Proteobacteria
A Microbial Biorealm page on the phylum Proteobacteria
Cell Structure
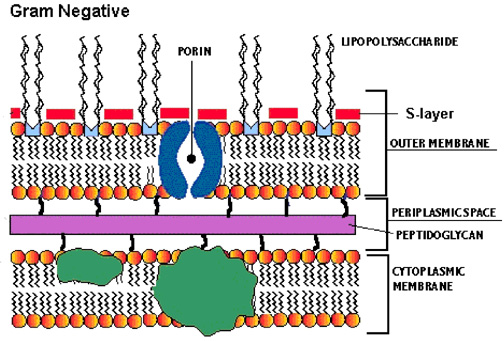
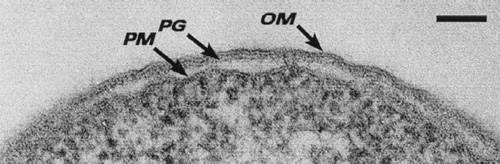
The structural link between gram-negative bacteria is the presence of an outer membrane (OM), a peptidoglycan layer, and a periplasm, which all together constitute the gram-negative envelope. The outer membrane contains two kinds of lipids, lipopolosaccharide (LPS) and phospholipids, as well as some unique proteins. The phospholipid composition of the OM is similar to that of cytoplasmic membrane except it has a slight augmentation in phophatidylethanolamine.
The LPS is composed of three parts, the proximal, hydrophobic lipid A region, the distal, hyrdophilic O-antigen polysaccharide region, and the core oligosaccharide that connects the two. The lipid A region and proximal part of the core both contain many charged groups, most anionic. The lipid A region is thought to be essential for OM assembly while the proximal end of the core is responsible for maintaining the barrier property of the OM. The proximal part of the core keeps out hydrophobic compounds including antibodies, mutagens, and detergents. The O-antigen region appears to help virulence, escaping phagocytosis, and sticks out into the surrounding medium.
Murein lipoprotein is a small protein at 7200 Da that exists in great numbers with 7x105 copies per cell. Three fatty acid groups penetrate the inner leaflet of the OM while the rest is hydrophilic and is most likely located in the periplasm. About one-third of the murein lipoprotein molecules are covalently bound to the peptidoglycan layer, thus fixing the OM to the underlying peptidoglycan and increasing stability of the envelope.
Porins are relatively non-specific pores/channels that allow the rapid passage of small hydrophilic molecues across the OM. The porins have an antiparallel B-sheet shape and do not contain any long stretches of hydrophobic amino acids, resulting in it transversing the membrane 16 times. The external loops to the sheets contain many charged amino acid residues that are thought to interact with the negatively charged LPS through divalent cation bridges. Porins can be found in a trimer with the center being hydrophobic. The channels have a wide entrance, wide exit, and a short central constriction that allows near maximum diffusion rates by minimizing friction and still serves as a barrier to larger solutes.
References
Beveridge, Terry J. 1999. Structures of Gram-Negative Cell Walls and Their Derived Membrane Vesicles. Journal of Bacteriology 181 16: 4725-4733.
F.C. Neidhardt et al. (ed.). 1996. Escherichia coli and Salmonella: cellular and molecular biology, 2nd ed., vol. 1. American Society for Microbiology, Washington, D.C.