collins M. Phylogenetic Analysis of Species of the meso-Diaminopimelic Acid-Containing Genera Brevibacterium and Demabacter. Int J Syst Evol Microbiol 44(3):583-585.}
Edited by W. David Nesher of Dr. Lisa R. Moore, University of Southern Maine, Department of Biological Sciences, http://www.usm.maine.edu/bio