Methanococcus: Difference between revisions
No edit summary |
No edit summary |
||
| Line 1: | Line 1: | ||
{{Curated}} | |||
{{Biorealm Genus}} | {{Biorealm Genus}} | ||
Revision as of 21:10, 6 August 2010
A Microbial Biorealm page on the genus Methanococcus
Methanogens overview
Classification
Higher order taxa:
Archaea; Euryarchaeota; Methanococci; Methanococcales; Methanocaldococcaceae
Species:
Methanococcus jannaschii, M. maripaludis, M. voltae PS
Description and Significance
M. jannaschii was the first Archaea to have it's genome sequenced, which opened the doors for comparison between the genomes of the three domains. It was originally located from a sediment sample collected from the sea floor at the base of a "white smoker" chimney on the East Pacific Rise.
Methanocaldococcus jannaschii was formally in the genus Methanococcus, but due to it's ability to grow in high temperatures it was re-classified. There were a few thermophilic species in Methanococcus that were reorganized, and this reorganization was supported by a low 16S rRNA sequence similarity between the thermophilics and mesophilics. The species left in Methanococcus are mesophilic and are related on the genus level by close DNA reassociation levels.
No difference is noted in the G + C content of thermophilics and mesophilics of Methanococcus. However, there is a difference in the proteins. The thermophilic proteins have higher residue volume, higher residue hydrophobicity, more charged amino acids, and fewer uncharged polar residues than the mesophilic proteins.
Genome Structure
M. jannaschii contains a main circular chromosome consisting of 1,664,970 bp, a large extra-chromosomal element with 58,407 bp, and a small extra-chromosomal element of 16,550 bp. There were similarities found in the genome with other domains, but there were also unique sequences.
The organism M. voltae was found to contain a system of gene transfer similar to general transduction except that the bacteriophage component (in terms of virus replication) is defective or absent. VTA (voltae transfer agent) is responsible for the transfer and it's 4.4kb fragments of DNA are resistant to DNase.


The majority of genes in M. jannaschii cannot be identified with Bacterial genomes or eukaryotic sequenced data. Of the similarities, Methanocaldococcus shared the anabolic genes with Bacteria (especially those involved in energy production and nitrogen fixation) and shared more of the cellular information processing and secretion systems with Eukaryotes.
Cell Structure and Metabolism
This autotrophic organism is strictly anaerobic and gets it's energy by the reduction of CO2 with H2 to generate methane. In addition to it's anabolic pathway, it also contains several scavenging molecules that most likely play a role in importing small organic compounds such as amino acids. Although Methanococcus spp. have a nifH-like gene they cannot fix N2, with the exception of M. maripaludis that can.
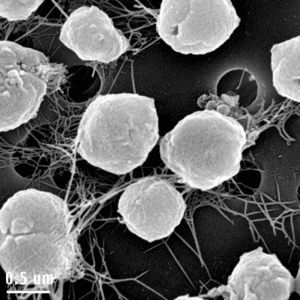
Structurally, the species consist of having two bundles of flagella at the same cellular pole along with no cell membrane, but a thin S-layer covering the plasma membrane. They are cocci in shape, about 1.0 microns in diameter, and are Gram stain negative.
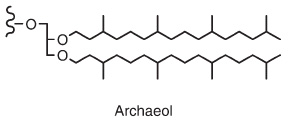
All Archaea have lipids with links between the head group and side chains, making the lipids more resistant to heat and acidity than bacterial and eukaryotic ester-linked lipids.The glycerol headgroup with two ether-linked side-chains (instead of esters) is known as archaeol.:
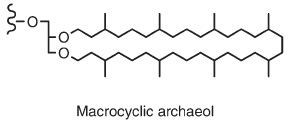
For more information on Intact Polar Lipids (IPLs) in Archaea, including tetraethers, check the description here.
Ecology
M. jannaschii can grow in habitats with pressure up to more than 200 atm and a temperature range between 48 and 94oC, with an optimum growth temperature being 85oC.
References
Bult et al. 1996. Complete Genome Sequence of the Methanogenic Archaeon, Methanococcus jannaschii. Science 273: 1058-1072.
Eiserling et al. 1999. Bacteriophage-like particles associated with the gene transfer agent of Methanococcus voltae PS. Journal of General Virology 80: 3305-3308.
Haney et al. 1999. Thermal adaptation analyzed by comparison of protein sequences from mesophilic and extremely thermophilic Methanococcus species. PNAS 96: 3578-3583.