Infection Mechanism of Genus Ebolavirus: Difference between revisions
No edit summary |
No edit summary |
||
| Line 1: | Line 1: | ||
{{Curated}} | |||
{{Viral Biorealm Family}} | {{Viral Biorealm Family}} | ||
Revision as of 00:31, 9 April 2011
A Viral Biorealm page on the family Infection Mechanism of Genus Ebolavirus
Ebola hemorrhagic fever is a severe, often fatal, disease in humans and non-human primates caused by the Ebola virus (Fig 1). Even though the virus first appeared in the Democratic Republic of the Congo (formerly Zaire) in the summer of 1976 and has only sporadically reemerged, it has still managed to capture the attention of the world due to death rates that can be as high as 90% as well as the visceral manner in which it kills [7,24]. Few viruses have the ability to turn internal organs into a soup that promptly flows out of the body, so those that do tend to capture the public eye.
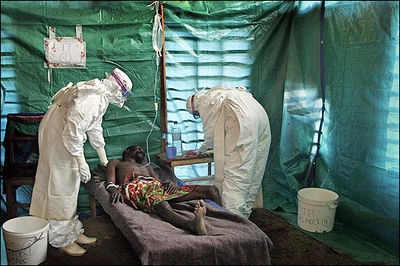
Despite the low incidence of infection, the lethality and potential airborne transmission of Ebola virus makes it a severe biological threat. This is compounded by the possibility of the virus being obtained from the wild for use as a biological weapon, something Japanese terrorist cult Aum Shinrikyo once unsuccessfully attempted [5]. As of the present, there are no licensed vaccines or specific antiviral treatments available for Ebola virus infections, with significant progress only made in the development of vaccines for nonhuman primates [34]. Therefore, the elucidation of the viral replication cycle as well as complete understanding of viral proteins is necessary for the production of human treatments.
Ebola Hemorrhagic Fever
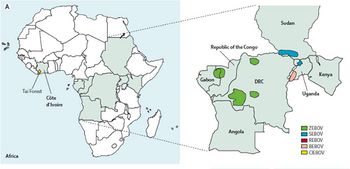
This mysterious hemorrhagic fever was first described in two separate 1976 outbreaks (Fig 2): first in southern Sudan and subsequently in northern Zaire, now Democratic Republic of the Congo. A causative agent was isolated from patients in both epidemics and named Ebola virus after a small river in northwestern Zaire. Only years later did researchers recognize that the plagues were caused by two distinct species of Ebola virus, Sudan Ebola virus and Zaire Ebola virus. The third African species, Cote d'Ivoire Ebola virus was isolated in 1994 from an infected ethnologist who had done a necropsy on a chimpanzee from the Tai Forest. Only in 2007 was a fourth African species, Bundibugyo Ebola virus, isolated [13].
An additional viral species, Reston Ebola virus, has origins in the Philippines. It was first discovered in a quarantine facility for Cynomolgus monkeys (Macaca fascicularis) in Reston, VA, USA, and made popular by Richard Preston's best-selling novel The Hot Zone. Luckily for the monkey caretakers, this is the only strain that is non-pathogenic in humans. Recently, Reston Ebola virus has been found infecting pigs in the Philippines. This raises important concerns for public health and food safety in this country, causing some to worry that this could turn into a serious problem for parts of Asia [13].
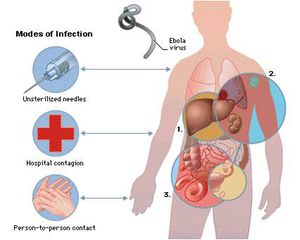
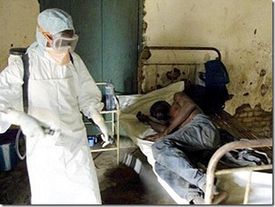
The Ebola virus enters the host primarily through mucosal surfaces or skin abrasions, with most human infections occurring after direct contact with patients or cadavers. In addition, there have been several cases in which hunters are infected after eating contaminated meat found in the forest. Also, most Ebola outbreaks are centered in hospitals in Africa where practices of basic hygiene and sanitation are considered luxuries (Fig 3). However, in modern hospitals with disposable needles and knowledge of basic hygiene and barrier nursing techniques, the virus rarely spreads on a large scale. Airborne transmission between monkeys was demonstrated during the outbreak of Reston Ebola virus in Virginia, but there is limited evidence of airborne transmission in any human epidemics [13].
One of the reasons that Ebola is so dangerous is that its symptoms are varied and appear quickly, yet resemble those of other viruses so much that the hemorrhagic fever is not rapidly diagnosed. Generally, the abrupt onset of symptoms follows an incubation period of 2-21 days and is characterized by high fever, chills and myalgia. Subsequent signs of infection indicate multisystem involvement and include gastrointestinal (nausea, vomiting, diarrhea), respiratory (chest pain, cough) and neurological (headache) manifestations. The unfortunate thing about these symptoms is that they are easily mistaken for malaria, typhoid fever, dysentery, influenza or various bacterial infections, all of which are far more common in regions where Ebola is prevent, but also less fatal [7].
After these initial symptoms, the fever often progresses to cause more serious ones: diarrhea, dark or bloody feces, vomiting blood, red eyes due to distention and hemorrhage of sclerotic arterioles, petechia, maculopapular rash and purpura. There is often gastrointestinal bleeding from the mouth and rectum, sometimes leading to the sloughing of the gut and venting from the anus. Internal and external hemorrhage from orifices, such as the nose and mouth, may also occur. As the virus progresses, bleeding in the brain can lead to severe depression, seizures and delirium [7].
The span of time from onset of symptoms to death is usually between 6 and 16 days. By the second week of the infection, the patient will either experience a full recovery or undergo systemic multiorgan failure. Mortality rates are generally high, ranging from 50-90% depending on the specific strain. The cause of death is normally due to hypovolemic shock or organ failure (Fig 4). While hemorrhages can be severe and have been the calling card of this virus, they are actually present in fewer than half of patients [53]. Here, Chief Epidemiologist Philippe Calain from the CDC Special Pathogens Branch sums up the appearance of his patients during the 1995 Ebola outbreak in Kikwit:
"At the end of the disease the patient does not look, from the outside, as horrible as you can read in some books. They are not melting. They are not full of blood. They're in shock, muscular shock. They are not unconscious, but you would say 'obtunded', dull, quiet, very tired. Very few were hemorrhaging. Hemorrhage is not the main symptom. Less than half of the patients had some kind of hemorrhage. But the ones that had bled, died" [28]
Virion Structure
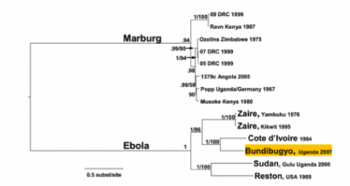
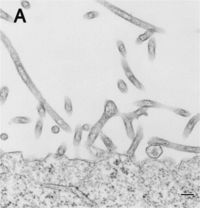
Filoviridae, of which Ebola virus is a member, is a family of viruses that contain single, linear, negative-sense ssRNA genomes. The family name was derived from the Latin word filum, which alludes to the thread-like appearance of the virions when viewed under an electron microscope (Fig 1). Filoviruses have been divided into two genera: Ebola-like viruses with species Zaire, Sudan, Reston, Cote d’Ivoire and Bundibugyo; and Marburg-like viruses with the single species Marburg (Fig 5). All of these are responsible for hemorrhagic fevers in primates that are characterized by often fatal bleeding and coagulation abnormalities [25].
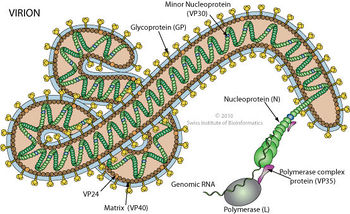
The tubular Ebola virions are generally 80 nm in diameter and 800 nm long. In the center of the particle is the viral nucleocapsid which consists of the helical ssRNA genome wrapped about the NP, VP35, VP30 and L proteins. This structure is then surrounded by an outer viral envelope derived from the host cell membrane that is studded with 10 nm long viral glycoprotein (GP) spikes. Between the capsid and envelope are viral proteins VP40 and VP24 [14] (Fig 6).
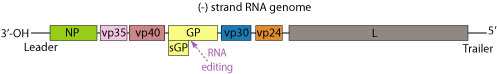
The genome of each virion is around 19kb in length, and codes for seven structural and one non-structural proteins. The gene order is as follows: 3′ – leader – NP – VP35 – VP40 – GP/sGP – VP30 – VP24 – L – trailer – 5′ (Fig 7). The leader and trailer regions are not transcribed, but carry important signals that control transcription, replication and packaging of the genome into new virions [9]. The seven genes each consist of their respective open reading frame as well as long non-translated sequences of unknown purpose that flank the coding regions [25].
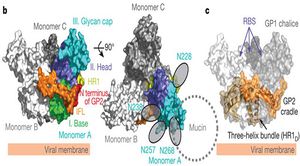
Ebola actually encodes two forms of its glycoprotein gene. The small, non-structural, dimeric soluble form (sGP) is transcribed directly from the viral mRNA and its function remains mostly unknown [43]. This protein is not found in virus particles, but is instead secreted from infected cells into the blood [47]. A second glycoprotein results from transcriptional editing of the glycoprotein origin of replication and encodes a trimeric, membrane-bound form. This envelope GP spike is expressed at the cell surface, and is incorporated into the virion to drive viral attachment and membrane fusion. It has also been shown as the crucial factor for Ebola virus pathogenicity [55]. GP is actually post-translationally cleaved by the proprotein convertase furin to yield disulphide-linked GP1 and GP2 subunits. GP1 allows for attachment to host cells, while GP2 mediates fusion of viral and host membranes [48]. This protein assembles as a trimer of heterodimers on the viral envelope, and ultimately undergoes an irreversible conformation change to merge the two membranes [27].
The product of the third gene, VP40, is located beneath the viral envelope where it helps to maintain structural integrity. It has also been associated with late endosomes and likely mediates filovirus budding due to its ability to induce its own release from cells in the absence of all other viral proteins [21]. The second matrix protein, VP24, has been shown to suppress interferon production. However, interferon interference may not be its only function. Other experiments have shown that this protein, along with VP35 and NP are sufficient to form nucleocapsid structures [19]. Lastly, VP24 is necessary for the correct assembly of a functional nucleocapsid, as a lack of VP24 leads to reduced transcription/translation of VP30 [18].
The remaining structural proteins form the nucleocapsid, and are thus intimately associated with the viral genome. These are the nucleoprotein NP, the polymerase cofactor VP35, the viral-specific transcription activator VP30 and the viral RNA polymerase L. These nucleocapsid proteins have a dual function in the viral replication cycle: they are involved as structural components, but also catalyze replication and transcription of the genome. While NP, VP35 and L are sufficient for replication, transcription initiation will not proceed without VP30 [33].
Immune System Evasion
One of the reasons that Ebola is so deadly is that it has multiple ways of interfering with or avoiding the human immune system. While the virus is busy destroying the human body, the immune system is either still in the process of discovering that there is a problem, or is in such disarray that it would be next to impossible to mobilize a unified effort to fight off the invader.
Among the five strains of Ebola virus, the Zaire strain appears to be the most virulent, with a mortality rate of up to 90%. Despite extensive research, the molecular basis for this virulence has not been determined. However, this question came a bit closer to being answered thanks to the work of Takada et al. [45]. They were able to demonstrate that the infectivity of the Zaire strain of virus was markedly enhanced in the presence of mouse monoclonal antibodies, suggesting an antibody-dependent enhancement of infection (Fig 8A). Furthermore, serum samples collected from patients infected with the Ebola Zaire virus during the 1995 outbreak in Kikwit, Democratic Republic of the Congo (no. 2 and 6), enhanced the infectivity of virus in human embryonated kidney cells [45] (Fig 8B).
The authors had previously shown that antibody-dependent enhancement requires a heat-labile serum factor that can interact with the Fc portion of antibodies since heat treatment reduced the infectivity-enhancing ability of antiserum to GP. Protein A, a surface protein from the cell wall of the bacteria Staphylococcus aureus that binds the Fc region of immunoglobulins, had the same effect. The authors thus decided to study complement component 1 (C1), an initial component of the classical complement pathway, since it is heat labile, binds to Fc and also interacts with cell surface molecules. This complex consists of C1q and two serine protease proenzymes, C1r and C1s. Experiments proved that the infectivity of Ebola virus was enhanced in the presence of mouse monoclonal antibodies and purified C1q, demonstrating that antibody-dependent enhancement is mediated by the C1q molecule [45].
These findings all suggest a novel mechanism of enhancement: multiple IgG antibodies bind to GP epitopes in close proximity, allowing the binding of C1 to the Fc region of the antibodies (Fig 9). This complex binds C1q ligands on the cell surface and stabilizes the interaction between the virus and its receptor, increasing the likelihood of viral attachment. C1q ligands have been identified on many cell types, including the monocytes/macrophages and endothelial cells that are preferentially targeted by this hemorrhagic virus. Therefore, while antibodies normally protect the body, this virus is able to use them for faster and easier attachment to target cells [45].
In addition being an essential cofactor for the viral RNA polymerase complex, VP35 has been identified as an inhibitor of multiple components of the interferon (IFN) pathways [1] (Fig 10,12). These are a family of cytokines produced in response to viral infection that exert antiviral, cell growth-inhibitory and immunoregulatory activities. Upon viral infection, the IFN response can be triggered by sensors such as retinoic acid-inducible gene I (RIG-I) protein and the melanoma differentiation-associated gene 5 (MDA-5) protein that recognize dsRNA or ribonucleoprotein complexes (Fig 10). However, VP35 is able to disrupt this pathway just as it begins by competing with RIG-1 for the binding of dsRNA [6].
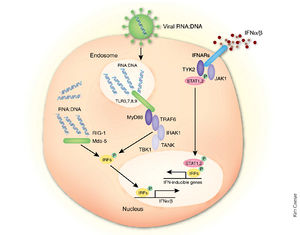
Under normal circumstances, signal propagation occurs through the mitochondrion-associated adapter IFN-β promoter stimulator 1 (IPS-1) which subsequently activates inhibitor of κB kinase epsilon (IKKε) and TANK-binding kinase (TBK-1). These in turn phosphorylate the otherwise inactive interferon regulatory factors 3 (IRF-3) to promote IFN- α/β secretion (Fig 10). IRF-3 is a constitutively expressed transcription factor that is predominantly cytoplasmic when in its inactive form. Following its activation by serine/threonine phosphorylation near its carboxy terminus, the protein dimerizes and moves into the nucleus to bind DNA and interact with histone acetyltransferases. This factor contributes to the immediate activation of IFN-β gene expression and of select IFN-α genes as well as several other genes with potential antiviral activity [26] (Fig 10).
However, VP35 inhibits IRF-3 activation primarily by inhibiting its phosphorylation. Results indicate that in the absence of this antagonist activity, large amounts of IFN-α/β would be produced that would inhibit viral spread [2]. In addition, these interferons influence the production of other immunoregulatory cytokines. This mechanism therefore provides a possible explanation for the blocking of dendritic cell maturation visualized during Ebola infection which impairs T-cell activation and proliferation [22].
Another mechanism of inhibition involves a physical interaction between VP35 and the cellular kinases IKKε and TBK-1. Immunoprecipitation data suggests that VP35 may succeed in targeting these proteins by interacting with their relatively similar kinase domains. In addition, the viral protein can disrupt IKKε and IPS-1 interactions, partially preventing the activation of these kinases at high virion concentrations [35].
Once bound to their receptors, interferons activate the Janus tyrosine kinase / signal transducer and activator (JAK/STAT) signaling pathway that stimulates transcription of genes encoding antiviral proteins such as protein kinase R (PKR). Upon binding to dsRNA, PKR is activated and phosphorylates translation initiation factor eIF-2, which then arrests protein synthesis and inhibits viral replication [15].
VP35 is also able to counteract the antiviral action of interferon in infected cells by suppressing the pathway regulated by double-stranded RNA-dependent protein kinase PKR. It accomplishes this goal by inhibiting phosphorylation of PKR and eIF-2α. This ability seems independent to the dsRNA binding activity of the protein, and might involve inhibition of the activity of PACT, a cellular protein that activates PKR under stress conditions [15]. Recently, the C-terminal IRF inhibitory domain has been shown to be crucial in blocking PKR activation, going so far as to reverse phosphorylation of the host protein [41].
Fatal Ebola infections are marked by unchecked viral replication combined with an inadequate antiviral response. In order for this to occur, the early antiviral innate immune response must be delayed or inhibited. As evidenced by all of these proposed mechanisms, VP35 clearly plays a critical role in the pathogenesis of this extremely deadly virus.
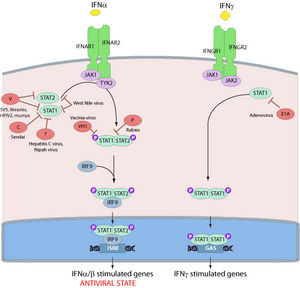
As previously mentioned, the second matrix protein, VP24, suppresses interferon production as well. The binding of IFN-α/β and IFN-γ to their respective receptors activates STAT complexes, a family of transcription factors that regulate immune system gene expression. Normally, STAT1:STAT2 heterodimers and STAT1 homodimers are transported to the nucleus by karyopherin α1 and activate numerous genes involved in antiviral activities (Fig 11). However, VP24 competes with STAT1 to bind karyopherin α1, blocking nuclear accumulation and leading to inhibition of IFN signaling [37].
During infection, monocytes/macrophages in the lymphoid tissues are early and sustained targets of this deadly virus. Since these cells usually elicit the response cascade in the acute phage of inflammation, their early infection helps Ebola evade the immune system while subsequently spreading throughout the host. In addition, infected macrophages release increased amounts of nitric oxide (NO), a gaseous hormone that normally functions in cell communication. However, in high concentrations, NO depresses the mitochondrial membrane potential, causing apoptosis in nearby natural killer cells [17] (Fig 12).
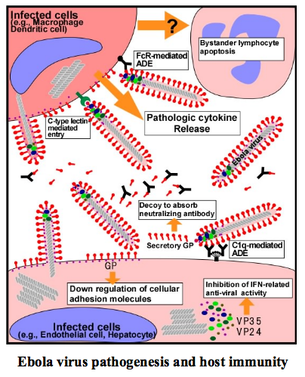
So far, most of the processes by which Ebola escapes or hampers immune response involve viral structure proteins. However, other studies have implicated a role of sGP in immune system evasion. During viral infection, large amounts of proinflammatory cytokines like tumor necrosis factor alpha (TNF-α) are secreted from infected macrophages and cause disruption of the endothelial barrier (Fig 12). Surprisingly, when sGP was administered simultaneously with TNF-α, it caused a recovery of barrier function [49].
This suggests an anti-inflammatory role of sGP that helps to explain somewhat mysterious observations during infection. When a patient is suffering from Ebola, tissue destruction can be seen in multiple organs. However, these areas are not infiltrated by leukocytes, which instead appear to congregate within the adjacent vascular system. It appears that the recruitment of neutrophils via an activated endothelium has occurred, but the transmigration process is blocked by the anti-inflammatory effect of sGP [49]. In addition, sGP shares many neutralizing epitopes with GP, suggesting that this secreted protein may serve as a decoy that absorbs antibodies [20] (Fig 12).
Viral Entry
As the first step of the viral life cycle, entry into the host cell is a popular drug target as infection could be stopped before replication disrupts cell function. However, the entry mechanism of Ebola virus is poorly understood. Many enveloped viruses, Ebola virus included, rely upon endocytosis to infect cells. Several distinct endocytic mechanisms exist in mammalian cells, and can be distinguished by the type of cargo they carry as well as the proteins involved in their regulation [10]. However, all mechanisms ultimately transport virions through successive endocytic vesicles until a compartment with adequate conditions, low pH in the case of Ebola, is reached [42]. Upon membrane fusion, the capsid moves into the cell cytoplasm at a site where replication proceeds optimally.
Previous studies to clarify the entry mechanism of Ebola have produced conflicting results, with involvement of clathrin-mediated endocytosis [4], caveolin-mediated endocytosis [12] and a Rho GTPase-dependent pathway that may suggest macropinocytosis all being implicated [36]. However, all of these experiments were performed using retrovirus-based pseudotypes in which GP is coated onto the surface of a retrovirus capsid containing a recombinant genome. This removes the need to work in a biosafety level four laboratory, but suffers from non-ideal biochemical characteristics. Only recently have experiments been performed with wild-type Ebola virus Zaire that demonstrate that cellular entry involves uptake by a macropinocytosis-like mechanism [38] (Fig 13).
Throughout these experiments, the researchers relied heavily upon both infectious Zaire Ebola virus (ZEBOV) as well as morphologically comparable virus-like particles (VLPs) composed of VP40, GP and Nef-luciferase fusion protein. Immunofluorescence staining for matrix protein VP40 was used to determine infection when infectious virions were used, but these experiments are not specific for viral uptake. Importantly, a cell entry assay utilizing VLPs was used to discriminate between blocks in virus entry versus blocks in later steps in the infection cycle. In this case, virus-endosomal membrane fusion was monitored by luciferase release from VLPs into the cytoplasm [38].
Saeed et al. first sought to further elucidate the role of clathrin-mediated endocytosis (CME) and caveolin-mediated endocytosis (CavME) in cellular entry of Ebola virus by utilizing specific dominant-negative mutations (DN), alterations that produce an altered gene product that acts antagonistically to the wild-type allele, of Eps15 and caveolin-1. Eps15 and caveolin-1 are required for the formation of CME and CavME vesicles respectively, so their DN mutations inhibit their respective endocytosis pathways with high specificity. They determined that neither DN mutation had a significant impact on Ebola infection or entry, indicating that neither CME not CavME play a major role in these processes (Fig 14). Further experimentation in the same vein proved that cell entry of the virus is independent of dynamin, a large GTPase that plays a critical role in CME and CavME [38].
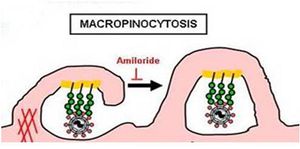
Saeed et al. later determined that Ebola virus associates with lipid rafts during entry by visualizing extensive co-localization of gfpZEBO-VLPs with the lipid raft protein CTxB (Fig 15). They therefore knew that they were looking for a dynamin-independent, lipid raft-dependent, non-clathrin/non-caveolar endocytic mechanism of entry [38]. Macropinocytosis is one such path that has already been shown to be important for the uptake of vaccinia virus [30]. This mechanism is associated with outward extensions of the plasma membrane formed by actin polymerization. These so called membrane ruffles can fold back upon themselves and form a macropinosome upon fusion of the distal loop ends [51] (Fig 13).
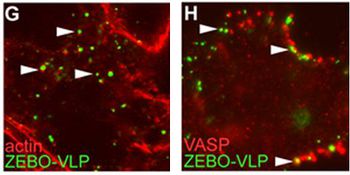
The involvement of macropinocytosis was tested through the use of EIPA (5-(N-ethyl-N-isopropyl amiloride), a potent and specific inhibitor of Na+/H+ exchanger activity important for macropinosome formation. They discovered a dose-dependent inhibition of gfpZEBOV infection as well as severe inhibition of ZEBO-VLP entry in the presence of this amiloride (Fig 16). Further experiments determined that Ebola virions colocalize with internalized dextran, a complex polysaccharide taken in by macropinocytosis, and requires the activity of p53-activated kinase 1, another hallmark of this entry pathway. Lastly, it was noted that gfpZEBO-VLPs were associated with Arp2 and vasodilator-stimulated phosphoprotein (VASP), two actin-associated proteins that promote actin assembly (Fig 17). This also points towards macropinocytosis, since actin is required for the formation of plasma membrane ruffles as well as vesicle trafficking [38].
All of these findings points towards a macropinocytosis-like pathway as the primary internalization method of Ebola. They also indicate a role of actin in viral entry and suggest that the virus can actively promote localized actin remodeling through its interaction with Arp2 and VASP. The mechanism by which the virus causes macropinocytosis is not understood, but most likely involves the interaction of GP with cell surface receptors. The receptor tyrosine kinase Ax1 and integrin βI have been implicated as viral receptors, with evidence that several other receptor tyrosine kinases and integrins can trigger macropinocytosis [38].
Filovirus Transcription
During transcription, the RNA genome is transcribed into seven monocistronic mRNAs whose length is determined by highly conserved start and stop signals. These start signals are predicted to form stable stem-loop structures [40]. Just as in other negative sense RNA viruses, the transcription process begins with the binding of the polymerase complex to a single binding site located within the leader region of the genome. The complex then slides along the RNA template and sequentially transcribes the individual genes in their 3’ to 5’ order. However, the polymerase is released from the template following mRNA formation, so reinitiation at downstream genes is attenuated. Thus, the first gene, NP, is transcribed at the highest levels, whereas the last gene, L, is transcribed at the lowest [11].
VP30 is assumed to be a transcription activation factor that is essential for the viral life cycle. While the mechanism is not completely understood, it is suggested that this protein is involved in initiation because VP30-dependent transcription is regulated by RNA of the first transcription start signal. The first 23 nucleotides of this NP mRNA are involved in stem-loop structure formation which might interfere with the progression of transcription by physically hampering polymerase movement [52]. However, the N-terminus of VP30 contains a Zn+2 binding Cys-His motif [31] and is rich in basic amino acids, allowing it to directly interact with RNA [23]. Therefore, the protein could either resolve or cover this secondary structure and allow transcription to proceed. Further research has also shown that VP30 is important in transcription reinitiation, and may bind stem-loops formed by the promoter of each Ebola virus gene [29].
Interestingly, VP24 has also been shown to inhibit transcription and replication of the Ebola virus genome. While the exact mechanism has not yet been elucidated, it is possible that VP24 binds to NP and hampers the function of VP35, VP30 or L. This interference could be important in converting the viral genome from a transcription or replication competent form to one that is ready for virion assembly and egress [50].
Disruption of Cell Adhesion
Expression of Ebola GP in cultured cells causes a disruption in cell adhesion that results in a loss of cell-cell contacts, as well as a loss of attachment to the culture substrate. The effects of GP are caused by the mucin domain, a highly glycosylated region of GP1 composed of roughly 150 amino acids and containing numerous N- and O- linked glycosylation sites (Fig 18). While this loss of endothelial cell attachment is key for the characteristic hemorrhaging, only recently has a mechanism for the disturbance of cell adhesion been proposed [16].
For years, staining by flow cytometry has associated Ebola infection with a reduction in membrane levels of β1 integrin, a receptor that mediates attachment with the extracellular matrix, as well as major histocompatibility complex 1 (MHC1), a molecule important in immune system recognition [44]. While these effects were previously assumed to result from removal of surface proteins, Francica et al. propose that GP-mediated loss of surface protein recognition occurs via steric shielding of surface epitopes [16].
Observations that this down regulation is relieved by enzymatic removal of carbohydrate modification suggests that the steric occlusion is mediated by N- and O-linked modification of GP. In fact, the O-linked glycosylation of the mucin domain may promote an extended conformation that allows this domain to serve as a 150 residue long flexible rod that can mask domains in the immediate vicinity. Inherent in this mechanism is the fact that GP must localize in close proximity to the affected proteins, possibly explaining the variety of cell receptors regulated by this viral protein. This mechanism also helps the virus in immune system avoidance. The ability of GP to mask MHC1 may be a strategy for avoiding CD8 T cell-mediated killing of Ebola infected cells [16].
Viral Budding
The VP40 matrix protein is a necessary for virion egress, and can actually form virus-like particles that bud from the cell surface in the absence of all other viral proteins (Fig 19). This process is mediated by viral L-domains in the N-terminus of the protein that interact with host factors such as Nedd4 and Tsg101. The human ubiquitin ligase Nedd4 is part of an ubiquitination enzyme cascade that covalently links multiple ubiquitin copies to VP40. In contrast, Tsg101 is involved in selecting ubiquitylated cargo for incorporation in multi-vesicular bodies that are subsequently secreted by the cell [46].
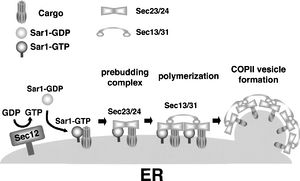
However, VP40 has to be transported to the plasma membrane before it can induce budding. Recent evidence suggests that the viral protein utilizes the COPII transport system through interaction with host protein Sec24C during intracellular transport to the plasma membrane (Fig 20). This system is responsible for vesicular transport from the ER to the Golgi, so VP40 most likely encounters Sec24C at an early point in its intracellular transport. However, the process of cargo transport to the plasma membrane as well as the process of VP40 dissociation from the complex must still be clarified [54].
While the plasma membrane was initially viewed as a randomly arranged lipid bilayer, it has recently been shown that this cell barrier is much more complex in nature. Certain cholesterol enriched regions known as lipid rafts have been implicated in such diverse functions as channeling external signals into specific second messenger pathways [8] and mediating cell-cell interactions [32]. These processes are possible because these regions concentrate relevant molecules in specialized microenvironments. While these properties are certainly important for cell function, these lipid platforms also serve as a coordination site for the intimate reactions of Ebola virus proteins required for assembly and budding. This increases the efficiency of virus budding, while lowering the chance that defective, noninfectious particles will be released [3].
These rafts may also impact viral pathogenicity as well as host immune response. Cell signaling molecules normally located in lipid rafts can be incorporated into the envelope of budding virions and used to ensure more efficient replication by affecting biochemical processes of newly infected cells [3]. Adhesion proteins could enhance viral entry, while incorporation of certain glycosylphosphatidylinositol (GP1) linked proteins can inhibit the complement pathways CD55 and CD59, which help the virus evade complement-mediated lysis [39].
Connecting the Cellular Mechanism to the Hemorrhagic Fever
The Ebola virus is normally transmitted by direct contact with infected body fluids or skin/mucus membrane contact. Once inside the body, the virus attacks macrophages and monocytes, relying upon host antibodies and complement component 1 for efficient infection [45]. The white blood cells respond by releasing large amounts of proinflammatory cytokines that increase permeability of the vascular endothelium, which facilitates easier entry into the virus's secondary targets, endothelial cells (Fig 20). These cytokines also recruit more macrophages to the area, maximizing the number of cells that Ebola can use to spread throughout the body. In the meantime, hepatocytes are being destroyed by the virus, ensuring that these cell signals cannot be cleared from the bloodstream [17].
Following GP-mediated receptor binding, Ebola virions are taken into endothelial cells via macropinocytosis. After their formation, macropinosomes move further into the cytoplasm to acquire new markers or fuse with other vesicles of the standard endolysosomal pathway. This eventually moves the Ebola virions to more acidic compartments such as early and late endosomes that assist in the pH dependent fusion of viral and cellular membranes [38]. During this process, the cell detaches from its neighbors and loses contact with its basement membrane thanks to a mechanism of glycan mediated steric occlusion by GP [16]. The newly created particles then leave via lipid rafts, leaving a destabilized vascular system responsible for the massive blood loss characteristic of Ebola patients [3].
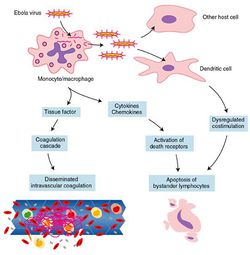
Meanwhile, the immune system is going haywire (Fig 21). Interferons are not being made because VP35 interferes with nearly every step in the process [2]. White blood cells are trapped inside the circulatory system because sGP limits their movement [49]. Macrophages and monocytes are releasing a cocktail of proinflammatory cytokines that destroy the vascular endothelium, but also activate the coagulation cascade. This puts your body in a paradoxical state in which you can die of hypovolemic shock from massive hemorrhage, or from catastrophic thrombosis, the formation of blood clots around the body [17].
References
1) Basler, C.F., Wang, X., Mühlberger, E., Volchkov, V., Paragas, J., Klenk, H.D., Garcia-Sastre, A. & Palese, P. (2000). The Ebola virus VP35 protein functions as a type I IFN antagonist. PNAS, 97, 12289-12294.
2) Basler, C.F., Mikulasova, A., Martinez-Sobrido, L., Paragas, J., Mühlberger, E., Bray, M., Klenk, H.D., Palese, P. & García-Sastre, A. (2003). The Ebola Virus VP35 Protein Inhibits Activation of Interferon Regulatory Factor 3. Journal of Virology, 77(14), 7945-7956.
3) Bavari, S., Bosio, C.M., Wiegand, E., Ruthel, G., Will, A.B., Geisbert, T.W., Hevey, M., Schmaljohn, C., Schmaljohn, A. & Aman, M.J. (2002). Lipid Raft Microdomains: A Gateway for Compartmentalized Trafficking of Ebola and Marburg Viruses. The Journal of Experimental Medicine, 195(5), 593-602.
4) Bhattacharyya, S., Warfield, K.L., Ruthel, G., Bavari, S., Aman, M.J. & Hope, T.J. (2010). Ebola virus uses clathrin-mediated endocytosis as an entry pathway. Virology, 401, 18-28.
5) Borio, L., Inglesby, T., Peters, C.J., Schmaljohn, A.L., Hughes, J.M., Jahrling, P.B., Ksiazek, T., Johnson, K.M., Meyerhoff, A., O’Toole, T., Ascher, M.S., Bartlett, J., Breman, J.G., Eitzen, Jr., E.M., Hamburg, M., Hauer, J., Henderson, D.A., Johnson, R.T., Kwik, G., Layton, M., Lillibridge, S., Nabel, G.J., Osterholm, M.T., Perl, T.M., Russell, P. & Tonat, K. (2002). Hemorrhagic fever viruses as biological weapons: medical and public health management. Journal of the American Medical Association, 287, 2391-2405.
6) Cárdenas, W.B., Loo, Y.M., Gale, M., Hartman, A.L., Kimberlin, C.R., Martínez-Sobrido, L., Saphire, E.O. & Basler, C.F. (2006). Ebola Virus VP35 Protein Binds Double-Stranded RNA and Inhibits Alpha/Beta Interferon Production Induced by RIG-I Signaling. Journal of Virology, 80(11), 5168-5178.
7) CDC Special Pathogens Branch. (2010). Ebola Hemorrhagic Fever Case Count and Location List. Ebola Hemorrhagic Fever Information Packet.
8) Cheng, P.C., Cherukuri, A., Dykstra, M., Malapati, S., Sproul, T., Chen, M.R. & Pierce, S.K. (2001). Floating the raft hypothesis: the roles of lipid rafts in B cell antigen receptor function. Semininars in Immunology, 13, 107-114.
9) Crary, S.M., Towner, J.S., Honig, J.E., Shoemaker, T.R. & Nichol, S.T. (2003). Analysis of the role of predicted RNA secondary structures in Ebola virus replication. Virology, 306(2), 210-218.
10) Doherty, G.J. & McMahon, H.T. (2009). Mechanisms of Endocytosis. Annual Review of Biochemistry, 78, 857–902.
11) Emerson, S.U. (1982). Reconstitution studies detect a single polymerase entry site on the vesicular stomatitis virus genome. Cell, 31, 635-642.
12) Empig, C.J. & Goldsmith, M.A. (2002). Association of the caveola vesicular system with cellular entry by filoviruses. Journal of Virology, 76, 5266-5270.
13) Feldmann, H. & Geisbert, T.W. (2010). Ebola haemorrhagic fever. DOI:10.1016/S0140-6736(10)60667-8.
14) Feldmann, H., Klenk, H.D. & Sanchez, A. (1993). Molecular biology and evolution of filoviruses. Archives of Virology, Supplementum, 7, 81-100.
15) Feng, Z., Cerveny, M., Yan, Z. & He, B. (2007). The VP35 Protein of Ebola Virus Inhibits the Antiviral Effect Mediated by Double-Stranded RNA-Dependent Protein Kinas PKR. Journal of Virology, 81(1), 182-192.
16) Francica, J.R., Varela-Rohena, A., Medvec, A., Plesa, G., Riley, J.L. & Bates, P. (2010). Steric Shielding of Surface Epitopes and Impaired Immune Recognition Induced by the Ebola Virus Glycoprotein. PLOS Pathogens, 6(9), e1001098.
17) Geisbert, T., Hensley, L., Larsen, T., Young, H., Reed, D., Geisbert, J., Scott, D., Kagan, E., Jahrling, P. & Davis, K. (2003). Pathogenesis of Ebola Hemorrhagic Fever in Cynomolgus Macaques. American Journal of Pathology, 163, 2347-2370.
18) Hoenen, T., Groseth, A., Kolesnikova, L., Theriault, S., Ebihara, H., Hartlieb, B., Bamberg, S., Feldmann, H., Ströher, U. & Becker, S. (2006). Infection of Naïve Target Cells with Virus-Like Particles: Implications for the Function of Ebola Virus VP24. Journal of Virology, 80(14), 7260-7264.
19) Huang, Y., Xu, L., Sun, Y. & Nabel, G.J. (2002). The assembly of Ebola virus nucleocapsid requires virion-associated proteins 35 and 24 and posttranslational modification of nucleoprotein. Molecular Cell, 10, 307-316.
20) Ito, H., Watanabe, S., Takada, A. & Kawaoka, Y. (2001). Ebola Virus Glycoprotein: Proteolytic Processing, Acylation, Cell Tropism, and Detection of Neutralizing Antibodies. Journal of Virology, 75(3), 1576-1580.
21) Jasenosky, L.D., Neumann, G., Lukashevich, I. & Kawaoka, Y. (2001). Ebola Virus VP40-Induced Particle Formatin and Association with the Lipid BIlayer. Journal of Virology, 75(11), 5205-5214.
22) Jin, H.L., Yan, Z.P., Prabhakar, B.S., Feng, Z.D., Ma, Y.J., Verpooten, D., Ganesh, B. & He, B. (2010). The VP35 protein of Ebola virus impairs dendritic cell maturation induced by virus and lipopolysaccharide. Journal of General Virology, 91, 352-361.
23) John, S.P., Wang, T., Steffen, S., Longhi, S., Schmaljohn, C.S. & Jonsson, C.B. (2007). Ebola Virus VP30 Is an RNA Binding Protein. Journal of Virology, 81(17), 8967-8976.
24) Johnson, K.M. & Breman, J.G. (1978) Ebola haemorrhagic fever in Zaire, 1976. Bulletin of the World Health Organization, 56(2), 271-293.
25) Klenk, H.-D. & Feldmann, H. (2004). Ebola and Marburg Viruses: Molecular and Cellular Biology. Wymondham: Horizon Bioscience.
26) Kumar, K.P., McBride, K.M., Weaver, B.K., Dingwall, C. & Reich, N.C. (2000). Regulated nuclear-cytoplasmic localization of interferon regulatory factor 3, a subunit of double-stranded RNA-activated factor 1. Molecular and Cellular Biology, 20, 4159–4168.
27) Lee, J.E., Fusco, M.L., Hessell, A.J., Oswald, W.B., Burton, D.R. & Saphire, E.O. (2008). Structure of the Ebola virus glycoprotein bound to an antibody from a human survivor. Nature, 454, 177-181.
28) Leffel, E.K. & Reed, D.S. (2004). Marburg and Ebola Viruses as Aerosol Threats. Biosecurity and Bioterrorism: Biodefense Strategy, Practice, and Science, 2(3), 186-191.
29) Martínez, M.J., Biedenkopf, N., Volchkova, V., Hartlieb, B., Alazard-Dany, N., Reynard, O., Becker, S. & Volchkov, V. (2008). Role of Ebola Virus VP30 in Transcription Reinitiation. Journal of Virology, 82(24), 12569-12573.
30) Mercer, J. & Helenius, A. (2009). Virus entry by macropinocytosis. Nature Cell Biology, 11, 510-520.
31) Modrof, J., Becker, S. & Muhlberger, E. (2003). Ebola virus transcription activator VP30 is a zinc-binding protein. Journal of Virology, 77, 3334-3338.
32) Moran, M. & Miceli, M.C. (1998). Engagement of GPI-linked CD48 contributes to TCR signals and cytoskeletal reorganization: a role for lipid rafts in T cell activation. Immunity, 9, 787-796.
33) Mühlberger, E.,Weik, M., Volchkov, V.E., Klenk, H.D. & Becker, S. (1999). Comparison of the transcription and replication strategies of Marburg virus and Ebola virus by using artificial replication systems. Journal of Virology, 73, 2333-2342.
34) Pratt, W.D., Wang, D., Nichols, D.K., Luo, M., Woraratanadharm, J., Dye, J.M., Holman, D.H. & Dong, J.Y. (2010). Protection of Nonhuman Primates against Two Species of Ebola Virus Infection with a Single Complex Adenovirus Vector. Clinical and Vaccine Immunology, 17(4), 572-581.
35) Prins, K.C., Cárdenas, W.B. & Basler, C.F. (2009). Ebola Virus Protein VP35 Impairs the Function of Interferon Regulatory Fact-Activating Kinases IKKε and TBK-1. Journal of Virology, 83(7), 3069-3077.
36) Quinn, K., Brindley, M.A., Weller, M.L., Kaludov, N., Kondratowicz, A., Hunt, C.L., Sinn, P.L., McCray, P.B., Stein, C.S., Davidson, B.L., Flick, R., Mandell, R., Staplin, W., Maury, W. & Chiorini, J.A. (2009). Rho GTPases Modulate Entry of Ebola Virus and Vesicular Stomatitis Virus Pseudotyped Vectors. Journal of Virology, 83(19), 10176-10186.
37) Reid, S.P., Leung, L.W., Hartman, A.L., Martinez, O., Shaw, M.L., Carbonnelle, C., Volchkov, V.E., Nichol, S.T. & Basler, C.F. (2006). Ebola Virus VP24 Binds Karyopherin α1 and Blocks STAT1 Nuclear Accumulation. Journal of Virology, 80(11), 5156-5167.
38) Saeed, M.F., Kolokoltsov, A.A., Albrecht, T. & Davey, R.A. (2010). Cellular Entry of Ebola Virus Involves Uptake by a Macropinocytosis-Like Mechanism and Subsequent Trafficking through Early and Late Endosomes. PLOS Pathogens, 6(9).
39) Saifuddin, M., Hedayati, T., Atkinson, J.P., Holguin, M.H., Parker, C.J. & Spear, G.T. (1997). Human immunodeficiency virus type 1 incorporates both glycosyl phosphatidylinositol-anchored CD55 and CD59 and integral membrane CD46 at levels that protect from complement-mediated destruction. Journal of General Virology, 78, 1907-1911.
40) Sanchez, A., Kiley, M.P., Holloway, B.P. & Auperin, D.D. (1993). Sequence analysis of the Ebola virus genome: organization, genetic elements, and comparison with the genome of Marburg virus. Virus Research, 29, 215-240.
41) Schümann, M., Gantke, T. & Mühlberger, E. (2009). Ebola Virus VP35 Antagonizes PKR Activity through Its C-Terminal Interferon Inhibitory Domain. Journal of Virology, 83(17), 8993-8997.
42) Schornberg, K., Matsuyama, S., Kabsch, K., Delos, S., Bouton, A. & White, J. (2006). Role of endosomal cathepsins in entry mediated by the Ebola virus glycoprotein. Journal of Virology, 80(8), 4174-4178.
43) Simmons, G., Wool-Lewis, R., Baribaud, F., Netter, R. & Bates, P. (2002). Ebola Virus Glycoproteins Induce Global Surface Protein Down-Modulation and Loss of Cell Adherence. Journal of Virology, 76(5), 2518-2528.
44) Sullivan, N., Peterson, M., Yang, Z., Kong, W., Duckers, H., Nabel, E. & Nabel, G. (2005). Ebola Virus Glycoprotein Toxicity Is Mediated by a Dynamin-Dependent Protein-Trafficking Pathway. Journal of Virology, 79(1), 547-553.
45) Takada, A., Feldmann, H., Ksiazek, T. & Kawaoka, Y. (2003). Antibody-Dependent Enhancement of Ebola Virus Infection. Journal of Virology, 77(13), 7539–7544.
46) Timmins, J., Schoehn, G., Ricard-Blum, S., Scianimanico, S., Verent, T., Ruigrok, R.W.H. & Weissenhorn, W. (2003). Ebola Virus Matrix Protein VP40 Interaction with Human Cellular Factors Tsg101 and Nedd4. Jouranl of Molecular Biology, 326, 493-502.
47) Volchkov, V.E., Becker, S., Vochkova, V.A., Ternovoj, V.A., Kotov, A.N., Netesov, S.V. & Klenk, H.D. (1995). GP mRNA of Ebola virus is edited by the Ebola virus polymerase and by T7 and vaccinia virus polymerases. Virology, 214, 421-430.
48) Volchkov, V.E., Feldmann, H., Volchkova, V.A., & Klenk, H.D. (1998). Processing oft eh Ebola virus glycoprotein by the proprotein convertase furin. PNAS, 95, 5762-5767.
49) Wahl-Jensen, V., Afanasieva, T., Seebach, J., Ströher, U., Feldmann, H. & Schnittler, H. (2005). Effects of Ebola Virus Glycoproteins on Endothelial Cell Activation and Barrier Function. Journal of Virology, 79(16), 10442-10450.
50) Watanabe, S., Noda, T., Halfmann, P., Jasenosky, L. & Kawaoka, Y. (2007). Ebola Virus (EBOV) VP24 Inhibits Transcription and Replication of the EBOV Genome. The Journal of Infectious Diseases, 196, S284-290.
51) Weed, S.A. & Parsons, J.T. (2001). Cortactin: coupling membrane dynamics to cortical actin assembly. Oncogene, 20, 6418-6434.
52) Weik, M., Modrof, J., Klenk, H.D., Becker, S. & Mühlberger, E. (2002). Ebola Virus VP30-Mediated Transcription Is Regulated by RNA Secondary Structure Formation. Journal of Virology, 76(17), 8532-8539.
53) World Health Organization. (2010). Ebola Hemorrhagic Fever. Epidemic and Pandemic Alert and Response (EPR).
54) Yamayoshi, S., Noda, T., Ebihara, H., Goto, H., Morikawa, Y., Lukashevich, I.S., Neumann, G., Feldmann, H. & Kawaoka, Y. (2008). Ebola Virus Matrix Protein VP40 Uses the COPII Transport System for Its Intracellular Transport. Cell Host and Microbe, 3, 168-177.
55) Yonezawa, A., Cavrois, M. & Greene, W.C. (2005). Studies of Ebola Virus Glycoprotein-Mediated Entry and Fusion by Using Pseudotyped Human Immunodeficiency Virus Type 1 Virions: Involvement of Cytoskeletal Proteins and Enhancement by Tumor Necrosis Factor Alpha. Journal of Virology, 79(2), 918-926.
Page authored for BIOL 375 Virology, September 2010