Hamiltonella defensa: Difference between revisions
| Line 38: | Line 38: | ||
==References== | ==References== | ||
1. [https:// | 1.[https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5512215/ Ankrah, Nana Y D et al. “Cooperative Metabolism in a Three-Partner Insect-Bacterial Symbiosis Revealed by Metabolic Modeling.” Journal of bacteriology vol. 199,15 e00872-16. 11 Jul. 2017, doi:10.1128/JB.00872-16] | ||
2. [https:// | 2. [https://academic.oup.com/gbe/article/10/3/786/4857210 Chevignon, Germain et al. “Culture-Facilitated Comparative Genomics of the Facultative Symbiont Hamiltonella defensa.” ''Genome Biology and Evolution'' vol. 10,3 (2018): 786-802. doi.:10.1093/gbe/evy036] | ||
3. [https://www.ncbi.nlm.nih.gov/pmc/articles/ | 3. [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2576707/ Degnan, Patrick H, and Nancy A Moran. “Diverse phage-encoded toxins in a protective insect endosymbiont.” Applied and environmental microbiology vol. 74,21 (2008): 6782-91. doi:10.1128/AEM.01285-08] | ||
4. [https:// | 4. [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2690004/ Degnan, Patrick H et al. “Hamiltonella defensa, genome evolution of protective bacterial endosymbiont from pathogenic ancestors.” ''Proceedings of the National Academy of Sciences of the United States of America'' vol. 106,22 (2009): 9063-8. doi:10.1073/pnas.0900194106] | ||
5. [https:// | 5. [https://aem.asm.org/content/80/18/5818/article-info#sec-2 Dykstra, Hannah R et al. “Factors Limiting the Spread of the Protective Symbiont Hamiltonella defensa in Aphis craccivora Aphids.” ''Applied and Environmental Microbiology'', vol. 80,18 (2014): 5818-27. doi:10.1128/aem.01775-14.] | ||
6. [https://www.ncbi.nlm.nih.gov/pmc/articles/ | 6. [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1151865/ Moran, Nancy A et al. “Evolutionary relationships of three new species of Enterobacteriaceae living as symbionts of aphids and other insects.” ''Applied and environmental microbiology'' vol. 71,6 (2005): 3302-10. doi:10.1128/AEM.71.6.3302-3310.2005] | ||
7. [https://www. | 7. [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1287993/ Moran, Nancy A et al. “The players in a mutualistic symbiosis: insects, bacteria, viruses, and virulence genes.” Proceedings of the National Academy of Sciences of the United States of America vol. 102,47 (2005): 16919-26. doi:10.1073/pnas.0507029102] | ||
8. Yong, Ed. ''I Contain Multitudes: the Microbes Within Us and a Grander View of Life. Ecco, an Imprint of HarperCollinsPublishers, 2016. | 8. [https://www.pnas.org/content/102/36/12795 Oliver, K. M., et al. "Variation in Resistance to Parasitism in Aphids Is Due to Symbionts Not Host Genotype." ''Proceedings of the National Academy of Sciences,'' vol. 102,36 (2005): 12795-800. doi:10.1073/pnas.0506131102.] | ||
9. Yong, Ed. ''I Contain Multitudes: the Microbes Within Us and a Grander View of Life. Ecco, an Imprint of HarperCollinsPublishers, 2016. | |||
==Author== | ==Author== | ||
Revision as of 03:00, 1 May 2020
Classification
Domain: Bacteria; Phylum: Proteobacteria; Class: Gammaproteobacteria; Order: Enterobacterales; family: Enterbacteriaceae; Genus: Hamiltonella
Species
Hamiltonella defensa
Description and Significance
Hamiltonella defensa is a host-restricted mutualist symbiont. It is conditionally beneficial, which means that a couple of factors have to be present in order for H. defensa to serve its purpose. Its main purpose is protecting aphids from braconid wasp parasitoids. It is an endosymbiont of aphids, so it lives inside these sap-sucking insects. This bacterium is important because it has the unique ability to defend its aphid host from these invasive parasitoids.
H. defensa has an extremely dynamic and diverse genome. It is so diverse, in fact, that scientists had difficulty classifying it, due to how many similarities it has with other bacteria that are actually distantly related, as well as how many differences there are between it and its seemingly closely relatives.
Genome Structure
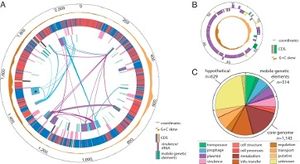
Hamiltonella defensa has an extremely dynamic genome. It is relatively small, only 2.1 Mb with circular chromosome. It encodes 2,100 protein-coding genes, has a relatively large number of pseudogenes, and is littered with mobile DNA, insertion sequences, and phage remnants. Horizontal gene transfer (HGT) plays a role in its dynamic genome. An example of HGT at work in H. defensa's genome is the fact that approximately half of H. defensa's DNA comes from toxin-encoding bacteriophages called APSEs. APSEs are similar to lamda-like phages, which are bacterial viruses that infect E. coli (it has other similarities to the E. coli bacterium, including a similar number of pseudogenes). DNA from different strains of APSEs can be horizontally transferred to H. defensa. Depending on which strain of APSE H. defensa obtains its DNA from, the development stage at which the parasitoid wasps are killed varies. However, not much is known about trait variation among strains due to the fact that only one isolate has been fully sequenced [1]. H. defensa was isolated from TN5 cells, allowing it to have no contamination from aphids, Buchnera, or other bacteria.
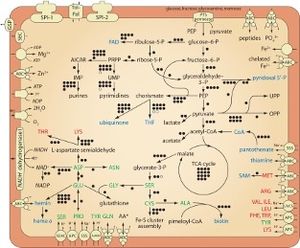
H. defensa is auxotrophic for 8 of its 10 essential amino acids. It relies on Buchnera, a primary endosymbiont of aphids, to synthesize these amino acids. However, despite H. defensa's limited biosynthetic capabilities, it has considerably more cell structural, DNA replication, recombination, and repair genes than do obligate endosymbionts. This is likely due to its extremely varied sources of DNA. Additionally, its diverse genome allows for a wide variety of regulatory genes which help H. defensa to cope with changes to its environment, such as attack of hosts by parasitoids or an invasion of a new host species. Additionally, its genome reflects many pathogenicity factors related to host invasion [3].
H. defensa's genome tells a story of a long-term relationship with insects through its reduced size and compositional bias. It contains genes for toxins, effector proteins, and other things that are critical in its mutualistic relationship with its host. This close endosymbiotic relationship has led to gene losses and inactivations for H. defensa, but these losses are remedied by the genes it procures from other sources through HGT [3].
Cell Structure, Metabolism and Life Cycle
Hamiltonella defensa can be maternally transmitted or passed through horizontal gene transfer between aphids [3]. It is found in approximately 34% of aphids. This percentage may seem low based on how beneficial this bacterium can be to its host. However, the reason it is not found in a higher percentage of aphids is because its presence can be costly to its host if there is no threat present. Specific costs to a host vary among aphid species, but generally, survivorship and life span were lower in aphids infected with H. defensa when no threat was present.
Additionally, H. defensa can conduct glycolysis on its own, which is a very unique ability for endosymbionts. This is important because it allows for energy production, even if H. defensa is living in oxygen-limited environments [4]. It also has the ability to export 14 unique metabolites to its host [1].
Protective Mechanism and Pathogenesis
As previously mentioned, Hamiltonella defensa has the ability to kill braconid wasp parasitoids by blocking their larval development [3]. Braconid wasp parasitoids target aphids by laying their eggs inside of an aphid host, thereby killing it. Research shows that aphids which contain H. defensa, specifically with APSE DNA, can resist these parasitoids and survive. They do this by killing the wasps before they grow large enough to kill the aphid host. The exact mechanism by which H. defensa does this is unknown. Some proposed theories are that the phages could be directly poisoning the wasps or that the phages could be splitting the Hamiltonella apart causing their toxins to spill over the wasps, or a combination of the two [8]. In order for this bacterium to be beneficial to its aphid host, a parasite threat has to be present and the bacteriophage APSE DNA has to be a part of H. defensa's genome. This is why it is conditionally beneficial. Without these two conditions present, H. defensa is useless to its host.
Despite H. defensa's amazing protective ability, it exhibits many similarities to enteric pathogens, which are present in the gut. Similar to enteric pathogens, it is necessary for H. defensa to invade a novel host and, as previously mentioned, contains many pathogenicity factors in its genome [6].
References
9. Yong, Ed. I Contain Multitudes: the Microbes Within Us and a Grander View of Life. Ecco, an Imprint of HarperCollinsPublishers, 2016.
Author
Page authored by Isabella Valli, student of Prof. Jay Lennon at IndianaUniversity.