Desulfuromonas
Classification
Higher order taxa:
Bacteria; Proteobacteria; delta/epsilon subdivisions; Deltaproteobacteria; Desulfuromonadales; Desulfuromonadaceae
Species:
D. acetexigens, D. acetoxidans D. chloroethenica, D. michiganensis, D. palmitatis, D. thiophila, D. sp.
Description and Significance
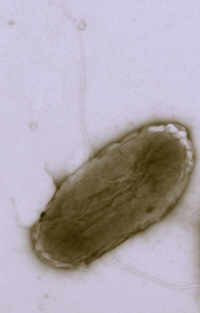
Desulfuromonasis a Gram-negative bacteria, known for being a sulfur reducing eubacteria through conversion of elemental sulfur into sulfide. Unlike some other sulfur-reducing bacteria, it does not reduce sulfate. A recent study has demonstrated that Desulfuromonas is a distinct phylogenetic cluster (one of three, the others being Geobacter and Dulfuromusa) within the the phylogenetically coherenent family Geobaceraceae, which falls within the delta subclass Proteobacteria. Click here for more information.
Genome Structure
Projects to sequence the genome of D. acetoxidansorganisms are beginning by organizations such as the Biomax Informatics project, but so far only a preliminary analysis has been completed. The unfinished sequence of Desulfuromonas acetexigens can be found here.
Cell Structure and Metabolism
Desulfuromonas, like other sulfur-reducing bacteria, has a very unique and complex metabolism. Desulfuromonasuses a variety of enzymes and organic compounds in order to reduce sulfate into sulfide. The reduction process is extremely complex, and the flow of reducing equivalents from electron donors is often linked to respiratory energy conservation utilizing a range of electron carriers as well. The reduction to sulfide from sulfate process has eight-electron steps and uses a variety of intermediates. Desulfuromonas typically would not excrete until the final oxidation state with the product of sulfide (R. Rabus et al., 2004).

Ecology
Desulfuromonasspecies are typically scarce in freshwater, but found frequently in anoxic marine and brackish water sediments because of the high suflur concentration ( R. Rabus et al., 2004).
References
Biomax Informatics. 2004. PENDANT Genome Database.
Joint Genome Institute. 2004. Draft Genome. Desulfuromonas acetoxidans.
