Listeria Monocytogenes: A Pathological Perspective: Difference between revisions
No edit summary |
|||
| Line 1: | Line 1: | ||
{{Curated}} | |||
Edited By: Dominic Camperchioli | Edited By: Dominic Camperchioli | ||
<br> | <br> | ||
Latest revision as of 14:18, 1 October 2015
Edited By: Dominic Camperchioli
Physiology
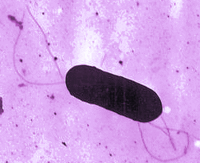
Listeria monocytogenes is an aerobic Gram-positive motile bacillus commonly found in pregnant women, newborns, and people with compromised immune systems (Figure 1). As L. monocytogenes bacteria commonly grow in the soil, they are often consumed by animals and from there become a food borne illness for humans. Usually listeriosis results from dairy consumption (1), but it can be transmitted through other sorts of foods, such as the Listeria outbreak that originated at a Colorado cantaloupe farm (2).
However, listeriosis is not as common as one might imagine, as most healthy individuals are not normally affected by small amounts of L. monocytogenes (1). Most infections begin in the gastrointestinal tract, specifically the small intestine, where macrophages are common. From there, the infection will generally spread to surrounding tissues, such as epithelial cells and hepatocytes, before migrating up to the brain, causing meningitis (3). Although rare, listeriosis is lethal in about 30% of cases (4).
L. monocytogenes bacteria prosper intracellularly, preferring conditions inside of macrophages, which are optimal for growth and replication; the bacteria survive much worse extracellularly because means for growth and replication are not as easily accessible. Although antibodies and vaccinations are not effective against L. monocytogenes, acquired immunity is possible but not common (5).
This page starts with a general overview of the life cycle of L. monocytogenes and its virulence factors and then analyzes effects of antibiotics and host cell defense methods, summarizing the status quo of Listeria monocytogenes from a pathological perspective.
Overview of L. monocytogenes
Scientists have been aware of Listeria monocytogenes since the 1920’s (4), yet much about it is still unknown. However, its life cycle has been studied carefully, as it is different from most pathogens. These differences lead to variations among effective treatment methods when infected with L. monocytogenes and other common gram-positive pathogens; its specific pathogenesis is unlike most known. L. monocytogenes continues to be a hot topic for research because of its infamous lethality through infection (3). To effectively analyze the mechanisms used by hosts to ward off infection, one must first understand the life cycle and virulence factors of L. monocytogenes.
Life Cycle
This enitre section is summarized by the diagram in Figure 3.
Entry into Host Cells
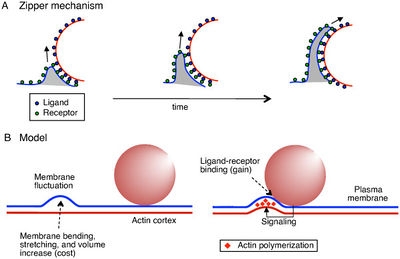
Infection usually begins in the small intestine, where macrophages easily engulf L. monocytogenes. However, L. monocytogenes can induce their own phagocytosis into nonphagocytic cells due to the internalin operon, which encodes InlA and InlB. InlA and InlB produce internalin A and B, which bind to receptors on the host cell wall, therefore inducing the phagocytosis of L. monocytogenes. Either one is sufficient on its own to induce phagocytosis as long as the needed receptors are present on the host cell (E-cadherin for InlA and the hepatocyte growth factor receptor, HGF-Rc/c-Met, for InlB). Host cells tricked by these proteins use the zipper mechanism of phagocytosis, in which receptors on the host bind with ligands on L. monocytogenes sequentially, eventually "zipping" the two membranes together and engulfing L. monocytogenes (Figure 2). In addition to InlA and InlB, Listeriolysin O (LLO), a pore-forming toxin similar to many other cholesterol-dependent cytolysins commonly found in gram-positive bacteria, also induces phagocytosis. LLO forms enough pores in the membrane to induce membrane internalization because of the host's ability to repair its plasma membrane, bringing L. monocytogenes into the host (6).
Lysis of Host Vacuole
Once introduced into the host cell, L. monocytogenes must escape from the vacuole in which it was internalized before entering the cytoplasm to grow and replicate. Again LLO is the protein responsible for pore formation and this time responsible for the escape of L. monocytogenes from the vacuole (7).
LLO also activates Ca2+ intracellular channels and channel independent Ca2+ mechanisms. These mechanisms result in host cell responses with diverse effects on both the host cell and L. monocytogenes, including host cell degranulation, pro-inflammatory mediators, apoptosis, and cytoskeletal rearrangement. L. monocytogenes benefit from Ca2+ influx into the host cell through the production of lipid rafts in the host cell’s membrane, to which more LLO can later bind, and through cytokine synthesis. L. monocytogenes later uses the cytokine it synthesizes to activate IP3R Ca2+ channels through tyrosine phosphorylation and G-protein activation of PLCs (8). IP3R channels release Ca2+ from the endoplasmic reticulum (9), additionally contributing to the excess of Ca2+ in the cytoplasm (8).
In addition to LLO, L. monocytogenes escapes from the vacuole by utilizing two phospholipase Cs (PI-PLC and PC-PLC, which are capable of lysing the vacuole without LLO). These PLCs lyse the individual phospholipids of the membrane into separate phosphates and ester tails, which are then dissolved in the cytoplasmic water of the host cell (12).
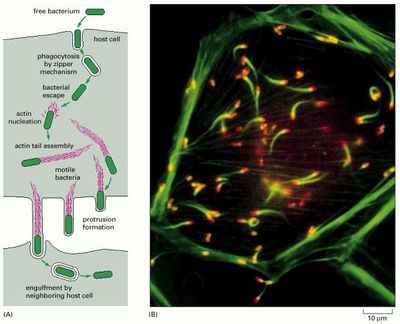
Actin Nucleation and Motility
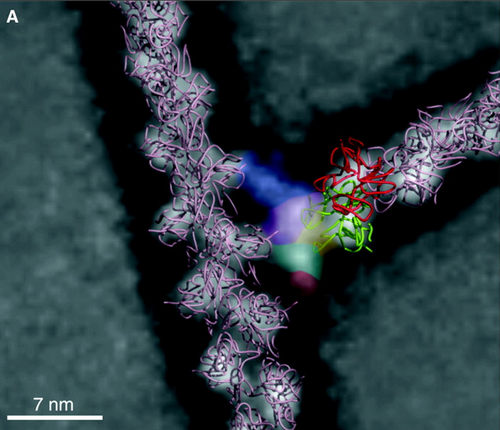
In order for L. monocytogenes to enter the next host cell, they must find a method for motility. ActA, the second gene of the lecithinase operon, carries the responsibility for both actin nucleation and motility (11). It produces ActA protein, a surface protein of L. monocytogenes that induces the production of actin within the host cell and focuses the production at one pole of the bacteria. Additionally, actA creates binding sites on the bacteria for host cell actin to attach, binding to the NH2 group of ActA protein (7). Once actin nucleates at a pole of L. monocytogenes, the Arp2/3 complex activates and assists in binding the actin filaments together at the tail, making the rotation of the actin capable of propelling the bacteria (Figure 4) (13). However, the nature of actin encourages constant binding and formation at their barbed ends. Therefore, a regulating protein near the Arp 2/3 complex capable of slowing growth near the junction while not preventing actin formation where needed is necessary. The enabled/vasodilator-stimulated phosphoproteins (Ena/VASP) bind to actin and delay F-actin capping, promoting the formation of linear actin filaments instead of highly branched ones (2, 14). Through this complex mechanism, L. monocytogenes moves not only through the host cell but also into adjacent host cells to repeat the cycle of growth and replication (Figure 3B) (11).
However, L. monocytogenes does not just stop once it reaches its destination; it stops when there is not longer production of the actin complex, which halts movement (11). This is because of the relative rigidity of actin, which allows for elastic Brownian ratchet movement to act as the driving force of the cell as more actin is produced (15).
Intercellular Spread
After L. monocytogenes enters the new host cell, two layers of membranes engulf it, one from the cell from which it came and one from the new cell it enters. Just as in the original host cell, L. monocytogenes must lyse both of these membranes in order to enter the host cell’s cytosol (16). To accomplish this, L. monocytogenes not only uses Listeriolysin O, PI-PLC, and PC-PLC, but also lecithinase, which is only produced after actA produces ActA protein in the lecithinase operon, suggesting there may be a specific need for lecithinase in helping with the destruction of the two host membranes but not in the destruction of just one (16). Other than the addition of lecithinase, the process is the same as the lysing of the singular cell membrane. From here, the process repeats itself as L. monocytogenes spreads throughout the host.
Virulence Factors
Of the many genes that contribute to the virulence of L. monocytogenes, the prfA sequence regulates five of them in the plcA-prfA operon. Although the virulence proteins regulated by the prfA sequence do not always have the same level of expression, they all have higher expression levels when in macrophages compared to epithelial or endothelial cells. Virulence expression also appears somewhat dependent upon environmental factors not necessarily specific to the cell. PrfA protein contains a leucine zipper structure that binds to DNA in order to activate the translation of the sequences responsible for coding the virulence factors (17). Although it is unkown what the exact method of prfA activation is (18), it appears positively self-regulated (19). It is clear that a lack of prfA expression lowers the cell’s virulence to almost none and constitutive expression of prfA is not beneficial to L. monocytogenes; proper regulation is needed (18). The genes regulated by prfA are:
• plcA(17), responsible for PI-PLC production. PI-PLC hydrolyzes PI and PI-glycan (11). Additionally, PI-PLC helps in the escape of L. monocytogenes from the host phagosomes (12).
• hly(17), responsible for the formation of listeriolysin O, the main factor in classification of L. monocytogenes virulence and a pore-forming toxin that assists in the induction of L. monocytogenes into the host cell and the release of it from the phagosomes (11).
• mpl(17), codes for a metalloprotease. Without this gene, about one half of the necessary genes are not coded. L. monocytogenes can survive without LLO if Mpl and PC-PLC are coded for (20).
• actA(17), responsible for everything involving actin (figure 4), including the formation of ActA, the protein responsible for nucleating actin formation at one pole of the bacterial cell and allowing actin to attach to the cell (11).
• plcB(17), responsible for PC-PLC production, which, similarly to PI-PLC, can help the escape of L. monocytogenes from the phagosome. It is commonly activated during intracellular spread (3).
Other virulence inducing genes that are not regulated by prfA include:
• inlA and inlB, which code for internalin A and internalin B. Interestingly, prfA seems to partially regulate these genes. The proteins help induce the phagocytosis of L. monocytogenes by non-phagocytotic cells, enabling the spread of the bacteria throughout more of the host’s cells (17).
• p60 gene, which codes for p60 protein. Although the exact function of p60 protein is unknown, it is essential for L. monocytogenes virulence and is believed to be responsible for shortening the length of bacterial chains and encouraging the invasion of host cells (21).
• fbpA, a more recently discovered gene, encodes for fbpA protein, which helps L. monocytogenes adhere to host cells, promoting induction. Additionally, it appears to increase the amount of LLO and internalin B, which increase the virulence of the bacteria (22).
The Use of Antibiotics Against Infection
Antibiotics commonly effective against Gram-positive bacteria has also been effective against L. monocytogenes. However, L. monocytogenes is starting to go the way of strains of Staphylococcus and Streptococcus; slowly but surely resistance to antibiotics is evolving. A single strain of L. monocytogenes was found resistant to chloramphenicol, erythromycin, and streptomycin (23). Additionally, species of Listeria closely related to L. monocytogenes but not pathogenic are resistant to many other common antibiotics, such as tetracycline, penicillin G, ampicillin, vancomycin, and gentamycin (4). Most of these resistance factors are found on plasmids capable of interspecies horizontal transfer, so it will not be long before resistance to these antibiotics is no stranger than Streptococcus resistance to them. In fact, many of the resistance genes found in L. monocytogenes were transferred from Streptococcus or Enterococcus (23).
Specific antibiotic resistance to streptomycin, erythromycin, tetracycline, and chloramphenicol in L. monocytogenes resides on plasmid pIP811, which easily transfers among common Gram-positive bacteria. The genes in this plasmid share many similarities to those with the same resistance capacities found in Enterococci or Streptococci; one gene (responsible for chloramphenicol resistance) is even homologous to its counterpart in Enterococci and Streptococci (23).
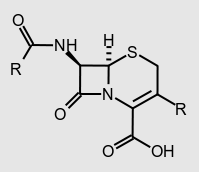
L. monocytogenes expresses uniquely high resistance to cephalosporins, which are typically used to treat Gram-positive infections of unknown origin due to their strength (Figure 5). The gene liaS causes resistance to cephalosporins, which simultaneously increases the susceptibility of the cell to nisin. Conversely, mutant lmo2229 increases cell resistance to nisin while decreasing resistance to cephalosporins. In order to make L. monocytogenes as susceptible to nisin as possible, research is constantly being conducted on the virulence factors of L. monocytogenes, looking for weaknesses to exploit. Recently, extreme susceptibility to both nisin and antibacterial medications has been discovered in the TelA gene. The eventual goal is to implicate the susceptible sequences into L. monocytogenes, lowering the chances of it to cause foodborne illness (24).
CesRK, another important gene discovered in L. monocytogenes that provides the bacteria with significant antibiotic resistance, forms many types of membrane proteins and has a two-part system. CesRK resistance extends beyond ß-lactam antibiotics to increased resistance to ethanol. Possibly, CesRK may eventually serve as a regulator for alcohol-induced genes within L. monocytogenes, becoming a problem regarding the effectiveness of many alcohol disinfectants. Unfortunately, alcohol based disinfectants will become ineffective against L. monocytogenes, one of the more dangerous species of bacteria (25).
Similar to the two-part functioning of CesRK, LisRK also divides into a two-part system. This time, however, the gene is in charge of osmoregulation of the cell, adapting easily to high levels of solutes in both its host cell and the outside environment. Sometimes another gene, HtrA, assists LisRK greatly when trying to adjust osmolarity while forming filaments (26). Additionally, LisRK can adapt to pH stresses and hydrogen peroxide stresses elegantly without facing too much of a burden from the adjustment (27). Obviously, technicalities are controlled by many genes and proteins that we have not yet discovered or attributed to a function and there is much more to be investigated that may hold a key to a new antibiotic.
Novel Host Cell Defense Mechanism
Because of its life cycle and habitat, L. monocytogenes is a tricky bacterium to work with. Its easy spread from one host to another makes it rather difficult to combat, especially because of its location within macrophages of the host (11). For now, a combination of antibiotics works fairly well, but, similarly in fashion to S. aureus, many strains are developing antibiotic resistances (4). Therefore, investigating defense mechanisms within hosts helps analyze where weaknesses within the immune system responses may be (28). Below are some current investigations in which host cell defense mechanisms for L. monocytogenes have been analyzed.
Antimicrobial Peptides (AMPs)
Antimicrobial peptides are diverse molecules responsible for immunity to a plethora of fungi, viruses, and bacteria. By attacking all sorts of cellular processes, they prevent infection of the host organism, and specific α-defensins have even been observed inhibiting bacterial toxins. In the case of L. monocytogenes, a human AMP, HNP-1, traps the bacteria within the host cell’s vacuole, preventing the proliferation of the species in the host. The AMPs inside human macrophages not only trap the foreign species but also induce higher levels of phagocytosis. The presence of L. monocytogenes induces α-defensin HNP-1 production within the host cell as an immune response to the bacteria’s presence. HNP-1 does not assist with the phagocytosis of L. monocytogenes, but it does help trap them within the vacuoles of the host cell (28). This prevents replication, growth, and spread, as L. monocytogenes needs to be within the host cell’s cytosol to begin these processes (3). The HNP-1 does so by colocalizing with the bacteria inside the host cell. Once this occurs, HNP-1 prevents the release of listeriolysin O from phagosomes and also halts its ability to form channels through the membranes. Hence, HNP-1 traps L. monocytogenes inside the phagosome until the host cell properly deals with the invader. AMPs may be the host cell’s single most effective method of repressing the infection (28).
Autophagy Mediating Pathways
Although macrophages constantly engulf L. monocytogenes, they do not usually destroy the bacteria. Macrophages eliminate L. monocytogenes through either destruction of the entire host cell (apoptosis) or the lysing of a certain part (autophagy). Autophagy is insufficient when either the TLR2 or the NOD pathway is not present in the cell because the ERK pathway cannot signal its induction (29).
Riboflavin Deficiency
Riboflavin (Vitamin B2) deficiency is common in the same demographics as listeriosis. Riboflavin kinase lyses riboflavin into flavin mononucleotide and flavin adenine dinucleotide, which are essential for proper NADPH oxidase 2 (Nox2) priming. Nox2 contributes significantly to the host cell’s immune response to bacterial infections, therefore making a deficiency of riboflavin detrimental to a host’s ability to fight infections. Specifically, Nox2 depletion causes lowered levels of neutrophils, macrophages, and phagocytes in the host, allowing easier L. monocytogenes survival (30).
MPYS Pathway Resistance
When L. monocytogenes enters a host cell, the host cell’s MPYS stimulator is triggered. This stimulation causes Ly6Chi monocytes to be summoned to the infected region using the CCR2 regulatory pathway. In this pathway, CCR2 (a cytokine receptor) receives MPYS and the signal is sent to the monocytes via MCP-1 or MCP-3 (both ligands of CCR2), causing their release from bone marrow. However, this pathway only induces the release of Ly6Chi; the locating of the site of infection is done through interaction of surface proteins of the monocyte and the host cell, CD11b and ICAM-1 respectively. Once in the infected area, Ly6Chi eliminates the bacteria via phagocytosis (5).
T Helper 17 Cytokines vs. Epstein Barr Virus-Induced 3
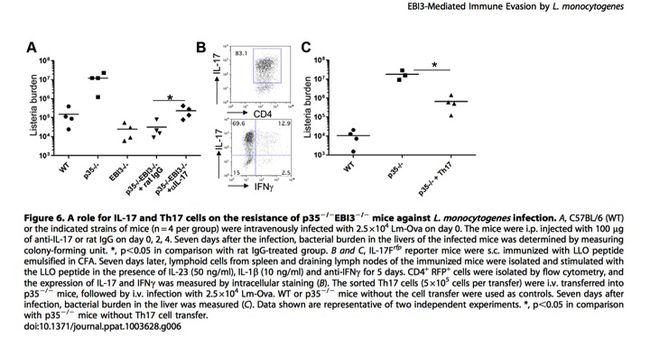
T helper 17 cells (Th17) are common in the human body and are often responsible for the destruction of foreign pathogenic microbes, yet they failed to do so when placed in cell culture with L. monocytogenes unless certain mutations were present in the host cell. In the instances where Th17 was successful in limiting growth of L. monocytogenes, there was a mutation in the host cell’s Epstein Barr Virus-Induced 3 (EBI3) gene (5). EBI3 protein, first found in B-lymphocytes, assists with the regulation of lymphocytic responses (31). Surprisingly, such a mutation would not benefit the immune system of the host. L. monocytogenes manipulates EBI3 into a defensive utility that cloaks its presence within the cell, therefore tricking the host into believing the pathogen is not present (Figure 6), especially in IL-12 deficient patients. IL-12 is a cytokine that either promotes or inhibits Th17 and Th1 (a cytokine similar to Th17) and therefore regulates immune responses within the host (Figure 6). When dysfunctional, L. monocytogenes easily takes advantage of EBI3 and uses it to hide its presence within the host cell. To combat this method of intracellular survival, researchers are investigating methods to alter EBI3 to make it impossible for L. monocytogenes to utilize it beneficially (5).
References
