Methanococcus jannaschii: Difference between revisions
No edit summary |
No edit summary |
||
| Line 1: | Line 1: | ||
{{Uncurated}} | |||
{{Biorealm Genus}} | {{Biorealm Genus}} | ||
Revision as of 15:40, 16 September 2010
A Microbial Biorealm page on the genus Methanococcus jannaschii
Classification
Higher order taxa
- Superkingdom: Archaea
- Phylum: Euryarchaeota
- Class: Methanococci
- Order: Methanococcales
- Family: Methanocaldococcaceae
- Genus: Methanocaldococcus (syn: Methanococcus)
Species
Genus species - Methanocaldococcus jannaschii
- synonym: Methanococcus jannaschii
Description and significance
Methanococcus jannaschii is an autotropic hyperthermophillic organism that belongs to the kingdom of Archaea. They were found to live in extreme environments such as hypothermal vents at the bottom of the oceans in which water reaches boiling temperature or pressure is extremely high (Bult, C.J. et. al., 1996). The evolutionary of these organisms and the biological mechanisms that they use to not only survive but also thrive in such extreme environments are of great interest to current researches.
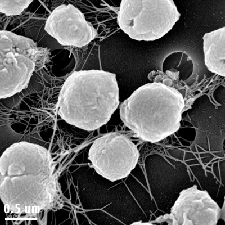
In 1982, M. jannaschii was first isolated in the East Pacific Rise, near the western coast of Mexico. The organism was found from a sample sediment taken from a 2600 m deep “white smoker” chimney. Its extreme habitat of temperature between 48oC-94oC, pressure at 200atm, a pH of 5.2-7, and 1.0-5.0% NaCl leads experts to a conclusion that these organisms must have modified and unique adaptations for optimal growth at high temperature, high pressure, and also moderate salinity level (Tsoka et. al, 2003). Moreover, cellular structure of M. jannaschii revealed that they are irregular cocci (have irregular spherical shape). Their motility is by utilizing polar bundles of flagella (Tsoka et. al., 2003). Characteristic to any organism that belongs to the kingdom Archaea, M. jannaschii lacks cell wall; however, they do possess cell envelope that is made up of cytoplasmic membrane and a protein surface layer (S-layer) (Sleytr et. al., 2007).
Methanococcus jannaschii belongs to a specific group called methanogens, or methane producers (Tumbula and Whitman, 1999). Methanogens are physiologically specialized to undergo fueling reactions to produce methane as the end product (Reeve, 1992). They are ultimately autotropic single-celled organisms. Being an autotropic organism, M. jannaschii is strictly anaerobic and uses only carbon dioxide as its sole carbon source. Its main pathway for energy production is through methanogenesis, a process during which hydrogen is used as an energy source to reduce carbon dioxide to methane (Zhu et. al., 2004). Methanogens are extremely important to anaerobic environments because they convert organic compounds into methane, which then rises into the aerobic environment. By doing so, these organisms provide a pathway for compounds that exist in anaerobic environment to escape into the atmosphere, where it acts as a natural gas resource (Reeve, 1992).
M. jannaschii became the first organism in the archaea to have its complete genome sequenced (Bult, C. J. et. al, 1996). Its genomic structure, along with the unique lifestyle, has gradually provided insights into understanding the organism’s adaptations to its extreme habitat. More importantly, the knowledge gained from studying the metabolic processes, the enzymes, and the proteins that are specifically involved in these processes will lead to deeper understanding of the evolution of Archaea. In studying the interaction between M. jannaschii and its environment, many evolution questions about our environments could potentially be answered.
Genome structure
Methanococcus jannaschii was the first organism of the kingdom Archaea to be completely sequenced in 1995 (Bult, C.J. et. al., 1996). Three characteristic elements that make up the genome of M. jannaschii are: a large circular chromosome, a large-circular extra chromosome, and a small circular extra-chromosome (Bult, C.J. et. al., 1996). Using whole-genome random sequencing, it was found that:
1. The large circular chromosome has a length of 1.66 mega base pair. Within the chromosome, there are a total of 1729 protein-coding regions, and the percentage of G+C content is 31.4%.
2. The large circular extra-chromosome has a length of 58 kilo base pair. The total protein-coding regions are 45, and the percentage of G+C content is reported as 28.2%.
3. The small circular extra-chromosome contains 16.5 kilo base pair. The predicted protein-coding regions are 12, and the G+C content percent is 28.9%.
The genome structure of Methanococcus jannaschii finally provided solid support for the hypothesis that Archaea is indeed a separate domain of life from Bacteria and Eukarya, and that Archaea is more closely related to Eukarya. This hypothesis was made several years before genome sequencing even existed (Tsoka et. al., 2003). Although the M. jannaschii genome is quite small compared to the genome of well-known species such as Eschericia coli (only 40% of E. coli genome), researchers believe that the genome is very specific to the autotropic lifestyle of the organism (Zhu et. al., 2004). Until now, only about 52% of the proteins in the genome have been identified and assigned to functions (Zhu et. al., 2004). However, their hyperthermophillic properties and their special mechanisms to their extreme environments leads to conclusion that these proteins are strictly involved in energy production, specifically methanogenesis, cell division, metabolism, and also maintaining the organism’s lifestyle (Zhu et. al., 2004).
Cell structure and metabolism
1. Features of cell structure of archaeal organisms.
Cellular structure and metabolism of M. jannaschii allow for a deeper understanding of the domain Archaea, and also the evolutionary relationship between this domain and the other two, Bacteria and Eukarya. M. jannaschii belongs to the domain Archaea, and therefore exhibit cellular structures that are characteristics of this domain. They are unique because they do not have a cell wall made of peptidoglycan (called murein) like most bacteria. Most archaeal species have walls made of proteins or glycoproteins, which form the outermost envelope of the cell called the surface layer (S-layer) (Sleytr. et. al., 2007). Because of the lack of a cell wall, this explains why Archaea are resistant to antibacterial antibiotics, which are often used as genetic markers (Tumbula and Whitman, 1999). More remarkably, they are different from all other organism due to the glycerol ethers that make up their cytoplasmic membrane, while other species’ membranes are composed of phospholipids (glycerol diesters) (Sleytr. et. al., 2007). Equally distinctive is the fact that no known Archaea have been identified as pathogens (Tumbula and Whitman, 1999).
2. Proteins that have been identified using shotgun proteomics method
Even though the genome of Methanococcus janaschii have been sequenced and the predicted protein functions have been assigned, the proteome, which provides the products of the actual expressed genome, will eventually give greater insights into studying the importance of these macromolecules in the organism’s lifestyle. Using a shotgun proteome method, which is called Multidimensional Protein Identification Technology (MudPIT), researchers were able to identify the actual amount of proteins, and also assigned them with specific functions (Zhu. et. al., 2004).
M. jannaschii protein mixtures were first digested in a more complicated protein mixtures by allowing M. jannaschii to grow on a complex medium enriched with H2. This mixture was further loaded on a “multidimensional chromatographic capillary column,” which is then separated and resolved. With this method, hundreds of proteins were identified at the same time (Zhu. et. al., 2004). This method of protein identification was extremely convenient for it provided a faster and more efficient way of searching for functional proteins without having to separate each protein ahead of time. Out of 1775 genes predicted from the sequenced genome, 963 (54.2%) were identified. These genes are identified to be involved in amino acid biosynthesis, biosynthesis of cofactors, prosthetic groups, and carriers, DNA metabolism, energy metabolism, synthesis of purines, pyridines, nucleosides, and nucleotides, fatty acid and phospholipids metabolism, transcription and other cellular processes. Due to the abundance of these proteins in the genome, the report concluded that these proteins are “housekeeping” proteins that are crucial in allowing cell maintenance and growth (Zhu. et. al., 2004).
Out of the 963 genes identified, 137 or 9% were proposed to be involved in energy metabolism. This group of proteins appears to have a high abundance within the proteome. These proteins include many enzymes used in the process of methanogenesis, and also many electron carrier proteins such as ferredoxin and subunits of proton transporting ATP synthase. The second large portion found in the genome includes genes that function in protein synthesis, specifically ribosomal proteins. 67% of the predicted genes are involved in the synthesis of cell envelope. The majority of proteins that are expressed by these genes are S-layer structural proteins. About 80% of the predicted genes are involved in fatty acids and phospholipids metabolism. Even though a large amount of genes have unknown functions or are hypothetical genes, it was reported that these genes are most likely crucial to cellular growth (Zhu et. al., 2004).
3. Methanogenesis pathway of Methanococcus jannaschii
As introduced earlier, methanogenesis is the main biochemical pathway from which Methanococcus jannaschii obtains energy. This pathway depends of a rich supply of H2 to reduce CO2 to CH4 (Teske et. al., 2003). There are seven steps involved in the pathway, and each step is catalyzed by a specific enzyme (Zhu et. al., 2004). Since these enzymes are synthesized by hyperthermophiles, they are called hyperthermophillic enzymes (Vieille and Zeikus, 2001). Most interestingly, these enzymes are optimally active at high temperatures, temperatures that would denature most enzymatic functions. In the first step of methanogenesis, carbon dioxide is introduced into the pathway, forming formyl-methanofuran (formyl-MF). This step is catalyzed by the enzyme formyl-methanofuran dehydrogenase (FMD), which is a membrane-bound protein complex. The second step of the pathway produces formyl-tetrahydromethanopterin by transferring the formyl group on the formyl-methanofuran. Formyl-MF:H4MPT formyl transferase (FTR) catalyzes this reaction. In the third step, formyl-H4MPT is reduced to methenyl-H4MPT by the removal of a water molecule. The enzyme involved in this step is H4MPT cyclohydrolase (MCH). The next step of the pathway involves two very functionally distinct enzymes that have been proven to be characteristic in the energy-producing metabolism of methanogens. They are coenzyme F420-dependent, N5, N10-methylene-H4MPT dehydrogenase (MTD), and the other is an H2-dependent, H2-forming N5-N10-methylene-H4MPT dehydrogenase (HMD). The product molecule in this pathway is methylene-H4MPT. N5, N10-methylene-H4MPT reductase (MER) is also a coenzyme F420-dependent, and in the fifth step, it catalyzes the production of methyl-H4MPT from methylene-H4MPT. Under rich hydrogen condition, this enzyme was shown to be highly expressed. In the sixth step, methyl S-coenzyme M is synthesized with the catalysis of the transmembrane enzyme complex, N5-methyl-H4MPT:Coenzyme M methyltransferase (MTR). This enzyme has proven to be extremely important in conserving energy for Methanococcus jannaschii. When MTR transfers a methyl group to conenzyme M, it also creates a proton gradient across the membrane by acting as a primary sodium pump. The Na+/H+ antiporter serves to aid the formation of methane in the next step. Methane is finally produced in the last step of the pathway by methyl conenzyme M reductase.
Ecology
In studying anaerobic microbial communities and their interaction, the sulfate reduction, methanogenesis, and anaerobic methane oxidation are often of great interest not only because there is a close relationship between these pathways, but also because they serve as potential genomic markers to study the evolution of the biochemical pathways of organisms that live in these extreme environments (environments that are believed to resemble that of the ancient Earth’s biosphere). Sulfate reduction and methanogenesis are both anaerobic pathways that convert bacterial organic products during fermentation or degradation to CO2 and methane (Teske et. al., 2003). The availability of sulfate determines the amount of sulfate-reducing organisms. When there is a depletion of sulfate, methanogenic archaea become the dominated species, metabolizing CO2 and H2 to produce methane (Teske et. al., 2003). If methane and sulfate both exist at the same time, anaerobic methane oxidation takes place; this is a process during which methane is oxidized to CO2, using sulfate as the electron acceptor (Teske et. al., 2003). These processes and their interactions become extremely crucial to the hydrothermal vent microbial community because the products of one process become the substrates for other organisms undergoing other biochemical pathways.
In methanogenesis, many substrates for Methanococcus jannaschii such as CO2, formate, and acetate are formed by fermentative bacteria that live in anaerobic environments (Reeve, 1992). By converting these compounds to a gaseous product, methane, Methanococcus jannaschii, or methanogens in general, provide a pathway for these organic compounds to escape from the hydrothermal vents into aerobic environments (Teske et. al, 2003). Thus, methanogenesis and the organisms that use this pathway are extremely important in the carbon cycle. A small portion of methane is used by anaerobic methane oxidizing organisms; however, a large portion of it escapes into the atmosphere, where it serves as a very potent greenhouse gas (Zhu et. al, 2004). Even though methane is a potential energy source, the vast amount of methane produced within the past few years have raised many concerns on the Earth’s atmosphere (Zhu et. al., 2004). The increasing amount of methane released in the ocean and in the atmosphere have had some implications on the increasing fuel-burning rate, as well as the increasing concentration of carbon dioxide that humans release. These facts contribute to people’s knowledge about the human damaging effects on global warming that we are now suffering from.
Application to Biotechnology
Methanococcus jannaschii has the ability to produce many unique cofactors, coenzymes, and enzymes during methanogenesis. These macromolecules have been the target of many biotechnological applications (Tumbulan and Whitman, 1999).
Recently, it has been found that coenzyme M (CoM) is strictly involved in the methanogenesis of Methanococcus jannaschii (Graham et. al., 2003). It is the only terminal methyl carrier that all methyl intermediates bind to in order to transfer their methyl groups. Coenzyme M is a very unique coenzyme due to its sulfonate group that allows enzymes such as methyl coenzyme M reductase to bind in order to do its job. CoM is important because its chemical synthesis has been therapeutically used as a “mucolytic” and “uroprotective” agent during chemotherapy. The biosynthesis of CoM was recently found to occur by the sulfite addition mechanism. As a result, a pathway of biosynthesis of CoM from phosphoenolpyruvate (PEP) has been proposed based upon analysis of the structure of CoM isolated from Methanococcus jannaschii (Graham et. al., 2003).
Many other biotechnical applications take advantages of other enzymes such as dehydrogenases
Current Research
Many in vivo genetic studies have been conducted with the goal to understand more about the lifestyle of not only Methanococcus jannaschii, but also the archaeal evolution and its relationship to the other two domains of life (Tumbula and Whitman, 1999).
One extremely interesting area of research is of course focusing on hyperthermophillic enzymes that Methanococcus jannaschii, as well as other methanogenic species, produces. These hyperthermophillic enzymes have become models to study enzyme evolution and enzyme catalytic mechanisms. Current research has been targeting on mutagenesis in order to understand more about the temperature limitation for enzyme catalytic activity. Since these enzymes are optimally active at high temperature, questions have been raised whether these enzymes could be modified to be also active at extremely low temperature (Vieille and Zeikus, 2001).
Another area of current research studies protein synthesis, especially proteins that are involved in making up the cell envelope, or the S-layer of archaeal species. In 2007, it has been discovered that glycoproteins forming the S-layers are the most common component in the cell envelope, making it the basic building blocks of surface layers in archaeal organisms (Sleytr et. al., 2007).
Current research has also focused on flagellin biosynthesis. So far, studies have shown that flagellins in archaeal species are unrelated to that of bacterial species. Further studies aim to see the minimum amount of flagella needed for motility, and also the assembly mechanism of flagella in Archaea (Tumbula and Whitman, 1999).
References
1. Bult, C.J., White, O., Olsen, G.J., Zhou, L., Fleischmann, R.D., Sutton, G.G., et al. “Complete genome sequence of the methanogenic archaeon, Methanococcus jannaschii.” Science 1996. Volume 273: 1058–1073.
2. Graham, D., Xu, H., and White, R. “Identification of coenzyme M biosynthetic phosphosulfolactate synthase: a new family of sulfonate-biosynthesizing enzymes.” J Biol Chem. 2002. Volume 277(16):13421-9.
3. Reeve, J.N. “Molecular biology of Methanogens.” Annu Rev Microbiol. 1992. Volume 46: 165–191.
4. Sleytr, UB., Egelseer, E., Ilk, N., Pum, D., and Schuster, B., “S-Layers as a basic building block in a molecular construction kit.” 2007. Volume 274(2): 323-334.
5. Teske, A., Dhillon, A., and Sogin, M., “Genomic Markers of Ancient Anaerobic Microbial Pathway: Sulfate Reduction, Methanogenesis, and Methane Oxidation.” Biol. Bull. 2003. Volume 204: 186-191.
6. Tsoka, S., Simon, D., and Ouzounis, C., “Automated Metabolic Reconstruction for Methanococcus jannaschii.” Archaea. 2004. Volume 1: 223-229.
7. Tumbula, D., and Whitman. W., “Genetics of Methanococcus: possibilities for functional genomics in Archaea.” Molecular Microbiology 1999. Volume 33: 1-7.
8. Vieille C, Zeikus GJ, “Hyperthermophilic enzymes: sources, uses, and molecular mechanisms for thermostability.” Microbiol Mol Biol Rev. 2001. Volume 65(1):1-43.
9. Zhu W, Reich CI, Olsen GJ, Giometti CS, Yates JR 3rd., J. Proteome Res. 2004. Volume 3: 538-548.