Methylacidiphilum fumariolicum: Difference between revisions
No edit summary |
No edit summary |
||
| Line 14: | Line 14: | ||
Species: Methylacidiphilum fumariolicum | Species: Methylacidiphilum fumariolicum | ||
-The Phylum of Verrucomicrobia is derived "verruca" meaning "wart" and has been recognized only because of 16S rRNA sequencing. | |||
==Description and Significance== | ==Description and Significance== | ||
Revision as of 18:56, 9 April 2014
Classification
Domain: Bacteria
Phylum: Verrucomicrobia
Class:
Order: Methylacidiphilales
Family: Methylacidiphilaceae
Genus: Methylacidiphilum
Species: Methylacidiphilum fumariolicum
-The Phylum of Verrucomicrobia is derived "verruca" meaning "wart" and has been recognized only because of 16S rRNA sequencing.
Description and Significance
Methylacidiphilum fumariolicum is an thermoacidophilic autotrophic microbe first discovered in 2007 in volcanic pools near Naples, Italy by Huub Op den Camp and other scientists. This microbe not only endures but thrives in very hot temperatures (50-60 C) and mud that is extremely acidic (pH 2-5) by oxidizing methane. [1] M. fumariolicum can use ammonium, nitrate or atmospheric nitrogen as a nitrogen source and also fixes carbon dioxide using methane as an energy source. [5] After studying the mudpot in which the microbe lived, it was found that M. fumariolicum is strictly dependent on the presence of rare earth metals such as lanthanides which act as cofactors to a key enzyme in methanotrophs, methanol dehydrogenase. [1] What is most interesting is that when these microbes are inhibited of growth, they store glycogen in large amounts (up to 36% of their dry weight), which indicates that carbon storage is important for survival when methane is absent. [5]
Genome Structure
Genome Details [2] :
-The genome of M. fumariolicum is 2.36 Mbp in size.
-GC content = 40.9%
-2,283 protein encoding genes
-Biosynthetic pathways and tRNAs for all 20 amino acids were present.
-All known genes encoding glycogen metabolism were present and transcribed. [5]
-All known genes required for Calvin-Benson cycle were transcribed. [4]
Metabolism
M. fumariolicum uses both carbon and nitrogen fixation in order to survive in its' environment. Studies have shown that when nitrogen is limited and carbon is in excess, M. fumariolicum accumulates carbon-rich polymers, such as glycogen. When methane is removed, the stored glycogen is consumed. Bacteria with stored glycogen in the absence of methane survive for much longer amounts of time than bacteria without glycogen. This is the first evidence of glycogen storage in the phylum Verrucomicrobia and shows that carbon storage is a survival strategy used by M. fumariolicum when methane is absent in the environment.
Energy
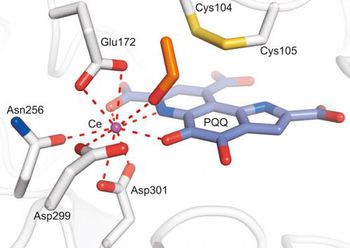
M. fumariolicum obtain their energy through the oxidation of methane in anaerobic conditions. Rare earth metals, such as lanthanum (Ln), cerium (Ce), praseodymium (Pr) and neodymium (Nd), are essential co-factors for the enzyme methanol dehydrogenase, which is used to metabolize the methanol produced from the oxidation of methane. [1]
Carbon Fixation
Methylacidiphilum fumariolicum has been found to be an autotroph. Its main carbon source is CO2 and is fixed via the Calvin Benson Bassham cycle. Studies have shown that no growth in a Methylacidiphilum fumariolicum colony is seen when CO2 concentrations are less than 0.3% (v/v). Further verification of Methylacidiphilum fumariolicum means of CO2 fixation was seen in a genome analysis.
Nitrogen Fixation
Methylacidiphilum fumariolicum has been found to fix nitrogen at low oxygen concentrations (less than 2% (v/v)) in chemostat cultures based on the following equation:
The optimal oxygen concentration for Methylacidiphilum fumariolicum nitorgen fixation is 0.5% (v/v). Studies show that this species poses the essential genes for nitrogen fixation (The genes encoding the structural protein (nifH, nifD and nifK) and the genes encoding cofactor biosynthesis (nifE,nifN and nifX)). Methylacidiphilum fumariolicum was also found to be more oxygen sensitive than most other proteobacterial methanotrophs. This could be due to the fact that during nitrogen fixation there was found to be nitrogenase activity, which is known to be oxygen sensitive. [3]
Ecology
References
[1] Pol, A., T. Barends, A. Dietl, A. Khadem, J. Eygensteyn, M. Jetten, and H. Op den Camp. "Rare Earth Metals Are Essential for Methanotrophic Life in Volcanic Mudpots." Environmental Microbiology (2013): N/a. Print.
[2] Khadem, A., A. Wieczorek, A. Pol, S. Vuilleumier, H. Harhangi, P. Dunfield, M. Kalyuzhnaya, J. Murrell, K. Francoijs, H. Stunnenberg, L. Stein, A. Dispirito, J. Semrau, A. Lajus, C. Medigue, M. Klotz, M. Jetten, and H. Op Den Camp. "Draft Genome Sequence of the Volcano-Inhabiting Thermoacidophilic Methanotroph Methylacidiphilum Fumariolicum Strain SolV." Journal of Bacteriology 194.14 (2012): 3729-730. Print.
[3] Khadem, A. F., A. Pol, M. S. M. Jetten, and H. J. M. Op Den Camp. "Nitrogen Fixation by the Verrucomicrobial Methanotroph 'Methylacidiphilum Fumariolicum' SolV."Microbiology 156.4 (2010): 1052-059. Print.
[4] Khadem, A. F., A. Pol, A. Wieczorek, S. S. Mohammadi, K.-J. Francoijs, H. G. Stunnenberg, M. S. M. Jetten, and H. J. M. Op Den Camp. "Autotrophic Methanotrophy in Verrucomicrobia: Methylacidiphilum FumariolicumSolV Uses the Calvin-Benson-Bassham Cycle for Carbon Dioxide Fixation." Journal of Bacteriology 193.17 (2011): 4438-446. Print.
[5] Khadem, A., M. Teeseling, L. Niftrik, M. Jetten, H. Op Den Camp, and A. Pol. "Genomic and Physiological Analysis of Carbon Storage in the Verrucomicrobial Methanotroph “Ca. Methylacidiphilum Fumariolicum” SolV." Frontiers in Microbiology 3 (2012): n. pag. Print.
Edited by Keely Chandler and Kelsey Sharples, students of Drs. Kaz Kashefi and Edward Walker for MMG 425 at Michigan State University, 2014.