Photobacterium profundum: Difference between revisions
No edit summary |
No edit summary |
||
| Line 1: | Line 1: | ||
{{Uncurated}} | |||
{{Biorealm Genus}} | {{Biorealm Genus}} | ||
Revision as of 19:03, 19 August 2010
A Microbial Biorealm page on the genus Photobacterium profundum
Classification
Higher order taxonomy
Bacteria; Proteobacteria, Gammaproteobacteria; Vibrionales; Vibrionaceae; Photobacterium
Properties
Presence of flagella: Yes
Number of membranes: 2
Number of inteins: 0
Genetic code: Translation table 11 (Bacterial and Plant Plastid)
Taxonomy ID: 298386
Rank: no rank
Other names:
deep-sea eubacterium SS9; Photobacterium sp. (strain SS9); Photobacterium sp. SS9; Photobacterium SS9; Photobacterium profundum strain SS9; Photobacterium profundum str. SS9
Description and significance
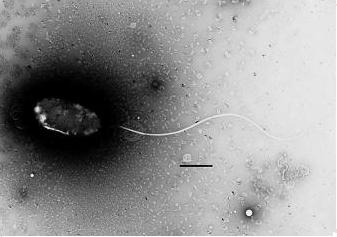
Members of the genus Photobacterium are bacteria in marine surroundings. Photobacterium profundum is a newly discovered species that was isolated from sediment of the Ryukya trench off of Japan. It is barophilic and can survive under high pressures. It is a Gram-negative rod that can potentially grow at temperature between 4oC and 18oC (optimal at 10oC) under atmospheric pressure, and at pressures between 0.1MPa and 70MPa (optimal at 10MPa) at 10oC. [1]
16S rRNA sequence of the Photobacterium profundum strain DSJ4 is closely related to the strain SS9 from the Vibrionaceae family within the Photobacterium subgroup. In the research study of Y. Nogi and et al., these strains are compared with Photobacterium angustum, Photobacterium damselae, Photobacterium phosphoreum, and Photobacterium histaminum. Strains DSJ4 and SS9 both have different temperature and pressure ranges for growth, cellular fatty acid composition and assorted physiological and biochemical characteristics, and DNA structures from the other Photobacterium species. [1] These differences lead to the creation of a new classification, Photobacterium profundum to represent this new deep-sea-type species.
Genome structure
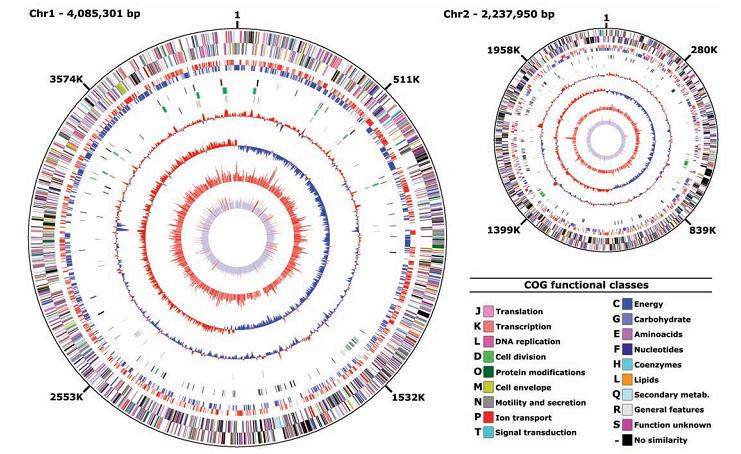
The genome of Photobacterium profundum consists of a 4.1-Mbp major circular chromosome, a 2.2-Mbp minor circular chromosome, and a 80-kbp circular plasmid. SS9 has 14 ribosomal RNA (rRNA) operons on chromosome 1, and 1 on chromosome 2; it has the maximal number of rRNA so far identified in a bacterial genome. This may implies its characteristic as an extremophile that responds to extreme growth conditions differently than the majority of the terrestrial mesophile organisms. Various operons could have evolved to operate under particular physiological conditions. [2]
Chromosome 1 contains the most “established” genes, and it is stable. Chromosome 2 serves as the “genetic melting pot” or a target of gene transfer, containing a high number of unknown genes. The lack of sunlight in deep sea causes its absence of the ORFs encoding light-activated photolyase genes. Photobacterium profundum contains a number of unique open reading frames (ORFs) that are not found in its Vibrionaceae family. The majority of transcription takes place in chromosome 1, where a minimal amount of transcription occurs in chromosome 2 since genes are poorly expressed there.
Cell structure and metabolism
The cells of Photobacterium profundum are Gram-negative, rod shape, 2-4μm long and 0.8-1.0 μm wide, with a single unsheathed flagella.
Strain DSJ4 is an anaerobic chemoorganotroph, with two types of metabolism – respiratory and fermentative types. Its microbial ability to use respiratory metabolism is proven with its positive result in catalase and cytochrome oxidase test. It uses nitrate as an proton acceptor which will be reduced to nitrite. This organism does not produce H2S as the fermentation product. Photobacterium profundum utilizes glycogen, D-galactose, D-glucose, maltose, D-mannitol, D-mannose, D-trehalose and glycerol as the sole carbon and energy sources. The presence of NaCl in the medium, the temperature between 4oC and 18oC, and the pressures between 0.1MPa and 70MPa are essential for strain DSJ4 to grow. [3] The fatty acid in DSJ4 is primarily hexadecenoic acid, and significant amounts of 14-methylpentadecanoic acid, octadecenoic acid, and icosapentaenoic acid. [1]
Several stress-response genes are activated at atmospheric pressure for SS9. Four genes (htpG, dnaK, dnaJ, and groEL) up-regulated at 0.1MPa, are involved in protein folding and in response to stress conditions. There is transcriptional induction of the glycolytic pathway and trehalose phosphotransferase system to be up-regulated under stress conditions.
Ecology
Omega-3 polyunsaturated fatty acid is essential for human health. Eicosapentaenoic acid (EPA) and docosahexaenoic acid (DHA) are needed for proper human development, prevention of some cancers and promotion of cardiovascular health. Photobacterium profundum strain SS9 includes four of the five genes required for EPA biosynthesis, and includes large multidomain proteins in its amino acids sequences that will catalyse EPA biosynthesis. [7]
Phylogenetic analysis
The phylogenetic analyses of the 16S rRNA gene sequence reveals that Photobacterium profundum is closely related to Hyphomicrobium indicum. gyrB, a DNA gyrase gene, is also employed as another phylogenetic marker to provide more reliable information of the evolutionary position of Photoprofundum and H. indicum. In comparison, Photobacterium profundum has a unique signature nucleotide sequence (5’-TTCATTACGAGCGG-3’) at positions 345-359, according to the E. coli numbering system. [5] These closely related organisms are differentiated by their various phenotypic characteristics, namely the optimum growth environment, morphology, catalase, oxidase, metabolic pathways, sucrose utilization and H2S production.
Pathology
The transmembrane DNA binding protein ToxR and the associated membrane protein ToxS can be found in many ocean species that are human or fish pathogens, and may cause gastroenteritis associated with seafood consumption. Photobacterium profundum contains ToxR; however, it is believe to be nonpathogenic to mammals. [8]
Application to Biotechnology
The sequencing of Photobacterium profundum was completed by the use of molecular methodologies. Molecular methodologies, the technique of extracting DNA from a sample, amplifying the interested gene using the polymerase chain reaction (PCR), and cloning and sequencing the resulting product, can help better understand the physiological and evolutionary diversity within the marine microbial community. Genome sequence of this unicellular marine organism provides example of how species with similar marine habitats can have evolved different gene complements that allow them to coexist with non-competitive, complementary functions. [4] This simple organism also provides a potential to discover hidden genetic and physiological complexity that might be applicable to future biotechnology.
Researchers at Scripps Institution of Oceanography at the University of California, San Diego have stated in their genomic research that these deep sea life forms, including Photobacterium profundum, “may help lay the groundwork for a variety of research outside of the deep ocean, including the application of deep sea bacterial genes for improving human nutrition and degrading pollutants, and calculating possibility for life in pressurized environments elsewhere in the solar system.” [6]
Current Research
Eric Allen, Assistant Professor of Marine Biology from Scripps Institution of Oceanography, University of California, San Diego is conducting a research on Photobacterium profundum. His emphases of research include the adaptation of deep-sea microbes to extreme environment, functional genomics, virulence determinants of the organism, and detection of pathogens in the marine environment. Allen’s recent publications are listed below:
• 1999. "Monounsaturated but not polyunsaturated fatty acids are required for growth of the deep-sea bacterium Photobacterium profundum SS9 at high pressure and low temperature" Appl. Environ. Microbiol. 65, 1710-1720.
• 2000. "FabF is required for piezoregulation of cis-vaccenic acid levels and piezophilic growth of the deep-sea bacterium, Photobacterium profundum strain SS9" J. Bacteriol. 182, 1264-1271.
• 2002. "Structure and regulation of the omega-3 polyunsaturated fatty acid synthase genes from the deep-sea bacterium Photobacterium profundum strain SS9" Microbiology 148, 1903-1914.
References
[1] Yuichi Nogi, Noriaki Masui and Chiaki Kato. 1998. “Photobacterium profundum sp. nov., a new, moderately barophilic bacterial species isolated from a deep-sea sediment.” Extremophiles, 2: 1-7.
[2] Vezzi, A., Campanaro, M., D’Angelo, F., Simonato, F., Vitulo, N., Lauro, F.M., Cestaro, A., Malacrida, G., Simionati, B., Cannata, N., Romualdi, C., Bartlett, D.H., Valle, G. “Life at Depth: Photobacterium profundum Genome Sequence and Expression Analysis.” Science. 2005. Volume 307. p. 1459-1461.
[3] Campanaro, M., Vezzi, A., Vitulo, N., Lauro, F.M., D’Angelo, F., Simonato, F., Cestaro, A., Malacrida, G., Bertoloni, G., Valle, G., Bartlett, D.H. “Laterally transferred elements and high pressure adaptation in Photobacterium profundum strains.” BMC Genomics. 2005. p. 1-15.
[4] The Agouron Institute. “Microbial Oceanography: Challenge and Opportunities.” http://www.agi.org/pdf/MicrobialOceanography.pdf
[5] Xie, Cheng-Hui and Yokota, Akira. “Transfer of Hyphomicrobium indicum to the genus Photobacterium as Photobacerium indicum comb. nov.” International Journal of Systematic Evolutionary Microbiology. 2004, 54, 2113-2116.
[6] “ScienceDaily: Advancements in Genomics Foster Deep Sea Discoveries.” April 18, 2005. http://www.sciencedaily.com/releases/2005/04/050411210332.htm
[7] Allen, Eric E., Bartlett, Douglas H. “Structure and regulation of the omega-3 polyunsaturated fatty acid synthase genes from the deep-sea bacterium Photobacterium profundum strain SS9.” Microbiology (2002), 148, 1903-1913.
[8] Bidle, Kelly A. and Barlett, Douglas H. “RNA Arbitrarily Primed PCR Survey of Genes Regulated by ToxR in the Deep-Sea Bacterium Photobacterium profundum Strain SS9.” Journal of Bacteriology (2001), 183(5) 1688-1693.
Edited by Karen Lai, student of Rachel Larsen at UCSD.
KMG