Leptospirosis: A Worldwide Zoonotic Disease
By Ian Veitch
Introduction
Leptospires are Gram-negative spirochetes that all share a distinctive cell structure consisting of a long, tight spiral that hooks at both ends (1, figure 1).
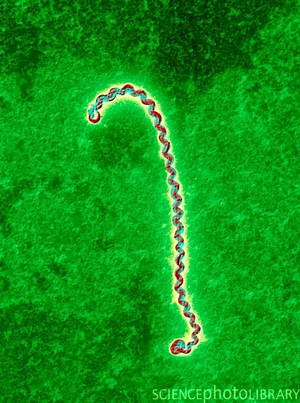
Leptospires are a well-known spirochete mainly because they cause leptospirosis, the most widespread zoonotic disease in the world that has been found in almost all species of mammals examined (2). Chronic carriers of leptospirosis are usually wild or domestic animals, such as rats, dogs, cattle, and pigs (3). Humans can contract the disease; however, leptospirosis in humans is always acquired from an animal source. Pathogenic leptospires reside in the proximal renal tubules of the kidneys of carriers and are subsequently excreted in urine. Urine tainted with leptospires can then contaminate soils, surface water, streams, and rivers (3). It is pertinent to note that not all leptospira species are pathogenic. There are twenty documented species in the genus Leptospira (figure 2). These twenty species are classified as pathogenic, non-pathogenic, or intermediate/opportunistic. While only nine of twenty species are labeled as pathogenic, more than 500,000 cases of severe leptospirosis are reported each year, with mortality rates exceeding 10% (4). Furthermore, the incidence of leptospirosis is higher in the tropics than in temperate regions because of longer survival of leptospires due to warm and humid conditions (5). Preventing and diagnosing leptospirosis can be a complicated process, partly due to the sheer diversity of bacteria within the genus Leptospira .
Leptospira Classification Systems
From a distance, all leptospira appear as spiral-shaped bacteria with distinctive end hooks. If examined closely, through various laboratory methods, the vast diversity of all leptospira bacteria can be appreciated. Two main classification systems have been implemented in an attempt to bring order to the sheer diversity found in the Leptospira genus.
DNA-based Classification
This form of classification is based on DNA-DNA relatedness established through hybridization (2). DNA-Based classification yielded twenty different species in the Leptospira genus (figure 2).
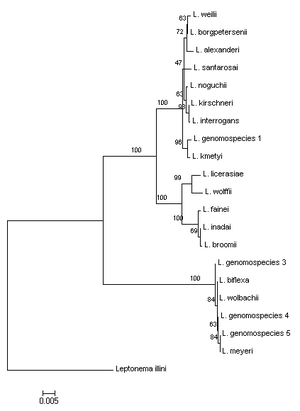
Nine species are pathogenic, five species are intermediates, and six species are saprophytic.
Serological Classification
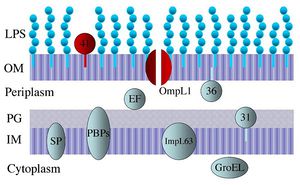
Serological classification organizes individual bacterial strains of a species into smaller groups based on their cell surface antigens. Antigens are foreign substances that induce an immune system response from the body, such as the heightened production of antibodies. Currently, there are more than two hundred serovars within the Leptospira genus. The basis for this method of classification lies in the cellular structure of leptospires. Like other Gram-negative bacteria, leptospires have a triple-layered cell envelope, which consists of an outer membrane, a peptidoglycan cell wall, and an inner membrane. The outer membrane lipids contain long sugar polymer extensions called lipopolysaccharides, or LPS (1, figure 3). LPS is a major antigen recognized by the sera of infected humans and animals (3). Serovar classification does not always match the genetic classification of leptospira species. For instance, DNA-similar strains can show up in different serogroups, and two serovars in the same serogroup can have drastically different DNA marker variations (6). Serological classification has shown that specific serovars are host-adapted to specific reservoir species; therefore, those serovars don’t cause disease in those hosts (7). For instance, L. interrogans serovar Canicola is host-adapted to dogs, L. interrogans serovar Bratslavia is host-adapted to horses, L. interrogans serovar Pomona is host-adapted to pigs, and L. interrogans serovar Icterohaemorrhagiae is host-adapted to rats. Furthermore, serological classification is helpful epidemiologically as the production of leptospira vaccines is directly related to antigen diversity.
Prevalence of Leptospirosis and Preventative Measures
Leptospirosis is one of the major zoonotic diseases worldwide and it has a large impact on both human and veterinary public health (8). Leptospirosis is prevalent in both developed and developing countries; however, the disease is primarily associated with poor communities that lack proper sanitation facilities and flood-prone regions (4). Pro-active prevention methods for reducing the spread of leptospirosis can be difficult due to the sheer diversity of leptospirosis-inducing bacterial strains.
Prevalence
Pathogenic leptospires live in the proximal renal tubules of the kidneys of carriers and are spread through the excretion of urine. Infections of animals or humans occur from direct contact with urine or indirectly from contaminated water (3, figure 4). There is a worldwide occupational association with human contraction of leptospirosis as direct occupational exposure occurs with many farmers, veterinarians, meat inspectors, and rodent control workers. (3,9). Indirect occupational exposure occurs with many sewer workers, miners, soldiers, septic tank cleaners, fish farmers, and game keepers (9).

Tropical, developing countries are constantly battling outbreaks of leptospirosis. In developing countries, many cases of leptospirosis in humans come about from activities of daily life, such as walking barefoot in damp conditions or by drinking contaminated water (9). Furthermore, there are many stray dogs in most developing countries and they have been classified as a significant reservoir for human infection (10). Another obstacle developing countries face is the spread of contaminated water during flooding. Many studies have documented instances where numbers of leptospirosis infected individuals increased soon after flooding (9). This phenomenon is likely due to the increased exposure to contaminated water in flooded regions.
Faine (1994) developed three epidemiological patterns of leptospirosis. The first pattern occurs in temperate climates where few serovars are involved and human infection almost always occurs through direct contact with infected animals through farming of cattle and pigs (11). The second pattern occurs in wet tropical areas that have many more serovars infecting humans and animals, along with larger numbers of reservoir species (11). The last pattern occurs in urban environments and involves rodent-borne infection (11). Because of heightened rodent control measures, this last pattern is rarely seen in developed countries; however, the pattern is still seen in outbreaks occurring in slum areas in developing countries (11).
Prevention
Occupational and recreational precautions, such as wearing protective clothing and avoiding splashes from urine or water, can help curb leptospirosis; however, complete prevention of leptospirosis without vaccination is quite hard (3). Immunity from leptospirosis is mostly humorally related in humans and animals and the first line of defense against leptospires is the innate immune system, which aims to remove foreign leptospires without the use of antigen presentation (4). In fact, non-pathogenic leptospires can be killed and eliminated from the body within minutes due to the immediate response of the innate immune system (4). The innate immune system is not enough to combat pathogenic leptospires, however, as they are able to survive due to their resistance to the action of the innate immune system. Since leptospires are extracellular pathogens, the acquired immune response depends on the production of antibodies (4). The adaptive immune system of infected humans or animals undergoes a process known as antigen presentation, where antibodies are produced that act against the antigens present on the LPS of the infecting leptospires. Antigen presentation is the basis for vaccinations as passive immunization with polyclonal or monoclonal anti-LPS antibodies is able to give protection against the leptospirosis disease (4).
Leptospirosis vaccination is a high priority in the veterinary field and vaccines have been used since the 1920’s (3). Current available vaccines are inactivated leptospires or outer membrane fractions (4). Furthermore, the vaccines do not produce a T cell dependent response, which makes annual booster shots a necessity. Another complication with leptospira vaccines is that there is no cross-protection against leptospiral serovars not included in the vaccine preparation. This means that when new serovars emerge, an entirely new vaccine must be produced. Because different geographical populations have such varied pathogenic serovars, the continuation of epidemiological studies is of the utmost importance for a successful vaccination program. Canine vaccines generally contain the serovars Canicola, Icterohaemorrhagiae, Grippotyphosa, and Pomona (figure 5), while pig leptospiral vaccinations generally contain the serovars Pomona, Grippoyphosa, Bratislava, Canicola, and Icterohaemorrhagiae (3).
Various research has been conducted in an attempt to find a vaccine that can generate cross-protective immunity against a number of serovars. The leptospiral outer membrane is dominated by the lipoprotein LipL32 (3). This outer membrane protein has been a prime candidate for use in a vaccine that would provide cross-protective immunity because it is a highly conserved outer membrane protein in leptospires (9). Various studies on the efficacy of immunization with LipL32 have shown promising results; however, the efficacy levels are not high enough for LipL32 immunization to be commercially produced. Adler and Moctezuma (2010) came up with a possible hypothesis for the variable results seen with LipL32 immunization. They surmise that LipL32 antibodies may be directed against epitomes that are not accessible on the leptospiral surface. This hypothesis is currently speculative at best because little is known about the structure or membrane topology of LipL32 (3).
Another candidate for a recombinant protein vaccine involves leptospiral immunoglobulin-like (Lig) proteins that belong to a family of surface-exposed determinants (4). Lig proteins are expressed during host infection and can bind to a variety of extracellular matrix components, therefore creating adhesion between leptospiral cells and host cells (4). Recent experimental studies on the effectiveness of a Lig protein vaccine have shown promise. Immunization in various studies did not grant sterilizing immunity; however, the C-terminal portion of LigA is still the most promising candidate to date for a cross-protection vaccine (4).
Case Study on Immunogenic and Antigenic Potential of Lipoproteins from Leptospira interrogans
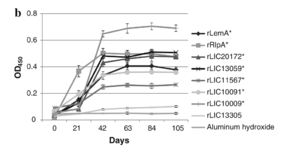
In a recent study, Hartwig et al. (2011) identified potentially protective lipoproteins from Leptospira interrogans serovar Copenhageni and evaulated the immunological properties of these lipoproteins. Their rational for this study was that current available leptospirosis vaccines require annual booster shots and confer only servovar specific immunity (Hartwig et al., 2011). The following eight lipoproteins were selected and isolated from a patient with severe leptospirosis: LemA, RlpA, LIC10009, LIC10091, LIC11567, LIC13305, LIC13059, and LIC20172. The eight chosen lipoproteins were cloned and expressed in Esherichia coli , and then inoculated in mice to test their humoral immune response. Importantly, immunized mice produced a significant anti-leptospiral IgG response to all of the recombinant proteins except LIC13305 (15, Figure 6). IgG antibodies are the most common antibodies and are extremely important in fighting bacterial and viral infections (16). Hartwig et al. (2011) found that on average, the IgG response peaked on day 42 post-immunization, and was maintained throughout the study period of 105 days. Furthermore, all of the recombinant proteins were recognized by antibodies present in the sera from patients with severe leptospirosis (15). The importance of this study lies in the fact that Hartwig et al. (2011) found 7 potential lipoproteins (LIC13305 failed to produce an IgG response) that are potential vaccine candidates that would induce cross-serovar immunity.
Pathology of Leptospirosis and Diagnostic Methods
Pathology
While much remains to be discovered about the mechanisms by which leptospires cause disease, some key parts are understood. Pathogenic leptospires enter the body through small abrasions or via mucous membranes (3). The bacteria then circulate in the blood stream for a period up to 7 days after which the number of leptospires in the blood and tissues will eventually reach a critical level, which is concurrent with renal tubular necrosis (3). The clinical presentation of leptospirosis is biphasic. The septicemic phase lasts about a week and is followed by the immune phase, which is characterized by antibody production and excretion of leptospires in the urine (9). Infections from leptospires are broken down into two main types. The first is anicteric leptospirosis, meaning leptospirosis not accompanied by jaundice. This type of infection is considered mild and may be accompanied by the following symptoms: chills, headache, fever, abdominal pain, and a skin rash (9). The second type of infection is icteric leptospirosis, or leptospirosis accompanied by jaundice. Icteric leptospirosis is considered a much more severe disease. The clinical course for this type of leptospirosis is usually rapidly progressive with multisystemic complications (9). Histopathology is most noticeable in the liver, kidneys, heart, and lungs. Icteric leptospirosis is a very common cause of acute renal failure.
Diagnosis of Leptospirosis
Because this disease is often mutlisystemic with a wide variety of clinical signs, diagnosis can be difficult. A variety of specific microbiological tests are necessary to confirm that a patient is infected with leptospirosis. Much like the two classification systems, isolated leptospires are identified either by serological methods or molecular techniques (9). Samples of blood, urine, or cerebral spinal fluid from infected patients can be visualized by dark-field microscopy to check for leptospires. This technique does work, however, it is insensitive and lacks specificity (9).
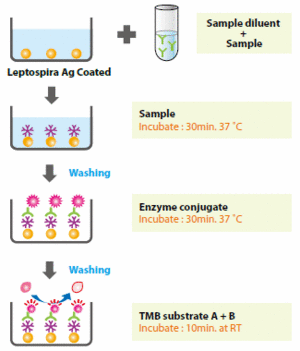
Another method often used during differential diagnostics is an enzyme-linked immunoabsorbent assay (ELISA). ELISA is used to detect the presence of leptospiral antigens (figure 6). To conduct an ELISA, an unknown amount of antigen is fixed to the bottom of a well plate and a specific antibody is applied over the surface so that it can bind to the antigen (12). The antibody is linked to an enzyme, which can produce a visible change (e.g. color change) that can be quantified in terms of optical density (12).
Another method used for serological diagnosis of leptospirosis is a microscopic agglutination test (MAT). MAT is considered the definitive serological test for leptospirosis (9). Patient sera is reacted with live antigen suspensions of leptospiral serovars and incubated. After incubation, the serum-antigen mixture is examined microscopically for leptospire agglutination and the titers are quantified (9). Usually the range of antigens used includes all the locally common serovars (9). MAT is analyzed by dark-field microscopy and the end point is the highest dilution of serum at which 50% agglutination occurs (9). The current CDC case definition for leptospirosis states that a titer > 200 with a clinically compatible illness is indicative for leptospirosis (9). It should be noted, however, that in regions where leptospirosis is endemic (e.g. tropical countries), a single titer > 800 in symptomatic patients is indicative of leptospirosis (9). One advantage of the MAT procedure is that it is specific for serovars. Despite their high sensitivity and specificity, MAT procedures are currently complex and costly, which reduces its worldwide distribution and use (13).
Because serological tests are often time consuming and expensive, alternative molecular diagnosis methods have been developed for characterizing leptospiral isolates. Modern molecular diagnosis methods include restriction fragment length polymorphism (RFLP) analysis, PCR based approaches, such as16S rRNA sequencing, and multiple locus sequence typing (MLST) (3). Depending on which primers are used, PCR analysis will yield only species level analysis or both species analysis and serovar level analysis. Most PCR-based diagnoses of leptospirosis fail to identify the infecting serovar. This usually isn’t problematic for individual patient management; however, identifying serovars are important for epidemiological and public health value (9). A study by Bezerra da Silva et al. (2011) designed primers that were complementary to polymorphic regions in the rfb, the nucleotide sequence of the LPS biosynthetic operon. The rfb varies among many Leptospira serovars within the same species because the LPS is highly immunogenic and is responsible for serovar specificity (2,9). While their method was not entirely optimal, Bezerra da Silva et al. (2011) designed a promising method that was able to distinguish serovars.
Yet another new, promising method for leptospiral isolate characterization is the Check-Points method. This method came about because of the associated drawbacks with current leptospiral diagnosis methods. Serological and molecular diagnostic methods often have low reproducibility, with a lack of directly generating electronic portable data (13). Furthermore, most current methods have a need for viable leptospiral cultures and are labor intensive, costly, and time consuming (13). The Check-Point system is essentially a multiplex ligase-depended probe amplification in which the distinct products are detected by hybridization to specific probes that are immobilized (14). More specifically, DNA was extracted from leptospira strains and run against specific probes designed on the basis of single nucleotide polymorphisms selected to be specific for distinct species. While the use of the Check-Points method for leptospira characterization is up and coming, the method does not require expensive or complex equipment and would allow leptospira characterization to be conducted in many more facilities than is currently possible (13).
Conclusion
Infection caused by pathogenic leptospires is an important zoonotic disease worldwide that infects a multitude of mammals, ranging from rats to humans. The complexity of leptospirosis is due to there being over 200 serovars, with multiple serovars in individual leptospira species. Leptospirosis is a worldwide zoonotic disease that can infect nearly every mammal on earth. The strains of pathogenic leptospires humans are most at risk for contracting are dependent on geographic location and which reservoir species humans are most likely to be exposed to (e.g. rats are typically reservoir species for L. interrogans serovar Icterohaemorrhagiae). For this reason there are serological, as well as molecular diagnostic tests that are often used to determine which particular strain caused the disease. Serological tests are extremely important in the field of epidemiology because current vaccinations rely on knowing which serovars a population is most likely to be exposed to. Current studies have goals to determine alternative vaccines that can produce cross-protective immunity against a number of serovars to further heighten protection against leptospirosis. Unfortunately, the symptoms of leptospirosis are often multisystemic and can be difficult to differentiate between other illnesses, such as the influenza virus. Many methods used to successfully diagnose leptospirosis involve very costly equipment, are very time consuming, and require a relatively high microbiological skill set. This can be problematic because high incidences of leptospirosis often occur in developing countries that experience more contact with contaminated drinking water and infected animals. Certain alternative methods for diagnosing leptospirosis, such as the Check-Point method, are being developed to bring simpler and less expensive methods to scientists and doctors working in developing countries.
References
(1) Slonczewski, J.L. and J.W. Foster. 2011. Microbiology: An Evolving Science . W.W. Norton & Company, Inc., New York, 1097 pp.
(2) Bezerra da Silva, J., E. Carvalho, R.A. Hartskeeri, and P.L. Ho. 2011. Evaluation of the use of selective PCR amplification of LPS biosynthesis genes for molecular typing of Leptospira at the serovar level. Current Microbiology . 62:518-524.
(3) Adler, B. and A.P. Moctezuma. 2010. Leptospira and leptospirosis. Veterinary Microbiology . 140:287-296.
(4) Fraga, T.R., A.S. Barbosa, and L. Isaac. 2011. Leptospirosis: aspects of innate immunity, immunopathogenesis, and immune evasion from the complement system. Scandinavian Journal of Immunology . 73:408-419.
(5) Barmettler, R., A. Schweighauser, S. Bigler, A.M. Grooters, and T. Francey. 2011. Assessment of exposure to Leptospira serovars in veterinary staff and dog owners in contact with infected dogs. JAVMA . 238:183-188.
(6) "Bacteriological Information." 2009. http://www.leptospirosis.org/.
(7) "Leptospirosis in Humans." 2000. http://www.vetmed.wisc.edu/pbs/zoonoses/leptospira/leptohuman.html.
(8) Collares-Pereira, M., H. Korver, B.V. Cao Thi, M. Santos-Reis, E. Bellenger, G. Baranton, and W.J. Terpstra. 2000. Analysis of Leptospira isolates from mainland Portugal and the Azores Islands. FEMS Microbiology Letters . 185:181-187.
(9). Levett, P.N. 2001. Leptospirosis. Clinical Microbiology Reviews . 14:296-326.
(10). Weekes, C.C., C.O. Everard, and P.N. Levett. 1997. Seroepidemiology of canine leptospirosis on the island of Barbados. Veterinary Microbiology . 51:215-222.
(11). Faine, S. 1994. Leptospira and leptospirosis. . CRC Press, Boca Raton, Florida.
(12) "ELISA Method." 2002. http://www.bio.davidson.edu/courses/genomics/method/ELISA.html.
(13). Ahmed, A., R.M. Anthony, and R.A.Hartskeerl. 2010. A simple and rapid molecular method for Leptospira species identification. Infection, Genetics,and Evolution . 10:955-962.
(14). Bergval, I.L., R.N.Vijzelaar, E.R. Dalla Costa, A.R. Schuitema, L. Oskam, A.L. Kritski, P.R. Klatser, and R.M. Anthony. 2008. Development of multiplex assay for rapid characterization of Mycobacterium tuberculosis . Journal of Clinical Microbiology . 46:689-699.
(15). Hartwig, D.D., F.K Seixas, G.M. Cerqueira, A.J. McBride, and O.A. Dellagostin. 2011. Characterization of the immunogenic and antigenic potential of putative lipoproteins from Leptospira interrogans . Current Microbiology . 62:1337-1341.
(16). WebMD. 2011. "Immunoglobulins." http://www.webmd.com/a-to-z-guides/immunoglobulins.
Edited by Ian Veitch, student of Joan Slonczewski for BIOL 238 Microbiology, 2011, Kenyon College.
