Pseudoalteromonas atlantica: Difference between revisions
| Line 19: | Line 19: | ||
Describe the appearance, habitat, etc. of the organism, and why it is important enough to have its genome sequenced. Describe how and where it was isolated. | Describe the appearance, habitat, etc. of the organism, and why it is important enough to have its genome sequenced. Describe how and where it was isolated. | ||
Include a picture or two (with sources) if you can find them. | Include a picture or two (with sources) if you can find them. | ||
[[Image:Pseat.jpg|right]] | |||
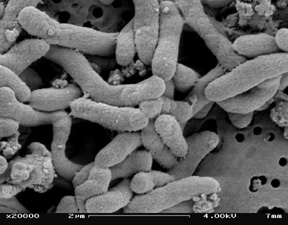
Pseudoalteromonas atlantica was first isolated from a marine algae in the Antarctic coastal marine environment. It is a rod-shaped, motile, gram-negative bacteria that is also found in the ocean world wide. The bacteria is motile through the means of a single polar flagellum, which helps it move back and forth between solid surfaces and the open ocean in search of food sources. The bacterial cells usually occur singly or in pairs. It obtains energy through chemoorganotrophic mechanisms. It is also biofilm-forming bacteria, which becomes important for bioremediation. It usually is found with marine eukaryotic hosts, such as crabs and seaweed. The genome of Pseudoalteromonas atlantica was sequenced due to its production of acidic extracellular polysaccharide during biofilm formation that shows potential in element recycling, detoxification, and materials production. | Pseudoalteromonas atlantica was first isolated from a marine algae in the Antarctic coastal marine environment. It is a rod-shaped, motile, gram-negative bacteria that is also found in the ocean world wide. The bacteria is motile through the means of a single polar flagellum, which helps it move back and forth between solid surfaces and the open ocean in search of food sources. The bacterial cells usually occur singly or in pairs. It obtains energy through chemoorganotrophic mechanisms. It is also biofilm-forming bacteria, which becomes important for bioremediation. It usually is found with marine eukaryotic hosts, such as crabs and seaweed. The genome of Pseudoalteromonas atlantica was sequenced due to its production of acidic extracellular polysaccharide during biofilm formation that shows potential in element recycling, detoxification, and materials production. | ||
==Genome structure== | ==Genome structure== | ||
Revision as of 05:31, 3 June 2007
Pseuderomonas Atlantica A Microbial Biorealm page on the genus Pseudoalteromonas atlantica
Classification
Higher order taxa
Bacteria; Proteobacteria; Gammaproteobacteria; Alteromonadales; Pseudoalteromonadaceae; Pseudoalteromonas Domain; Phylum; Class; Order; family [Others may be used. Use NCBI link to find]
Species
|
Pseudoalteromonas atlantica T6c Description and significanceDescribe the appearance, habitat, etc. of the organism, and why it is important enough to have its genome sequenced. Describe how and where it was isolated. Include a picture or two (with sources) if you can find them. Pseudoalteromonas atlantica was first isolated from a marine algae in the Antarctic coastal marine environment. It is a rod-shaped, motile, gram-negative bacteria that is also found in the ocean world wide. The bacteria is motile through the means of a single polar flagellum, which helps it move back and forth between solid surfaces and the open ocean in search of food sources. The bacterial cells usually occur singly or in pairs. It obtains energy through chemoorganotrophic mechanisms. It is also biofilm-forming bacteria, which becomes important for bioremediation. It usually is found with marine eukaryotic hosts, such as crabs and seaweed. The genome of Pseudoalteromonas atlantica was sequenced due to its production of acidic extracellular polysaccharide during biofilm formation that shows potential in element recycling, detoxification, and materials production. Genome structureDescribe the size and content of the genome. How many chromosomes? Circular or linear? Other interesting features? What is known about its sequence? Does it have any plasmids? Are they important to the organism's lifestyle? The size of its genome is 5.187007 megabases, and its GC content is 45%. Cell structure and metabolismDescribe any interesting features and/or cell structures; how it gains energy; what important molecules it produces. Some important molecules it produce include acidic extracellular polysaccharide(EPS), enzymes that hydrolyze agar, alginate, and carrageenan(such as agarase), signalling molecules, such as homoserine lactones, and proteases. It produces EPS in large amounts in order to concentrate nutrients and provide substrates within its biofilm for other marine microorganisms. In order to do this, it gains energy by colonizing solid surfaces quickly, and using its secreted enzymes to process and take up substrates. EcologyPseudoalteromonas atlantica's production of acidic extracellular polysaccharide in its biofim is particularly useful in bioremediation. The biofilm can be important in controlling toxic metal concentrations in marine environments because it can absorb 20-40% of trace metal lead. The bacteria has an unusual ability to regulate its EPS production. It has a DNA recombination system that involves reversible insertion of a mobile element, called IS 492. The insertion of IS 492 at an E{S site makes variable expressions of EPS possible. This system is responsive to changes in environmental conditions, therefore the bacteria can regulate its production of EPS in terms of the present environmental condition. PathologyHow does this organism cause disease? Human, animal, plant hosts? Virulence factors, as well as patient symptoms. It does not cause any disease. Current ResearchEnter summaries of the most recent research here--at least three required ReferencesEdited by student of Rachel Larsen and Kit Pogliano |