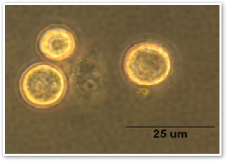
Prochloron didemni
A Microbial Biorealm page on the genus Prochloron didemni
Classification
Higher order taxa
Cellular Organisms; Bacteria; Cyanobacteria; Prochlorales; Prochloraceae; Prochloron
Species
|
NCBI: Taxonomy |
Prochloron Didemni
Description and significance
Prochloron Didemni is a unicellular cyanobacteria that is photosynthetic and oxygenic. It is an obligate symbiont that lives in the ascidian Lissoclinum patella and can be found in marine environments.
The study of Prochloron didemni and its genome can provide us with valuable information. The genome of P. didemni is currently incomplete. However, when completed, P. didemni's genome can be used for analysis to elucidate other genes responsible for symbiosis between bacteria and chordates, such as humans. Also, as a oxygenic, photosynthetic cyanobacteria, P. didemni is a significant player in global carbon fixation. The study of its metabolic pathways and unique photosynthetic processes would yield information beneficial to biotechnology.
R.A. Lewin first discovered Prochloron didemni in 1975, but identified it as Synechocystis didemni. In 1977, the presence of chlorophyll a/b and the lack of phycobilins caused it to be transfered to a new genus, Prochloron, and named Prochloron didemni.(4)
Genome structure
The complete genome for Prochloron didemni has not been sequenced but is estimated to be aproximately 5 million base pairs long with a molecular weight of 3.59x10^9 daltons.(8,10) This estimate was reached by assembly of 82,337 shotgun reads and Glimmer, a gene-finding software.(10) Denaturation kinetics revealed a Tm of 70.6°C, indicating a GC composition of 40.8 mol%.(8)
P. didemni possesses homologs to genes gap2 and gap3, glyceraldehyde-3-phosphate dehydrogenase genes, from Anabaena. These genes are needed to produce GADPH, which is used in the Calvin cycle to metabolize carbon.(6) Genes patA-patG have also been identified, which code for the machinery necessary for patellamide biosynthesis.(1)
Cell structure and metabolism
Prochloron didemni are photosynthetic cyanobacteria with a twist. They carry out photosynthesis similar to Eukaryotic plants, with two reaction centers in noncyclic electron flow.(7) However, unlike cyanobacteria, P. didemni have chlorophyll a and b and lack phycobiliprotiens.(2) Photosynthesis takes place in the thylakoids, with the thylakoids concentrated toward the periphery of the cell.(7)
The cell wall of Prochloron didemni consists of peptidoglycan and muramic acid of the A1 γ type. Approximately 40% cross-linkage exists in P. didemni's cell wall with a network of five to eight layers and a thickness of 12nm.(9)
Ecology
Prochloron didemni is an obligate symbiont that lives in the ascidian Lissoclinum patella. L. patella along with P. didemni can be found in tropical waters, such as the reefs in Hawaii and the Great Barrier Reef in Australia.(2,7)
Exchange of nutrients is the main reason for the symbiotic relationship between L. patella and P. didemni. Photosynthesis by P. didemni provides 60-100% of the organic carbon needed by L. patella. There is also evidence that suggests there is nitrogen cycling between the host and symbiont as well as nitrogen fixation. In exchange for the organic carbons and nitrogen cycling, L. patella provides optimal growth conditions by moving to areas with proper sunlight and a tunic covering.(1)
Patellamides, a product of P.didemni, also play in role in the symbiosis. The cytotoxic characteristic of patellamides may be used as feeding deterrents and patellamides' ability to chelate metals, such as copper and zinc, may be used in detoxification.(1)
Application to Biotechnology
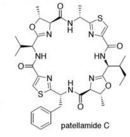
One of the major problems with treating cancer is multi drug resistance. A recent study on Prochloron didemni revealed that it produces Patellamides A and C, which can be used to reverse multi drug resistance in cancer cells.(1)
Patellamides are psuedosymmetrical, cyclic dimers. Each substructure of patellamide contains the specific sequence thiazole-nonpolar amino acid-oxazoline-nonpolar amino acid. Study show that patellamides A and C are ribosomally synthesized and encoded on a single ORF. Genes patA - patG are responsible for the biosynthesis of patellamides. It is currently known that patA codes for a Subtilisin-like protease, patD for an adenylating enzyme-hydrolase hybrid, patE the precursor peptide, and patG a oxidoreductase-protease hybrid. Functions of patB,C,& F are still unknown.(1)
Current Research
Anti-Cancer Compound
A promising study showed that patellamides, a product of Prochloron didemni that can reverse multi drug resistance, can be synthesized by transfer of a biosynthetic pathway. Because P. didemni is an obligate symbiont, it could not be cultivated. To bypass this problem, the genes responsible for patellamide biosynthesis were identified, cloned, and expressed in E.Coli.(1)
Searching for a nonribosomal peptide synthetase (NRPS) adenylation domain yielded a single gene, prnA. Further analysi revealed that this was the wrong domain architecture for patellamide biosynthesis. This pointed to ribosomal synthesis of patellamides. A TBLASTN search lead to patE, the gene for the patellamide precursor peptide. Surrounding patE were other genes necessary for patellamide biosynthesis, ranging form patA tot patG. patA codes for a protease, patD a adenylating enzyme-hydrolase hybrid, and patG a oxidoreductase-protease hybrid. The functions of patB, patC, and patF are still unknown.
Further study of P. didemni's genome will continue to reveal genes involved in patellamide production as well as symbiotic mechanisms and pathways.
Isolation of Photosystem II Supercomplex
It was recently shown that Prochlorococcus marinus SS120 had a 18-Pcb subunit, light-harvesting antenna ring around photosystem I. Investigation of Prochloron didemni revealed taht it had a similar supercomplex around photosystem II. The supercomplex is a photosystem II core dimer with five Pcb proteins on each side. Assuming that photosystem II core monomer binds 35 chlorophylls and each Pcb protein, like CP43, binds 13 chlorophylls, the supercomplex would increase the light-harvesting capacity of by ~200%. This increase in absorption power compensates for the lack of phycobiliproteins.(2)
Isolation of Thylakoids
References
(3) Lockhart, P., Beanland, T., Howe, C., and Larkum, A. "Sequence of Prochloron didemni atpBE and the inference of chloroplast origins". Proc. Natl. Acad. Sci. USA: Evolution. April, 1992. Volume 89. p. 2742-2746.]
(5) Christen, G., Stevens, G., Lukins, P., Renger, G., and Larkum, A. "Isolation and characterisation of oxygen evolving thylakoids from the marine prokaryote Prochloron didemni". FEBS. March, 1999. Letters 449 p. 264-268.
(6) Figge, R., Schubert, M., Brinkmann, H., and Cerff, R. "Glyceraldehyde-3-Phosphate Dehydrogenase Gene Diversity in Eubacteria and Eukaryotes: Evidence for Intra- and Inter-Kingdom Gene Transfer". Mol. Biol. Evol.. 1999. p. 429-440.
(7) Giddings Jr., T., Withers, N., and Staehelin, L. A. "Supramolecular structure of stacked and unstacked regions of the photosynthetic membranes of Prochloron sp., a prokaryote". Proc. Natl. Acad. Sci. USA: Botany. January, 1980. Volume 77. No. 1 p. 352-356.
(8) Herdman, M. "Deoxyribonucleic Acid Base Composition and Genome Size of Prochloron". Arch Microbiol. 1981. Volume 129. p. 314-316.
(9) Jurgens, U., Burger-Wiersma, T. "Peptidoglycan=Polysaccharide Complex in the Cell Wall of the Filamentous Prochlorophyte Prochlorothrix hollandica". Journal of Bacteriology. January, 1989. Volume 171. No. 1 p. 498-502.
(10) Delcher, A., Bratke, K., Powers, E., and Salzberg, S. "Identifying bacterial genes and endosymbiont DNA with Glimmer" Bioinformatics. January, 2007. Volume 23. No. 6 p. 673-679.
Edited by Edward Lee Eddie Lee, a student of Rachel Larsen Rachel Larsen