Mycobacterium avium subspecies paratuberculosis: Difference between revisions
| Line 41: | Line 41: | ||
==Cell structure and metabolism== | ==Cell structure and metabolism== | ||
''Map'' is a small, slow-growing, gram-positive, acid-fast positive, rod-shaped, | ''Map'' is a small, slow-growing, gram-positive, acid-fast positive, rod-shaped, multispecies pathogenic bacteria It is aerobic, has no mechanisms of motility and has a cell wall made up of a waxy mixture of lipids and polysaccharides [3, 10]. The cell wall of a pathogen must act as an interface between the cell and its host, and thus will structurally display that parasite’s virulence. In the case of ''Map'', a thick waxy cell wall made up of 60% complex lipids not only gives it the properties of an acid fast bacteria (the ability to resist decolorization by acidified alcohol), but it also creates hydrophobicity and an increased resistance to low pH, high temperature, and a variety of chemicals, increasing its longevity and pathogenic potential in harsh and varied environments [8, 13]. Studies have shown that virulence may be increased in ''Map'' by the presence of two copies of the five genes (fabG2, accD4, kasB, fabG3 and fabG5) involved in the fatty acid biosynthetic pathway through the combined action of these fatty acid synthases and polyketide synthases to create the complex lipid components of the membrane [9]. | ||
A thick cell wall also contributes to the slow growth rate of ''Map'' by decreasing the flow of nutrients into and out of the cell. ''Map'' is actually the slowest growing of all cultivated mycobacteria; under optimal conditions, it takes over 20 hours to generate. A ''Map'' isolate can produce small colonies within 3-6 weeks when grown in the optimal temperature of 37 degrees C under aerobic conditions [13]. As an aerobic organism, ''Map'' requires oxygen in its environment for metabolism; in order to survive in low-oxygen environments, it produces isocitrate lyase, an essential anapleurotic enzyme which assists in survival within microaerophilic conditions inside the host. In terms of metabolism, ''Map'' has a functional glycolytic pathway and tricarboxylic acid (TCA) cycle. The presence of an additional FAD binding domain in the ''fdhF'' gene, and the intact iron-sulfur domain within the ''fdxB'' gene possibly indicate an alternate survival strategy for ''Map'' which would be similar to that of the anaerobic survival of ''M. Tuberculosis'' under starvation [9]. | A thick cell wall also contributes to the slow growth rate of ''Map'' by decreasing the flow of nutrients into and out of the cell. ''Map'' is actually the slowest growing of all cultivated mycobacteria; under optimal conditions, it takes over 20 hours to generate. A ''Map'' isolate can produce small colonies within 3-6 weeks when grown in the optimal temperature of 37 degrees C under aerobic conditions [13]. As an aerobic organism, ''Map'' requires oxygen in its environment for metabolism; in order to survive in low-oxygen environments, it produces isocitrate lyase, an essential anapleurotic enzyme which assists in survival within microaerophilic conditions inside the host. In terms of metabolism, ''Map'' has a functional glycolytic pathway and tricarboxylic acid (TCA) cycle. The presence of an additional FAD binding domain in the ''fdhF'' gene, and the intact iron-sulfur domain within the ''fdxB'' gene possibly indicate an alternate survival strategy for ''Map'' which would be similar to that of the anaerobic survival of ''M. Tuberculosis'' under starvation [9]. | ||
Revision as of 13:36, 29 August 2007
A Microbial Biorealm page on the genus Mycobacterium avium subspecies paratuberculosis
.
Classification
Higher order taxa
Bacteria (Domain); Actinobacteria (Phylum); Actinobacteria (Class); Actinobacteridae (Subclass); Actinomycetales (Order); Corynebacterineae (Suborder); Mycobacteriaceae (Family); Mycobacterium (Genus); Mycobacterium avium complex (MAC) (Species group) [11]
Species
Mycobacterium avium
Subspecies
Mycobacterium avium subspecies paratuberculosis
Commonly referred to as MAP or Map
Also known as: Mycobacterium paratuberculosis, Mycobacterium johnei, Bacillus paratuberculosis, Bacterium paratuberculosis, Mycobacterium enteritidis, Darmtuberculose, and Mycobacterium avium paratuberculosis [11]
The classification of this organism has been subject to debate. Isolated over 100 years ago, it was originally termed Mycobacterium enteritidis chronicae pseudotuberculosae bovis johne [13]. Current classifications depend on genetic composition and phenotypic characteristics. With a genome almost identical to that of Mycobacterium avium (M. avium) and its subspecies, the International Association for Paratuberculosis has recognized the subspecies term (Map) as fitting. On the other hand, some researchers and organizations still use the name M. paratuberculosis, not only for simplicity, but as a result of its significant phenotypic variances from M. avium [3, 8].
Description and significance
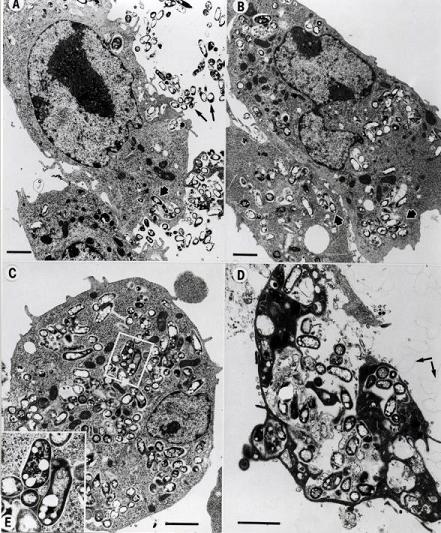
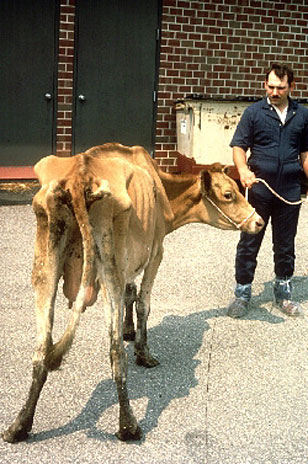
Mycobacterium avium subspecies paratuberculosis (Map) is the etiologic agent of Johne’s disease (pronounced “YO-nees” [8]) also known as paratuberculosis. It was first isolated by German scientists H.A. Johne and L. Frothingham over a century ago from diseased animals displaying inflammation of the ileum [13]. Johne’s disease, named for one of its finders, is a mammalian disease that resembles some clinical and pathological aspects of Crohn’s disease in humans [2], and which has become an increasingly relevant problem for cattle farmers and the economy worldwide. Map is found primarily in ruminants (cattle, sheep, goats, deer, antelope, bison, etc) and their three-chambered stomach relatives [3]. Johne’s disease has been studied extensively in dairy cattle, where it has been called one of the most serious diseases affecting these animals to date [4]. It is estimated that 20% of dairy herds and 8% of beef herds in the United States are infected [6], leading to a $1.5 billion loss for the cattle industry each year. Adding to this problem, Map has also been found to infect a variety of other wild animals, such as badger, fox, primates, rabbits, and swine [1]. Different strains of Map have been found equally distributed throughout the affected species (the same strain of bacteria is able to infect multiple types of animals), contributing to this wide host range and ease of transfer between species [8], all of which make the bacteria and disease very hard to keep track of. Also making Map hard to track is its long incubation period in host organisms. Though animals are infected at a young age, they don’t begin to show visible symptoms until adulthood, at which point they have reached the critical stages of disease and rarely survive [3, 5].
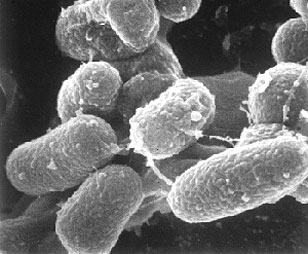
Map, an obligate intracellular pathogen, is a small (0.5 x 1.5 micron [3]), rod shaped bacteria which grows in circular colonies reaching about 1-2 mm in diameter [13]. It is usually found to be off-white or yellow depending on the medium [3], and has the ability to adapt easily to many environments. Map not only can evade host defenses and survive within phagosomal compartments for more than two weeks, but it has also been found to reprogram the pattern of gene expression of bovine macrophages to increase its chance of survival and pathogenesis within the host [1, 13]. Though it requires a suitable cell medium for growth, Map can survive in the environment well beyond its presence in an animal host, creating drastic problems for farmers trying to eradicate Johne’s disease from their herds.
Due to the steady increase of Map infection prevalence throughout cattle farms of the world and the economic impacts which result, methods of detecting Map from environmental cultures are currently being studied and may provide insight into the infection mechanisms of this pathogen and thus provide a basis for the development of drugs which may eradicate Johne’s disease altogether. Also, the close relation between characteristics of Johne’s disease in ruminants and Crohn’s disease in humans (a chronic inflammatory bowel disease) has created an ongoing debate of whether or not Map is a possible causative agent of this human disease as well. Though it has been fully sequenced, further functional characterization of the Map genome may solve this mystery between Map and Crohn’s by increasing our understanding of the molecular biology as well as the physiology, pathogenesis and host specificity of this bacterium [9].
Genome structure
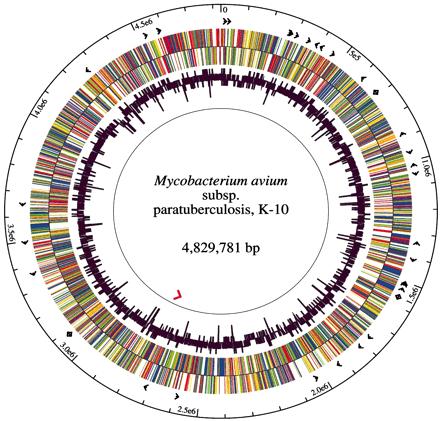
The Map strain used for sequencing, K-10, was originally isolated from a dairy herd in Wisconsin by investigators of the USDA National Animal Disease Center [10]. This strain is a typical bovine type II clinical isolate which was chosen because of its high efficiency of transformation with plasmid DNA [1], its virulence and its amenability to transposition mutagenesis. By random shotgun sequencing, 1.8- to 3.0-kb fragments were cloned into pUC18 and 66,129 sequences were used to create the final genome assembly. As represented in the figure, the K-10 Map genome has a single circular chromosome of 4,829,781 base pairs, 91.3% of which are protein coding. Map has a GC (guanine cytosine) content of 69.3%, an average gene density of 1,112 base pairs per gene and an average gene length of 1,015 base pairs. This genome encodes 4,350 ORFs, 45 tRNAs and 1 rRNA operon. 39 protein sequences were found within the Map genome which have no identifiable homolog recorded as of yet. About 1.5% of the Map genome consists of repetitive DNA [7].
Of the 19 insertion sequences identified in the K-10 genome, most represent homologs of other Mycobacterium IS elements. IS_MAP02 and IS_MAP04 are of interest because they have no homologs in other mycobacteria and thus could possibly be used as specific diagnostic targets for Map. The Map genome encodes for all proteins involved in the following metabolic pathways: glycolysis, the pentose phosphate pathway, the tricarboxylic acid cycle, and the glyoxalate cycle. Also, the large number of genes for regulatory function found within the genome is consistent with Map’s ability to survive in a wide range of environmental conditions. One major difference between Map and other mycobacteria is its inability to produce mycobactin in vitro, which is a siderophore responsible for the binding and transport of iron into the cell. A sequence comparison of the mycobactin gene cluster between Map K-10, M. avium strain 104 and M. tuberculosis strain H37Rv reveals a possible reason for this deficiency. Within this cluster, the mbtA gene is much shorter in Map than the other two, lacking more than 200 residues important for protein function. Though this remains to be tested more thoroughly, this truncated domain may be the limiting factor in mycobactin production for Map since MbtA has been found to initiate siderophore synthesis. One additional Map genome feature to note is the extentsive functional redundancy; a result of gene duplication, this redundancy is based on amino acid content and is usually high among genes involved in lipid metabolism and oxidoreduction [7]. A greater concentration proteins for lipid metabolism in Map compared to other species may reflect its requirement of a robust cell wall which allows it to survive and colonize in the ruminant intestine [9].
Cell structure and metabolism
Map is a small, slow-growing, gram-positive, acid-fast positive, rod-shaped, multispecies pathogenic bacteria It is aerobic, has no mechanisms of motility and has a cell wall made up of a waxy mixture of lipids and polysaccharides [3, 10]. The cell wall of a pathogen must act as an interface between the cell and its host, and thus will structurally display that parasite’s virulence. In the case of Map, a thick waxy cell wall made up of 60% complex lipids not only gives it the properties of an acid fast bacteria (the ability to resist decolorization by acidified alcohol), but it also creates hydrophobicity and an increased resistance to low pH, high temperature, and a variety of chemicals, increasing its longevity and pathogenic potential in harsh and varied environments [8, 13]. Studies have shown that virulence may be increased in Map by the presence of two copies of the five genes (fabG2, accD4, kasB, fabG3 and fabG5) involved in the fatty acid biosynthetic pathway through the combined action of these fatty acid synthases and polyketide synthases to create the complex lipid components of the membrane [9].
A thick cell wall also contributes to the slow growth rate of Map by decreasing the flow of nutrients into and out of the cell. Map is actually the slowest growing of all cultivated mycobacteria; under optimal conditions, it takes over 20 hours to generate. A Map isolate can produce small colonies within 3-6 weeks when grown in the optimal temperature of 37 degrees C under aerobic conditions [13]. As an aerobic organism, Map requires oxygen in its environment for metabolism; in order to survive in low-oxygen environments, it produces isocitrate lyase, an essential anapleurotic enzyme which assists in survival within microaerophilic conditions inside the host. In terms of metabolism, Map has a functional glycolytic pathway and tricarboxylic acid (TCA) cycle. The presence of an additional FAD binding domain in the fdhF gene, and the intact iron-sulfur domain within the fdxB gene possibly indicate an alternate survival strategy for Map which would be similar to that of the anaerobic survival of M. Tuberculosis under starvation [9].
Mycobactin is a siderophore unique to the mycobacterial family. This chemical is secreted from the membrane in order to break down and transport iron (an essential nutrient for growth) into the cell. Map is unique in that it does not produce mycobactin on its own like the rest of its genus, and thus requires a host cell to provide iron, which normally is found in cell macrophages. Because of this, in order to grow Map cells in in vitro culture, they must be provided with mycobactin in the medium or they will not grow or reproduce [3]. This mycobactin deficiency can also be seen as a contributor to the bacteria’s slow growth rate [13].
Ecology
Describe any interactions with other organisms (included eukaryotes), contributions to the environment, effect on environment, etc.
Pathology
.
How does this organism cause disease? Human, animal, plant hosts? Virulence factors, as well as patient symptoms.
Application to Biotechnology
Does this organism produce any useful compounds or enzymes? What are they and how are they used?
Current Research
MAP and Crohn's Disease
Methods of Detection
Enter summaries of the most recent research here--at least three required
References
[ 1 ] Chacon, Ofelia; Bermudez, Luiz E.; and Barletta, Raul G. “Johne’s Disease, Inflammatory Bowel Disease, and Mycobacterium paratuberculosis.” Annual Reviews of Microbiology. 2004. 58: 329-63.
[ 2 ] Collins, Michael T. “Paratuberculosis: Review of present knowledge.” Acta veterinaria Scandinavica. 2003. 44: 217-221.
[ 3 ] Collins, Michael T. “Update on paratuberculosis: 1. Epidemiology of Johne’s disease and the biology of Mycobacterium paratuberculosis.” Irish Veterinary Journal. 2003. 56(11): 565-574.
[ 4 ] Cook, Kimberly L. and Britt, Jenks S. “Optimization of methods for detecting Mycobacterium avium subsp. paratuberculosis in environmental samples using quantitative, real-time PCR.” Journal of Microbiological Methods. 2007. 69: 154-160.
[ 5 ] Grant, Irene R. “Mycobacterium avium ssp. paratuberculosis in foods: current evidence and potential consequences.” International Journal of Dairy Technology. 2006. 59(2): 112-117.
[ 6 ] Jaravata, Carmela V.; Smith, Wayne L.; Rensen, Gabriel J.; Ruzante, Juliana; and Cullor, James S. “Survey of Ground Beef for the Detection of Mycobacterium avium paratuberculosis.” Foodborne Pathogens and Disease. 2007. 4(1): 103-106.
[ 7 ] Li, Lingling; Bannantine, John P.; Zhang, Qing; Amonsin, Alongkorn; May, Barbara J.; Alt, David; Banerji, Nilanjana; Kanjilal, Sagarika; and Kapur, Vivek. “The complete genome sequence of Mycobacterium avium subspecies paratubersulosis.” PNAS. 2005. 102(35): 12344-12349.
[ 8 ] Manning, Elizabeth J.B. “Mycobacterium avium subspecies paratuberculosis: a review of current knowledge.” Journal of Zoo and Wildlife Medicine. 2001. 32(3): 293-304.
[ 9 ] Marri, Pradeep Reddy; Bannantine, John P.; and Golding, Geoffrey B. “Comparative genomics of metabolic pathways in Mycobacterium species: gene duplication, gene decay and lateral gene transfer.” FEMS Microbiology Reviews. 2006. 30(6): 906-925.
[ 10 ] NCBI. “Mycobacterium avium subsp. paratuberculosis K-10 k10 project at Univ. Minnesota.” Entrez Genome Project.
[ 11 ] NCBI. “Mycobacterium avium subsp. paratuberculosis.” Taxonomy Browser. <>
[ 12 ] Prantera, C. “Mycobacterium and Crohn’s disease: The endless story.” Digestive and Liver Disease. 2007. 39: 452-454.
[ 13 ] Rowe, M.T. and Grant, I.R. “Mycobacterium avium ssp. paratuberculosis and its potential survival tactics.” Letters in Applied Microbiology. 2006. 42: 305-311.
[ 14 ] Windsor, P. “Research into vaccination against ovine Johne’s disease in Australia.” Small Ruminant Research. 2006. 62: 139-142.
Edited by Kyla Holmes, student of Rachel Larsen
