Orthoreovirus
Baltimore Classification
Higher order taxa
Virus; Reoviridae (Family); Orthoreovirus.
Species
There are over 150 species of Reoviridae.
Description and Significance
The orthoreovirus infects invertebrates, plants and vertebrates.
Genome Structure
The genome of orthoreovirus is monomeric, segmented and consists of ten segments of linear double-stranded RNA. The complete genome is 23700 nucleotides long. The L1 is sequenced but only an estimate is available. The sequence is 3800-3900 nucleotides long. L2 is sequenced and the complete sequence is 3800-3900 nucleotides long. L3 is sequenced too but only an estimate is given and the complete sequence is 3800-3900 nucleotides long as well. M1 has been fully sequenced and the complete sequence is 2200-2300 nucleotides long. M2 has been sequenced as well and is a long as M1 but only an estimate of the sequence is presented. S2 and S3 are sequenced and are 1200-1400 nucleotides long but only an estimate is presented. S4 has also been completely sequenced and is 1200-1400 nucleotides long. The 5'-end of the genome has a methylated nucleotide cap. On the positive strand of each duplex, the negative strands have a phosphorylated terminus and the cap sequence type is m7G5ppp5'GmpNp. The multipartite genome is found in one type of particle only. Each virion contains a single copy of the genome but it may be a full lenght copy or shorter copies. (source: ICTV dB Descriptions)
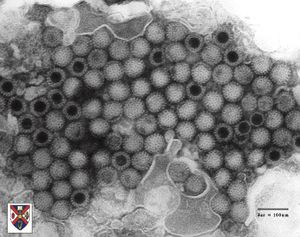
Virion Structure of an Orthoreovirus
The virions of an orthoreovirus consist of a capsid, a core and a nucleoprotein complex. The virus capsid is not envelpped, isometric with icosahedral symmetry and has a diameter of 80-82 nm. The capsid shells of virions are composed of two layers. All the shells are usually present but sometimes the outer shell is lost during preparation. The capsids appear round and the surface structure reveals a regular pattern with distinctive features. The capsomer arrangement is clearly visible and surface projections are not present. The inner capsids are the core and have a diametere of 60 nm. The core is spherical and consists of dsRNA genome with a diameter of 49 nm. The end of the fibers protrude almost through to the capsid surface. (source: ICTV dB Descriptions)
Reproductive Cycle of an Orthoreovirus in a Host Cell
The replication of the orthoreovirus occurs in the cytoplasm. The receptors are known to contain sialic acid, although most have not been definitively identified. The particles are internalized and partially uncoated in endolysosomes in the cytoplasm. They are resistant to protease digestion and the virus would be destroyed if completely uncoated. The sub-viral particle is the site of early transcription of the d/s RNA genome by viral polymerase. The various genome segments are transcribed or translated at different frequencies. The RNA is transcribed conservatively and only the (-)sense strands are used, resulting in the synthesis of (+)sense mRNAs that are capped inside the core. This whole process occurs without de novo protein synthesis. mRNAs leave the core and are translated into the cytoplasm.
Primary transcription results in capped transcriptions that are not polyadenylated and at least 5 enzymatic activities are present in orthoreovirus particles to carry out this process. Secondary transcription occurs later in infection in particles produced inside the infected cells and results in uncapped non-polyadenylated transcripts.
The genome is replicated in the cytoplasm in a conservative fashion. An excess if (+)sense strands are produced which serve as late mRNAs and as a template for (-)sense strand synthesis. The mechanism responsible for segregation of the various genome segments into developing particles is not known.
Viral Ecology & Pathology
The gastrointestinal and upper respiratory tracts are involved in most of the human orthoreovirus infections. The infections are asymptomic but occasionally produce mild febrile illness, or very rarely, serious complications.
