Photobacterium profundum: Difference between revisions
mNo edit summary |
No edit summary |
||
| Line 30: | Line 30: | ||
==Genome structure== | ==Genome structure== | ||
[[Image: | [[Image:CG16A.JPG |frame|Genomic organization of ''P. profundum'' strain SS9 chromosome 1 & 2. From www.ncbi.nlm.nih.gov.]] | ||
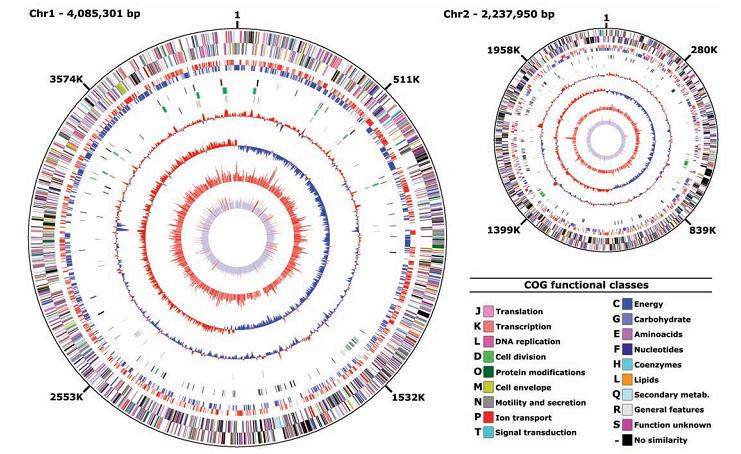
The genome of ‘’Photobacterium profundum’’ shows a tripartite structure consisting of a 4.1-Mbp major circular chromosome, a 2.2-Mbp minor circular chromosome, and a 80-kbp circular plasmid. SS9 has 14 ribosomal RNA (rRNA) operons on chromosome 1, and 1 on chromosome 2; it has the maximal number of rRNA so far identified in a bacterial genome. This may implies its characteristic as an extremophile, with the ability to respond rapidly to favorable changes in growth conditions. Various operons could have evolved to operate under particular physiological conditions. | The genome of ‘’Photobacterium profundum’’ shows a tripartite structure consisting of a 4.1-Mbp major circular chromosome, a 2.2-Mbp minor circular chromosome, and a 80-kbp circular plasmid. SS9 has 14 ribosomal RNA (rRNA) operons on chromosome 1, and 1 on chromosome 2; it has the maximal number of rRNA so far identified in a bacterial genome. This may implies its characteristic as an extremophile, with the ability to respond rapidly to favorable changes in growth conditions. Various operons could have evolved to operate under particular physiological conditions. | ||
Revision as of 06:11, 5 June 2007
A Microbial Biorealm page on the genus Photobacterium profundum
Classification
Higher order taxonomy
Bacteria; Proteobacteria, Gammaproteobacteria; Vibrionales; Vibrionaceae; Photobacterium
Properties
Presence of flagella: Yes
Interaction: No
Number of membranes: 2
Number of inteins: 0
Genetic code: Translation table 11 (Bacterial and Plant Plastid)
Taxonomy ID: 298386
Rank: no rank
Other names:
deep-sea eubacterium SS9; Photobacterium sp. (strain SS9); Photobacterium sp. SS9; Photobacterium SS9; Photobacterium profundum strain SS9; Photobacterium profundum str. SS9
Description and significance
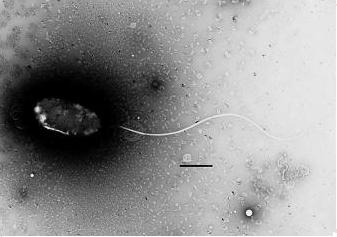
Members of the genus Photobacterium are common bacteria in marine environments. Photobacterium profundum is a newly discovered species that was isolated from sediment of the Ryukya trench off of Japan. It is barophillic and can survive under high pressures. It is a Gram-negative rod capable of growth between 4oC and 18oC (optimal at 10oC) under atmospheric pressure.
Genome structure
The genome of ‘’Photobacterium profundum’’ shows a tripartite structure consisting of a 4.1-Mbp major circular chromosome, a 2.2-Mbp minor circular chromosome, and a 80-kbp circular plasmid. SS9 has 14 ribosomal RNA (rRNA) operons on chromosome 1, and 1 on chromosome 2; it has the maximal number of rRNA so far identified in a bacterial genome. This may implies its characteristic as an extremophile, with the ability to respond rapidly to favorable changes in growth conditions. Various operons could have evolved to operate under particular physiological conditions.
SS9 has an unexpectedly high number of unique open reading frames (ORFs) in comparison to other several Vibrionaceae genomes sequenced. 38.6% of the ORFs are unique for chromosome 2, where 18.7% are unique for chromosome 1. Transposons are also found at a higher frequency on chromosome 2, suggesting the idea that chromosome 1 is more stable and containing the majority of the “established” genes, whereas chromosome 2 acts as a “genetic melting pot.”
Cell structure and metabolism
Several stress-response genes are activated at atmospheric pressure for SS9. Four genes, naming the ‘’htpG, dnaK, dnaJ, and groEL’’, up-regulated at 0.1MPa are involved in protein folding and in response to stress conditions. There is transcriptional induction of the glycolytic pathway and trehalose phosphotransferase system to be up-regulated under stress conditions.
The transport of amino acids such as Lys, His, Leu and Trp is reduced at high pressure, owing to the volume change of activation of the transport process. These transporters may have also evolved a particular protein structure to adapt to elevated pressure. The sensitivity of SS9 to low pressure is evident by the activation of different chaperones and DNA repair proteins at atmospheric pressure.
Ecology
Pathology
Application to Biotechnology
Current Research
Eric Allen, Assistant Professor of Marine Biology from Scripps Institution of Oceanography, University of California, San Diego is conducting a research on Photobacterium profundum. Allen’s recent publications are listed below:
• 1999. "Monounsaturated but not polyunsaturated fatty acids are required for growth of the deep-sea bacterium Photobacterium profundum SS9 at high pressure and low temperature" Appl. Environ. Microbiol. 65, 1710-1720.
• 2000. "FabF is required for piezoregulation of cis-vaccenic acid levels and piezophilic growth of the deep-sea bacterium, Photobacterium profundum strain SS9" J. Bacteriol. 182, 1264-1271.
• 2002. "Structure and regulation of the omega-3 polyunsaturated fatty acid synthase genes from the deep-sea bacterium Photobacterium profundum strain SS9" Microbiology 148, 1903-1914.
References
Yuichi Nogi, Noriaki Masui and Chiaki Kato. 1998. “Photobacterium profundum sp. nov., a new, moderately barophilic bacterial species isolated from a deep-sea sediment.” Extremophiles, 2: 1-7.
A. Vezzi, S. Campanaro, M. D'Angelo, F. Simonato, N. Vitulo, F.M. Lauro, A. Cestaro, G. Malacrida, B. Simionati, N. Cannata, C. Romualdi, D.H. Bartlett, G. Valle. 2005. "Life at Depth: Photobacterium profundum Genome Sequence and Expression Analysis." Science, 307: 1459-1461.
Edited by Karen Lai, student of Rachel Larsen at UCSD.