Leuconostoc
A Microbial Biorealm page on the genus Leuconostoc
Classification
Higher order taxa:
Bacteria; Firmicutes; Bacilli; Lactobacillales; Leuconostocaceae
Species:
L. carnosum, L. citreum, L. durionis, L. fallax, L. ficulneum, L. fructosum, L. garlicum, L. gasicomitatum,
L. gelidum, L. inhae, L. kimchii, L. lactis, L. mesenteroides, L. pseudoficulneum, L. pseudomesenteroides
|
NCBI: Taxonomy |
Description and Significance
Leuconostocs are traditionally found in association with plant matter, fermenting vegetables, milk, dairy products, and wines and meats. Leuconostocs were first isolated in 1878 by Cienkowski. Leuconostoc usually nonpathogenic acid-tolerant organisms with optimal temperature 18 and 25°C, but the group is quite diverse. For example, L. carnosum is an anaerobic bacterium found in spoiled, packaged meat. Optimal temperature for the organism is 2°C. L. carnosum able to inhibit growth of other (even closely related) bacteria. So, it may be used as biopreservative. Leuconostoc in general is important to fermentation of vegetables.
Genome Structure
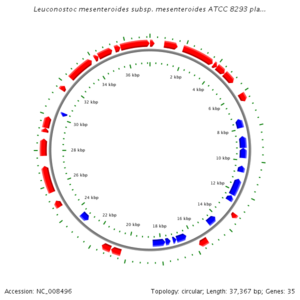
The genome of Leuconostoc is comprised of 2,038,396 bp with 2,075,763 nucleotides arranged in a circular design, and encodes for 2005 different proteins, with 54% of them being assigned putative biological functions. The G-C content of 37-40% has been reported as standard. As Luconostoc spp. is a lactic acid bacteria, the genome encoded all enzymes present in the 6-phosphogluconate phosphoketolase pathway. Genes were also found that encoded pyruvate dissipating enzymes that are predicted to catalyze the production of many metabolites leading to various end products of fermentation. Additionally, certain species of Leuconostoc were found to contain plasmids, randing from 1-7, ranging from 1.3 to 80Mda with genes that encode for bacteriocin production and citrate metabolism. (F. Breidt Jr. and Plengvidhya, Vethachai)
Cell Structure and Metabolism
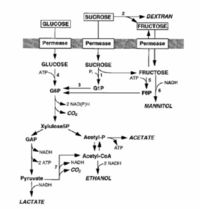
Leuconostoc is a gram-positive coccus, often lenticular on agar and usually occurs in pairs or chains. Leuconostoc is nonmotile, not spore forming chemoorganotrophic facultative anaerobe. Leuconostoc requires rich, complex media. It is catalase-negative nonproteolytic organism without cytochromes. Leuconostoc is nonhemolytic, vancomycin resistant organism. Leuconostoc requires nicotinic acid, thiamin, biotin, and pantothenic acid or one of its derivatives. It is heterofermentative (uses a combination of the pentose phosphate and phosphoketolase pathways). (Thunnel 1995) L. mesenteroides able to convert sugar to mannitol and dextran.
Ecology
Caulobacter generally live in a dilute aquatic environment where the most common limiting nutrient is phosphorus, an essential element for healthy growth. Lack of this nutrient induce Caulobacter to dramatically elongate its stalk up to 30 times longer than those in phosphate-rich medium (Brun et al. 2000).
Leuconostoc can often be found in the wild and is a part of the natural microflora in almost all farming fields. It is most commonly found in many different processed foods, either as a starter culture or as a contaminant. For many foods, such as fruit juice or milk products, Leuconostoc can contribute to off flavors due to diacetyl production. in certain alcohols, meats, vegetable, and fermented milk products like cheese or yogurt, though, it is often used as a starter culture for the same diacetyl production capacity.
Cell Division
Caulobacter asymmetrically divides to produce a motile swarmer cell and a stalk cell. The swarmer cell, which has a flagellum, swims for about 30-45 minutes before shedding the flagellum and differentiating into a stalk cell. The flagellum is ejected from the swamer by the destruction of the structures (MS ring) at the base of the flagellum. Within the swarmer cell, chromosomes do not replicate; however, chromosome replication begins immediately in the daughter cell with the stalk and when the swarmer loses its tail (Stanford). The stalk adheres to surfaces through an adhesive organelle called the holdfast.
Several two-component signal transduction proteins are involved in the cell cycle progression by accumulating at one or both poles "in a spatial and temporal pattern that is reproduced during each cycle" (Jacobs-Wagner 2003). In addition to this, DNA methylation is main component of signaling differentiation. Throughout the cell cycle, the chromosome progressively goes from being fully methylated to hemimethylated during DNA replication - this results in differential binding of regulatory proteins to activate or repress transcription. This was studied by using the CtrA gene, which encodes for an important cell cycle regulatory protein. The CtrA gene has two promoters, one of which "fires early" in the S phase and is recognized by the CcrM DNA methyltransferase. Analysis showed that this P1 promoter is actually repressed by DNA methylation (Reinsenauer and Shapiro 2002).