Lactobacillus casei: Difference between revisions
| Line 30: | Line 30: | ||
==Description and significance== | ==Description and significance== | ||
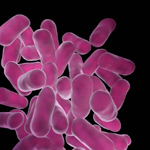
''Lactobacillus casei'' is one of the many species of bacteria belonging in the genus ''Lactobacillus''. It is a mesophilic bacteria that is gram positive, rod shaped, nonsporing, nonmotile, anaerobic, and contains no cytochromes (5). ''L. casei'' can be found in various environments such as raw and fermented dairy products, intestinal tracts and reproductive systems of humans and animals, and fresh and fermented plant products (1). The optimum pH for ''L. casei'' is 5.5. The lactic acid produced by ''L. casei'' through fermentation is very important since it can be used to make cheeses and yogurts, reduce cholesterol levels, enhance immune response, control diarrhea, alleviate lactose intolerance, inhibit intestinal pathogens, and serve as probiotics (8). Probiotics are viable microorganisms that promote or support a beneficial balance of microbes living in the gastrointestinal tract (5). In order for the probiotics to carry out their functions, the probiotic live cells must not be lower than 10^6-10^7 cfu/g (9). | |||
There are many strains/isolates of ''L.casei'' from different origins and geographical locations. That is why molecular typing of ''L.casei'' is crucial to understanding the evolutionary adaptation of this species to different ecological niches (1). Another reason for having ''L. casei's'' genome sequenced is to determine the phylogenetic relationships between various groups of bacteria in ''Lactobacillus''. ''L. casei, L. paracasei, L. rhamnosus'', and ''L. zeae'' form a closely related taxonomic group within ''Lactobacillus'' (3). Sometimes the classification ''L. casei'' is loosely applied to strains of any of these species by commercial companies. By having the genome sequenced, species boundaries could be drawn and names can be attached to those species (3). | |||
==Genome structure== | ==Genome structure== | ||
Revision as of 11:33, 29 August 2007
A Microbial Biorealm page on the genus Lactobacillus casei
Classification
Higher order taxa
Bacteria; Firmicutes; Bacilli; Lactobacillales; Lactobacillaceae; Lactobacillus
Species
|
NCBI: Taxonomy |
Lactobacillus casei
Strains
Listed are some of the many strains/isolates of Lactobacillus casei: (1)
HUMAN origin: L3, L6, L9, L14, L19, L25, L30, CRF28, Shirota
CHEESE origin: ASCC 428, ASCC 477, ASCC 1087, ASCC 1088, ASCC 1123, DPC 3971, DPC 3968, DPC 4249, DPC 4748
PLANT origin: 12A, 32G, BI0231, USDA-P
Description and significance
Lactobacillus casei is one of the many species of bacteria belonging in the genus Lactobacillus. It is a mesophilic bacteria that is gram positive, rod shaped, nonsporing, nonmotile, anaerobic, and contains no cytochromes (5). L. casei can be found in various environments such as raw and fermented dairy products, intestinal tracts and reproductive systems of humans and animals, and fresh and fermented plant products (1). The optimum pH for L. casei is 5.5. The lactic acid produced by L. casei through fermentation is very important since it can be used to make cheeses and yogurts, reduce cholesterol levels, enhance immune response, control diarrhea, alleviate lactose intolerance, inhibit intestinal pathogens, and serve as probiotics (8). Probiotics are viable microorganisms that promote or support a beneficial balance of microbes living in the gastrointestinal tract (5). In order for the probiotics to carry out their functions, the probiotic live cells must not be lower than 10^6-10^7 cfu/g (9).
There are many strains/isolates of L.casei from different origins and geographical locations. That is why molecular typing of L.casei is crucial to understanding the evolutionary adaptation of this species to different ecological niches (1). Another reason for having L. casei's genome sequenced is to determine the phylogenetic relationships between various groups of bacteria in Lactobacillus. L. casei, L. paracasei, L. rhamnosus, and L. zeae form a closely related taxonomic group within Lactobacillus (3). Sometimes the classification L. casei is loosely applied to strains of any of these species by commercial companies. By having the genome sequenced, species boundaries could be drawn and names can be attached to those species (3).
Genome structure
Describe the size and content of the genome. How many chromosomes? Circular or linear? Other interesting features? What is known about its sequence? Does it have any plasmids? Are they important to the organism's lifestyle?
Cell structure and metabolism
Describe any interesting features and/or cell structures; how it gains energy; what important molecules it produces.
Ecology
Describe any interactions with other organisms (included eukaryotes), contributions to the environment, effect on environment, etc.
Pathology
L. casei is generally considered nonpathogenic and safe. However, cases of sepsis, meningitis, and infections localized in organs have been reported (10).
Application to Biotechnology
Does this organism produce any useful compounds or enzymes? What are they and how are they used?
Current Research
Enter summaries of the most recent research here--at least three required
References
1. Cai, H., Rodriguez, B.T., Zhang, W., Broadbent, J.R., and Steele, J.L. “Genotypic and phenotypic characterization of Lactobacillus casei strains isolated from different ecological niches suggests frequent recombination and niche specificity”. Microbiology. 2007. Volume 153. p. 2655-2665.
2. Chan-Blanco,Y., Bonilla-Leiva, A.R., and Velazquez, A.C. “Using banana to generate lactic acid through batch process fermentation”. Applied Microbiology Biotechnology. 2003. Volume 63. p. 147-152.
3. Desai, A.R., Shah, N.P., and Powell, I.B. “Discrimination of Dairy Industry Isolates of the Lactobacillus casei group”. Journal of Dairy Science. 2006. Volume 89. p. 3345-3351.
4. Ha, M.Y., Kim, S.W., Lee, Y.W., Kim, M.Y., and Kim, S.J. "Kinetics analysis of growth and lactic acid production in pH-controlled batch cultures of Lactobacillus casei KH-1 using yeast extract/corn steep liquor/glucose medium. Journal of Bioscience and Bioengineering. 2003. Volume 96. p. 134-140.
5. Holzapfel, W.H., Haberer, P., Geisen, R., Bjorkroth, J., and Schillinger, U. “Taxonomy and important features of probiotic microorganisms in food and nutrition”. The American Journal of Clinical Nutrition. 2001. Volume 73. p. 365S-373S.
6. Marcos, A., Warnberg, J., Nova, E., Gomez, S., Alvarez, A., Alvarez, R., Mateos, J.A., and Cobo, J.M. “The effect of milk fermented by yogurt cultures plus Lactobacillus casei DN-114001 on the immune response of subjects under academic examination stress”. European Journal of Nutrition. 2004. Volume 43. p. 381-389.
7. Millette, M., Luquet, F.M., and Lacroix, M. "In vitro growth of selected pathogens by Lactobacillus acidophilus- and Lactobacillus casei- fermented milk". Letters in Applied Microbiology. 2007. Volume 44. p. 314-319.
8. Mishra, V. and Prasad, D.N. “Application of in vitro methods for selection of Lactobacillus casei strains as potential probiotics”. International Journal of Food Microbiology. 2005. Volume 103. p. 109-115.
9. Nebesny, E., Zyzelewicz, D., Motyl, I., and Libudzisz, Z. "Dark chocolates supplemented with Lactobacillus strains". European Food Resource Technology. 2007. Volume 225. p. 33-42.
10. Salvatore, S., Hauser, B., Devreker, T., Vierira, M.C., Luini, C., Arrigo, S., Nespoli, L., and Vandeplas, Y. “Probiotics and zinc in acute infectious gastroenteritis in children: are they effective?”. Nutrition. 2007. Volume 23. p. 498-506.
11. Sauvageot, N., Beaufils, S., Maze, A., Deutscher, J., and Hartke, A. “Cloning and characterization of a gene encoding a cold-shock protein in Lactobacillus casei”. FEMS Microbiology Letters. 2006. Volume 254. p. 55-62.
12. Takeda, K. and Okumura, K. “Effects of a fermented milk drink containing Lactobacillus casei strain Shirota on the human NK-cell activity”. The Journal of Nutrition. 2007. Volume 137. p. 791S-793S.
13. Ventura, M., Canchaya, C., Bernini, V., Altermann, E., Barrangou, R., McGrath, S., Claesson, M.J., Li, Y., Leahy, S., Walker, C.D., Zink, R., Neviani, E., Steele, J., Broadbent, J., Klaenhammer, T.R., Fitzgerald, G.F., O’Toole, P.W., and van Sinderen, D. “Comparative genomics and transcriptional analysis of prophages identified in the genomes of Lactobacillus gasseri, Lactobacillus salivarius, and Lactobacillus casei”. Applied and Environmental Microbiology. 2006. Volume 72. p. 3130-3146.
14. Yoon, K.Y., Woodams, E.E., and Hang, Y.D. “Production of probiotic cabbage juice by lactic acid bacteria”. Bioresource Technology. 2006. Volume 97. p. 1427-1430.
Edited by Alison Wong, student of Rachel Larsen