Lactobacillus plantarum and its biological implications
Introduction
By Katie Adlam
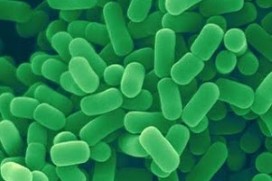
Lactobacillus plantarum (L. plantarum) is a rod-shaped, gram-positive lactic acid bacterium. It is commonly found in the human and other mammalian gastrointestinal tracts, saliva, and various food products. It can grow at temperatures between 15-45˚C and at pH levels as low as 3.2 [1]. L. plantarum is a facultative heterofermentative [2,3] that ferments sugars to produce lactic acid, ethanol or acetic acid, and carbon dioxide under certain conditions and selective substrates. Depending on the carbon source, these bacteria can switch from using heterofermentative and homofermentative ways of metabolism. This bacterium is acid and bile salt tolerant, which allows it to survive the passage through the gastrointestinal tract of humans. L. plantarum is of current interest to researchers and the food industry since it is considered a safe probiotic. It can help limit the amount of pathogenic bacteria or diseases that can have a negative impact on humans. In addition, recent research indicates that L. plantarum can be used as a vaccine vehicle.
Genome
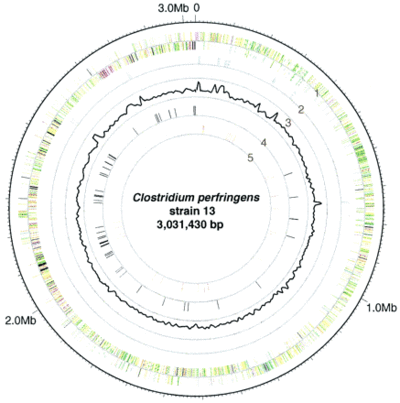
L. plantarum has a relatively large genome compared to other Lactobacillus spp. Its genome consists of a 3.3 Mb circular chromosome, which is the largest sequenced genome of any lactic acid bacteria [2]. The genome of L. plantarum consists of five rRNA operons, which are evenly distributed around the chromosome (FIGURE 2) [1]. A total of 62 tRNA encoding genes have been found and are in relation to some of the rRNA clusters. In addition, the genome encodes two classes of transposase regions, which are thought to encode mobile genetic elements [1].
The genome consists of 3,052 protein-encoding genes and only 39 of these genes are pseudogenes [1]. L. plantarum proteins are very similar to other Gram-positive bacteria since they have low GC content, have a peptidoglycan cell wall, and are organized collinearly. L. plantarum genome contains genes for the Embden-Meyerhoff-Parnas (EMP) pathway. Some of the genes correlate to enzymes that break down pentoses and hexoses. Furthermore, its genome shows that its genes encode for phosphotransferase, mannose, and fructose transport systems [1].
Metabolism
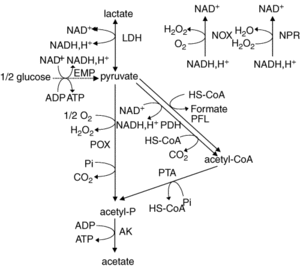
L. plantarum is a facultative anaerobe or a facultative heterofermentative, as it can grow in the presence and absence of oxygen. It has enzymes for fermentation in the Embden-Meyerhoff-Parnas pathway (Figure 3) and through the phosphoketolase pathway. This allows the bacterium to perform homolactic and heterolactic fermentation [1]. When oxygen is not present and this bacterium undergoes homolactic-like fermentation, L. plantarum converts carbon sources to D- and L- configurations of lactate and alcohols via lactate dehydrogenase. This normally occurs during its growth phase. It is important to note that L. plantarum can use a variety of carbon sources, which is hypothesized to be the result of horizontal gene transfer.
Under aerobic conditions, lactate is converted to acetate and one ATP is produced via lactate dehydrogenase, pyruvate oxidase, and acetate kinase. In addition to producing acetate, this pathway also forms hydrogen peroxide and carbon dioxide as byproducts. Hydrogen peroxide is formed by the conversion of oxygen via a manganese-dependent process. The production of this oxygen derived radical species can kill off other bacteria. Manganese-dependent processes use manganese as a catalase to lower the concentration of oxygen, which is beneficial to the aerotolerant bacterium [4]. L. plantarum can produce a variety of products from fermentation due to a large “pyruvate-dissipating potential” [1]. In addition, research has found that L. plantarum has a fumarate reductase, which implies that this bacterium has a basic electron transport chain for anaerobic bacteria [1]. It is proposed that this bacterium possesses a nitrate reduction system using nitrate as an electron acceptor. Furthermore, it does not contain the citric acid cycle; thus, does not generate activity through oxidation of acetate.
Biosynthesis and Secretion
L. plantarum, typically found in protein-rich environments like yogurt, has uptake systems for peptides. Once these peptides are ingested, various types of peptidases degrade them inside the bacterium. L. plantarum has 19 different genes encoding peptidases that have different specific functions, three of which can cleave N-terminal proline residues. Even though this bacterium has protein degradation mechanisms, it is still capable of producing most amino acids, except branched-chain amino acids such as valine, leucine, and isoleucine [1].
In addition to synthesizing amino acids, L. plantarum can perform a mRNA independent nonribosomal peptide synthesis [1]. Before this discovery, no other lactic acid bacteria was known to use this biosynthesis machinery. The L. plantarum nonribosomal peptide synthesis cluster contains two nonribosomal synthesis proteins, an important phosphopantetheinyl transferase, and proteins that are needed for precursors for regulation, transport, and enzymes [1].
Amino acids and peptides in L. plantarum are transported mainly through 57 ATP-binding cassette (ABC) transporters, of which 27 are importers and 30 are exporters. Since this bacterium cannot synthesize its own branched chain amino acids, it is able to obtain them through these transporters. In addition, L. plantarum has a high number of sugar import systems and regulatory proteins, which contribute to its flexible and versatile state as a bacterium. It can grow by using different carbon sources and can adapt to a wide array of environments [1].
L. plantarum is known to be able to adapt to stressful environments such as those in the gastrointestinal tract with a low pH or high salt content (Figure 4). Its genome encodes proteases that can degrade abnormal or nonfunctional proteins, as well as heat and cold-shock proteins to save energy under stress and to survive different climates. In order to survive in acidic environments, this bacterium uses the FoF1-ATPase and sodium-proton pumps to help regulate and maintain the intracellular pH. Furthermore, it has alkaline shock proteins to assist in pH tolerance. In addition, L. plantarum has developed ways to deal with oxidative stress by having catalases, peroxidases, and reductases, as well as a high intracellular concentration of manganese ions (Mn2+) to scavenge oxygen radicals [1]. This bacterium is able to obtain and accumulate manganese ions due to the P-type manganese translocating ATPase.
In addition to being able to adapt to the environment, L. plantarum has surface proteins that allow it to interact with the environment. It has a Sec transport system that is comprised of SecA/SecE/SecG/SecY/YajC (no SecDF) and it also has other proteins involved in secretion such as peptidases [1]. In addition, there are genes that encode for a sortase. Sortase is an enzyme that recognizes and cleaves carboxy-terminal sequences to modify the surface proteins to interact with different surfaces and substrates for growth. Lastly, L. plantarum has a protein machinery that is used to bind and uptake DNA from the environment. Scientists hypothesize that L. plantarum acquired its ability to adapt to many different environments from this uptake machinery. Researchers also have found very large regions in the genome of L. plantarum containing unusual base composition when comparing it to closely related species. An example of this are the genes for sugar uptake and catabolism (1). This reflects how L. plantarum is a very versatile species that can adjust to its environment.
Food Production
L. plantarum is commonly used in dairy, meat, and plant fermentations, and is found in food products such as yogurt, cheese, kimchi, sauerkraut, sourdough, and pickles [3, 5, 6]. The bacterium gives food certain tastes and flavors depending on the balance between acetate (volatile) and lactate (nonvolatile) organic acids [2]. By using oxygen and aerating food, producers can control the amount of acetic acid that is produced as the final product of fermentation. An example of this is sourdough fermentation. The bacterium is mainly used for dough acidification, but it can play a role in the leavening process.
A specific example of a food item that is produced by L. plantarum fermentation is sauerkraut. Sauerkraut is spontaneously produced from cabbage through lactic acid bacteria that are acid tolerant homolactic fermenters [7]. However, the taste of sauerkraut varies depending on the type of bacteria used, substrate for fermentation, salt concentration, and temperature of the fermentation. Because of the variation, a study set out to find starter cultures that could be universally used to produce sauerkraut. Leuconostoc mesenteroides (L. mesenteroides) was used for the first phase of fermentation and L. plantarum for the second. L. mesenteroides provides a mild, pleasant aromatic flavor to the sauerkraut while L. plantarum gives it an acidic, vinegar taste. The researchers found that by using both bacteria the fermentation time was shortened compared to other fermentation processes [7]. In addition,L. plantarum being dominant in the acidic, second phase of the fermentation resulted in the inhibition on the growth of other microorganisms. These two starter bacteria improved the final product of sauerkraut by reducing the fermentation time and the growth of pathogenic microbes.
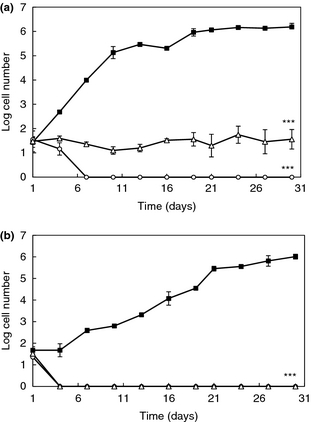
Natural preserved products are becoming increasingly desired by consumers. Because of this, scientists are studying antifungal lactic acid bacteria and their preservative properties. Since lactic acid bacteria typically can survive at wide range of temperatures and pH levels where some food spoilage yeasts are found, lactic acid bacteria are being studied to determine if they can inhibit growth of undesirable yeast. L. plantarum, a lactic acid bacterium, produces antifungal activity that can be substituted for potentially harmful preservatives that are found in food products [8]. A study in 2012 examined L. plantarum to determine if it had the ability to inhibit the growth of Rhodotorula mucilaginosa (R. mucilaginosa), a yeast that spoils dairy products and orange juice. After isolating and determining the viability of two strands of L. plantarum (strand 16 and 62), this group determined its affect on the activity of R. mucilaginosa by using the antiyeast bacteria as dairy starter adjuncts in yogurt and inoculants in orange juice. Inhibition of R. mucilaginosa by L. plantarum was tested against commercial, FDA approved preservatives such as sodium benzoate and potassium sorbate. The study demonstrated that L. plantarum, in fact, had a more significant impact on the inhibition of the growth of R. mucilaginosa compared to the FDA preservatives. It also decreased the rate of spoilage in both orange juice and yogurt (Figure 5) [8]. Since the inhibition of R. mucilaginosa was more effective than sodium benzoate and potassium sorbate, this bacterium has the potential to increase shelf life of yogurt and orange juice. However, it is important to note that the antifungal compounds produced by L. plantarum are unknown and currently being investigated.
Probiotics and Biotherapeutic Applications
According to the Merriam-Webster Dictionary, a probiotic is a dietary supplement that contains live bacteria or yeast and is used to maintain the normal gastrointestinal flora [9]. It is often taken orally and helps restore the flora after infection or antibiotic use. Some individuals, such as pregnant women, infants, and children with neurodevelopmental disorders, can benefit greatly from the use of probiotics [5].
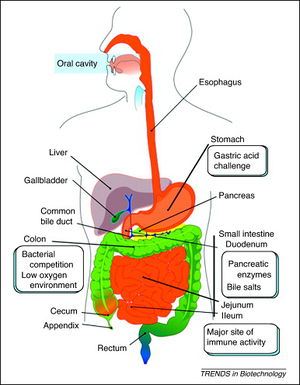
In order for probiotics to be effective, they must be able to reach their target site after ingestion (Figure 3). They have to travel through the gastrointestinal tract to the stomach. Low stomach pH could disrupt the proton motive force or enzyme function, consequently leading to a disruption of energy supply to the bacteria. In addition, probiotics must go through the small intestine that contains bile salts that can act as surfactants and can disturb cell membranes, DNA, and RNA [10]. Probiotics also must be somewhat resistant to antibiotics so that they can maintain the gastrointestinal tract homeostasis and aid in the recovery of intestinal microflora after treatment of antibiotics [11]. However, it is important to recognize that the antibiotic resistance genes could get transferred and form highly antibiotic resistant pathogenic bacteria.
L. plantarum is considered a probiotic because it secretes antimicrobial compounds, such as bacteriocin, that inhibit pathogenic gram-positive and gram-negative colonies from forming [5]. Bacteriocin, which is a Lactobacillus-inhibitory factor and toxin, inhibits the growth of similar bacteria and other antibiotic-like substances. L. plantarum also has a mannose-specific adhesion, which allows it to adhere to the epithelial lining in the human intestines and compete with both gram-positive and gram-negative pathogenic bacteria for nutrients [5,12]. These traits, in addition to its pH and temperature tolerance, make L. plantarum a potential probiotic to be used for therapy for those who suffer from gastrointestinal diseases such as Crohn’s Disease, inflammatory bowel disease (IBD), and colitis [5, 6, 11, 13, 14].
Gastrointestinal Tract
Probiotics help in lactose digestion and certain diseases such as Crohn’s Disease, inflammatory bowel disease (IBD), and colitis [5, 6, 11, 13, 14]. These conditions usually occur due to stress or homeostasis imbalances in the epithelial lining of the intestines. This leads to damage of the lining and increases permeability of the lining. Furthermore, this allows pathogenic bacteria to stimulate a mucosal immune response. The endotoxin and DNA from the pathogenic bacteria can cause an inflammatory response of the epithelial lining by bacterial translocation [14] and by bacterial translocation and stimulation of the release of cytokines and other pro-inflammatory mediators by endotoxins. For example, when tumor neurosis factor-α (TNF-α), a cytokine, becomes stimulated, it causes a potent interleukin (IL-8) to be released [15]. The secretion of the chemokine causes the movement of inflammatory cells into the lining. Furthermore, this response is mediated by an extracellular signal-regulated kinase (ERK). However, this inflammatory response can be repressed by species such as Lactobacilli, and specifically, L. plantarum [14].
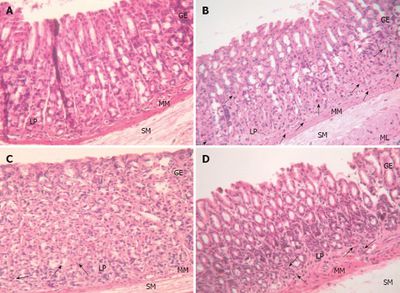
Current research is focused on determining the probiotic effects of L. plantarum on the epithelial lining of the intestines and how it can benefit humans [6, 11, 13, 14, 16]. A 2007 study established that L. plantarum could suppress an inflammatory response of the intestinal epithelial cells by inhibiting ERK activation of TNF-α, which could reduce the signaling of chemokines (IL-8) [15]. These results indicate that L. plantarum, as a probiotic, could help maintain the epithelial barrier and inhibit the inflammatory response (Figure 5). Furthermore, it was determined that L. plantarum could be used to prevent the development of colitis since it has a protective effect on tight junction (TJ) associated proteins and TJ ultra-structure [14]. It has been hypothesized that the TJs might form a barrier in the epithelial layer that could prevent bile from penetrating the blood stream [16]. In other words, L. plantarum is capable of providing protection on the intestinal epithelium by reducing the permeability of the intestinal lining and can reduce the amount of pro-inflammatory proteins in the gastrointestinal tract.
A specific bacterium that damages the mucosa and causes an inflammatory response by increasing TNF-α and interleukin is Helicobacter pylori (H. pylori). This bacterium colonizes in the human gastric mucosa and causes gastritis and ulcer diseases [6]. Its lipopolysaccharides (LPS) and surface proteins stimulate the production of TNF-α, interleukin, and reactive oxygen species (ROS). These do not stay localized to the area of infection. Instead, they spread throughout the body and cause damage. A study in 2012 showed that L. plantarum has anti-Helicobacter activity and can inhibit the growth of the pathogenic bacterium in acidic environments (pH 4) (Figure 6). Some fermentation products, such as lactic acid, inhibit the urease activity capability of H. pylori, and also inhibit Lactobacillus-inhibitory factors such as bacteriocin [6]. The anti-Helicobacter activity of L. plantarum can be used as a therapy for gastritis and other gastrointestinal tract diseases.
Vaccines
Recombinant strains of lactic acid bacteria are used to produce therapeutic proteins and to deliver these proteins to safe mucosal sites [17, 18]. These mucosal surfaces are a major site of pathogenic entry. Both systemic and mucosal immune responses can be induced at these surfaces [18]. They are also used to secrete other proteins such as interleukins and antibodies. In addition, mucosal vaccines can cause IgA secretion and a systemic immune response with T-cells to stimulate the immune system [19].
For example, bacteria expressing allergens provide an advantageous method of immunotherapy compared to subcutaneous and sublingual therapies [17, 20]. It is beneficial because the allergen will be protected from proteases since it is contained within the bacteria and is cost-effective since the antigen does not have to be processed. Also, the components of the allergen and the lactic acid bacteria are presented at the same time as the immune response. Studies have shown that certain strains of L. plantarum can be used as delivery vehicles with an allergen to prevent certain allergies, such as dust mite allergies [20]. If a recombinant strain has the dust mite allergen (Der p 1) for antigen delivery, it can reduce or prevent the stimulation of the ERK-pathway, which would lead to suppression of the chemokines that would cause inflammation.
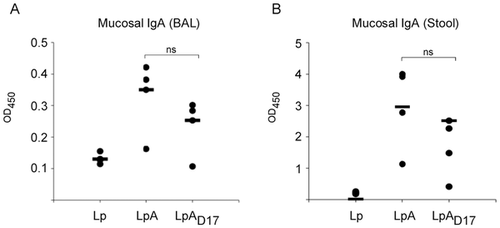
In addition, proinflammatory recombinant strains of L. plantarum are hypothesized to be effective mucosal delivery vehicles for vaccine antigens [18]. For example, a recent study used a strain of L. plantarum that expressed invasin from Yersinia pseudotuberculosis (Y. pseudotuberculosis). Invasin is a virulence factor that binds to β1-integrins on the surface of microfold cells (M cells) and stimulates the uptake of Y. pseudotuberculosis in the intestines. It can also cause inflammation of the host cell by activating the innate immune system [18]. By modifying L. plantarum so that the extracellular domain of invasin is anchored to the bacterial surface by N-terminal anchoring motifs, this strain of the bacterium can imitate the early infection symptoms of Y. pseudotuberculosis, leading to increase adjuvant characteristics. In other words, this strain of L. plantarum can, through the use of different N-terminal anchoring motifs, target Y. pseudotuberculosis invasin to the cell surface of the bacterium. Since this strain of L. plantarum can change the immunological tolerance of an immune response, it can be used to increase antigen immunogenicity and as a delivery vehicle for mucosal vaccines [18].
Lastly, strains of L. plantarum can be used to stimulate production of antibodies. A study using L. plantarum that expressed Borrelia burgdorferi (B. burgdorferi) OspA lipoprotein found that it stimulated the production of OspA-specific IgA and IgG antibodies as wells as pro- and anti-inflammatory cytokines (Figure 7) [19]. It did not lead to secretion of cytokine (IL-8) by epithelial cells and did not induce inflammatory effects. The recombinant L. plantarum is capable of stimulating a protective immune response via T-cell (Th1/Th2) mediated immunity.
Conclusion
L. plantarum is a bacterium that is very versatile and can adapt to various environmental conditions since it can ferment different types of carbohydrates and sugars. Specifically, it is able to withstand and grow in the harsh conditions of the gastrointestinal tract. Because of this, it can be used as a probiotic to benefit human health. A variety of strains or recombinants of L. plantarum can aid in restoring the homeostasis of the flora in the intestines, limit the amount of pathogenic bacteria, and could be potentially used as a vehicles for vaccines. However, not all the mechanisms for vaccinations with L. plantarum are known. As a result, many research groups are trying to determine possible mechanisms. Within the next few years, it is expected that more developments will be made to utilize the probiotic property of L. plantarum to benefit human health.
References
9. "Probiotic." Merriam-Webster. Merriam-Webster, 2013. Web. 22 Apr. 2013.
Edited by student of Joan Slonczewski for BIOL 238 Microbiology, 2013, Kenyon College.
