Holdemania filiformis: Difference between revisions
No edit summary |
|||
| Line 1: | Line 1: | ||
{{Uncurated}} | |||
A Microbial Biorealm page on the genus ''Holdemania filiformis'' | A Microbial Biorealm page on the genus ''Holdemania filiformis'' | ||
| Line 39: | Line 40: | ||
Edited by a student of Dr. Lisa R. Moore, University of Southern Maine, Department of Biological Sciences, http://www.usm.maine.edu/bio | Edited by a student of Dr. Lisa R. Moore, University of Southern Maine, Department of Biological Sciences, http://www.usm.maine.edu/bio | ||
Revision as of 13:36, 1 October 2015
A Microbial Biorealm page on the genus Holdemania filiformis
Higher order taxa
Domain: Bacteria, Phylum: Firmicutes, Subphylum: Clostridium, Class: Erysipelotrichia, Order: Erysipelotrichales, Family: Erysipelotrichaceae
Genus, Species
Holdemania filiformis
Description and significance
no photo available at the current time
Holdemania filiformis is a Gram-positive bacterium found in the gut of healthy humans. It is rod shaped and exists in pairs and short chains. Cells can be short or long (numerical measurements are currently unavailable), with longer cells possibly exhibiting central or terminal swellings. It has not been observed to be spore forming or heat resistant. At present, filiformis is the only species in the genus Holdemania.1 The amount existing in the gut and the effects of its presence is unknown. It is resistant to vancomycin.2
Genome structure
H. filiformis has a single circular chromosome consisting of 3,932,923 base pairs5, as submitted to the National Center for Biotechnology Information (NCBI) on December 19th, 2008 by the Genome Sequencing Center at Washington University School of Medicine.6 The genome was sequenced using shotgun assembly. 4223 hypothetical protein coding regions have been recognized.7 Some exampes may be found here.4
Cell and colony structure
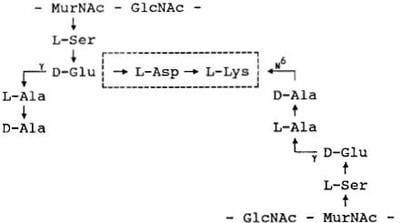
H. filiformis is a Gram positive-staining bacterium that easily decolorizes. The cell wall contains a group B murein type (B1δ(L-Ala)-D-Glu-L-Asp-L-Lys) that has not been observed in any other bacterial species.1 Colonies grown on brain heart infusion agar are 1.0 mm in diameter. They are circular, entire, low convex, and translucent with a granular appearance. PYT-glucose broth cultures were turbid and contained white sediment.2
Metabolism
Strictly anaerobic.2 It is saccharolytic. Good growth was exhibited in PYT broth media containing esculin, fructose, glucose, salicin, or sucrose. Moderate growth can sometimes be observed in peptone-yeast extract broth with maltose or lactose. Minimal growth was seen in PYT broths with no carbohydrates or with amydalin, arabinose, cellobiose, erythritol, glycogen, inositol, mannitol, mannose, melzitose, melibiose, raffinose, rhamnose, ribose, sorbitol, starch, trehalose, or xylose. In PYT-glucose broth, the consistent end products of fermentation were acetic acid, lactic acid, and succinic acid.2 Meat, gelatin, and milk are not digested or changed. Esculin is hydrolyzed; nitrate is not reduced; arginine is not deanimated; catalase and indole are not produced. It has been shown to grow in media ranging from a pH of 5.8 to 6.5 1,2 consistent with ranges found in the human large intestine.
Ecology
H. filiformis is part of the normal human gut flora. It was originally isolated from human feces.2 It appears to be mesophilic.
Pathology
H. filiformis has not been observed to be pathogenic.2
References
1 Dos, P. D., Garrity, G. M., Jones, D., Krieg, N. R., Ludwig, W., Rainey, F. A., Schleifer, K., Whitman, W. B., Parte, A. C., Goodfellow, M. 2009 Bergey’s manual of systematic bacteriology. Volume three, the firmicutes. Dordrecht, NY 1417 pp. doi# 10.1007/b92997
2 Willems, A., Moore, W. E. C., Weiss, N., Collins, M. D. 1997. Phenotypic and Phylogenetic Characterization of some Eubacteriurn-like isolates containing a novel type B wall murein from human feces: description of Holdemania filiformis gen. nov., sp. nov. International J. of Systematic Bacteriolol.47: 1201-1204 doi# 10.1099/00207713-47-4-1201
3 Holdemani filiformis overview. http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&id=61171&lvl=3&lin=f&keep=1&srchmode=1&unlock May 2, 2012
4Holdemania filiformis peptidase or homologue coding regions. http://merops.sanger.ac.uk/cgi-bin/speccards?sp=sp019394;type=P May 3 2012
5Holdemania filiformis overview: Genome. http://www.ncbi.nlm.nih.gov/genome/2036 May 2, 2012
6Holdemania filiformis master record for whole genome shotgun sequencing. http://www.ncbi.nlm.nih.gov/nuccore/223965545 May 2, 2012
7 Holdemania filiformis reference genome for the Human Microbiome Project. http://www.ncbi.nlm.nih.gov/bioproject/30361 May, 2, 2012
Edited by a student of Dr. Lisa R. Moore, University of Southern Maine, Department of Biological Sciences, http://www.usm.maine.edu/bio