Rickettsia japonica: Difference between revisions
No edit summary |
|||
| (14 intermediate revisions by 2 users not shown) | |||
| Line 12: | Line 12: | ||
*Species: R.japonica | *Species: R.japonica | ||
[[File: | |||
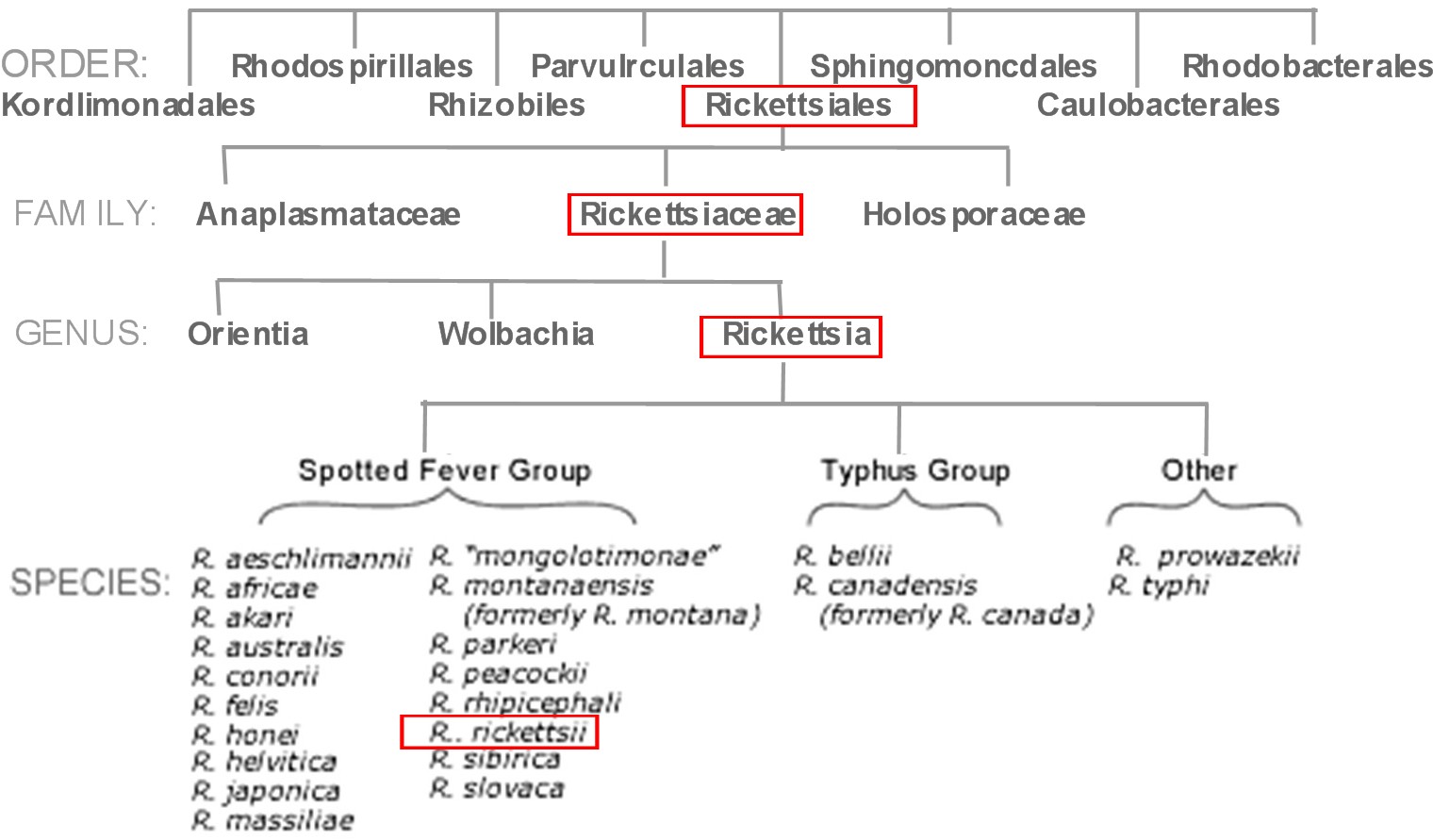
[[File:TreeOG.jpg|Figure 1: This unrooted phylogeny tree of the Rickettsia genus shows that R.japonica is closely related to Rickettsia rickettsii due to their genetic coding. The majority of the spotted fever group need two hosts and share the same characteristics which correlate with the difference in their genetic code (1). | |||
]] | ]] | ||
| Line 31: | Line 32: | ||
=7. Pathology= | =7. Pathology= | ||
''R. japonica'' causes Japanese spotted fever, in central Japan and Korea (3). Japanese spotted fever causes high fever, shaking chills, headache, and skin eruptions that rapidly spread throughout the body(18). Due to the variety of tick species the geographical region affected by the disease has expanding (4). Japanese spotted fever may result in death and has infected more people annually since first discovered in 1984(6). Because of ''R. japonica''’s pathogenic nature, it’s genome is reduced. This is demonstrated through the growth in host cells (8). A lineage of ''R. japonica'' contains non-synonymous | ''R. japonica'' causes [https://en.wikipedia.org/wiki/Japanese_spotted_fever Japanese spotted fever], in central Japan and Korea (3). Japanese spotted fever causes high fever, shaking chills, headache, and skin eruptions that rapidly spread throughout the body(18). Due to the variety of tick species the geographical region affected by the disease has expanding (4). Japanese spotted fever may result in death and has infected more people annually since first discovered in 1984(6). Because of ''R. japonica''’s pathogenic nature, it’s genome is reduced. This is demonstrated through the growth in host cells (8). A lineage of ''R. japonica'' contains non-synonymous [https://en.wikipedia.org/wiki/Single-nucleotide_polymorphism single nucleotide polymorphisms] with genes for rOmpB proteins (4). This protein aids in attaching and invading host cells, while increasing disease severity (12). A target open reading frame only existing in ''R. japonica'' bacteria can be used for rapid diagnosis and treatment of ''R. japonica'' infection(6). The disease can be treated with proper antibiotics in its early stages (6). | ||
=8. Current Research= | =8. Current Research= | ||
Latest revision as of 14:52, 10 December 2018
1. Classification
a. Higher order taxa
- Domain: Bacteria
- Kingdom: Prokaryote
- Phylum: Proteobacteria
- Class: Alphaproteobacteria
- Order: Rickettsiales
- Family: Rickettsiaceae
- Genus: Rickettsia
- Species: R.japonica
2. Description and significance
Rickettsia japonica belongs to the genus of Rickettsia, a small Gram-negative genus which are split into spotted fever and typhus groups (2). R.japonica, discovered in 1984, is different from other Rickettsia in that the species has distinct surface polypeptides and antibodies that only react tow R. japonica (3). This strain causes Japanese spotted fever, is highly pathogenic even fatal, and transferable through ticks, lice, and mites (4). R. japonica is an obligate intracellular pathogen with a small genome. As a result, the genomic diversity of this bacteria is low (4). Because of its genome size, many strains of R. japonica have been isolated and analyzed revealing few polymorphisms among the sequences.
R. japonica is pervasive in the Asian and Oceanic geographical regions (5). The bacteria is spread through animals traveling by air and ground specifically ones that are in higher abundance in warm and humid climates (2). Due to the rising influence of global climate change, it is possible that the transmission of this bacteria could continue to spread and evolve (6). There is also the possibility that household pets may become the most abundant victim. Fortunately, a treatment for early stages of Japanese spotted fever consisted of antibiotics, which proved successful and efficient (7).
3. Genome structure
R. japonica has a small genome, 1.1-1.6 MB, due to it being an obligate intracellular pathogen. Because genomic diversity is low, polymorphisms are not abundant across the other strains of Rickettsia (4). Mobile genetic elements like plasmids are present across the different strains of the species which is indicative of conjugation and horizontal gene transfer. The few amount of single nucleotide polymorphisms among the strains of R. japonica are representative of its low genomic diversity. R. japonica have one circular chromosome with a size of 1.3 MBP, one set of rRNA genes, and 1,186 regions that can express proteins (8). The guanine and cytosine content of the genome is roughly 32.7% and 31.8% (8). R. japonica has a specific open-reading frame , a 216 base pair specific set of genes, within the genome across the bacteria that assists with diagnosis of the disease through DNA sequencing. The nucleotide sequence within this open-reading frame was identical across all strains of R. japonica (6).
4. Cell structure
R. japonica is an obligate intracellular bacterium (4). It is a Gram negative rod that has a periplasmic space between the inner and outer membrane with lipopolysaccharides concentrated in the outer membrane. It is non-motile, does not form spores, and is considered pleomorphic meaning it is able to change its shape and size in adapting to a certain environment (9).
This bacterium has an outer layer composed of a unknown slime substance, producing a three-layered cell wall and are 0.4-0.5 x 0.8-1.5 micrometers in diameter (3). The inner membrane is thick while the outer membrane is thin (10). R. japonica has a plaque morphology when allowed to grow or cultured in eukaryotic cells. The cell wall is composed of peptidoglycan and is often localized to the nucleus, the endothelial and smooth muscle cytosol. It is classified as a zoonese, a type of disease that comes from animals pathogens which are transmissible to humans through mechanical or biological vectors (11). Replication methods are currently unknown, but this bacterium has major outer membrane proteins; rOmpB and rOmpA, which are implicated for the adhesion and protective immunity. rOmpB is the most common outer membrane protein which has functions of the S layer as well(12). rOmpA is involved in actin activity within the cell allowing the bacterium to invade and multiply within a host cell (12).
5. Metabolic processes
R. japonica requires a eukaryotic host to proliferate. Its optimal growth range lies between 32°C - 35 °C (13). R. japonica requires the shortest time of Rickettsia strains to form plaques which begins the infection cycle of 2-3 days. The bacterium will enter the host cell and lyse it to release progeny (13). Glutamic acid and glutamine are important organic molecules that assist in host cell penetration. Glutamate is oxidized in order to create ATP to assist in survival of the cell (13). Coupling energy creation with penetration allows the bacterial cell to integrate into the host cell (13). Pyruvates are also important organic molecules that enhance the pathogenicity of the cell. While the mechanism of this is unclear, it is known that pyruvate is essential for the respiration of the bacterium (13). In addition, incubating the cell with pyruvate is sufficient to restore the loss of biological activity (13). R. japonica multiply through transverse binary fission but low pH is unfavorable to cell survival(14). This bacterium is known to have transporter systems, which take host metabolites like NAD+ to power the TCA cycle and to acquire vitamins and cofactors (15).
6. Ecology
R. japonica has a life cycle closely dependent on Ixodidae and Argasidae ticks, which they use as transportation and hosts (16). The ticks and R.japonica have either a beneficial or harmful relationship. The Rickettsia infect the tick early in its life cycle and the two species grow together. The bacteria can lay inactive in the tick until it bites a human host. This contributes to the low genome diversity(4). However, little is known about the full extent of the relationship between tick and bacteria; due to the Rickettsia genus’ fatal relationship with other insects, there is the need for further research (4). Approximate replication time of R. japonica on its own is 6 hours, however when dependent on the host bacterial growth is slowed(4). R. japonica can switch between tick species frequently without altering the bacterial or host DNA. The species of tick with which the taxa interacts with determines the geographical distribution(17). For example, Dermacentor taiwanensis and Haemaphysalis flava ticks that prefer to live in environments like Japan are the preferred vectors for R. japonica(18). In addition to ticks, R. japonica antibodies are in bats determining that R. japonica can travel between species of hosts and to a larger geographical radius(5).
7. Pathology
R. japonica causes Japanese spotted fever, in central Japan and Korea (3). Japanese spotted fever causes high fever, shaking chills, headache, and skin eruptions that rapidly spread throughout the body(18). Due to the variety of tick species the geographical region affected by the disease has expanding (4). Japanese spotted fever may result in death and has infected more people annually since first discovered in 1984(6). Because of R. japonica’s pathogenic nature, it’s genome is reduced. This is demonstrated through the growth in host cells (8). A lineage of R. japonica contains non-synonymous single nucleotide polymorphisms with genes for rOmpB proteins (4). This protein aids in attaching and invading host cells, while increasing disease severity (12). A target open reading frame only existing in R. japonica bacteria can be used for rapid diagnosis and treatment of R. japonica infection(6). The disease can be treated with proper antibiotics in its early stages (6).
8. Current Research
Over the past decade, research has determined aspects of genetic diversity, growth, evolution, and the study of hosts (vectors) that revealed the pathogenicity of R. japonica. The bacteria’s replication and long generation time were found to dependent on and parallel to the life cycle of the host. Studies have shown potential for new vectors such as household dogs, which pose a bigger threat for humans to interact with R. japonica(18). Although interacting with potential vectors seemed controllable in the past, recent studies have exposed the spread of R. japonica by a wild flying carrier, the Australian Bat (5). The observed spread overseas of the original Japanese-based strain of R. japonica has incentivized further research in hopes to contain the fatal pathogen. There is currently a need for studying the future locations for this strain to arrive at a definitive radius of new infections. Then scientists will be able to determine the prevalence of R. japonica beyond coastal Asia and Oceania regions of the world in the future.
9. References
- Gibson, C. Rickettsia rickettsii. (2007). University of Wisconsin-La Crosse. Bioweb.uwlax.edu. Retrieved from http://bioweb.uwlax.edu/bio203/s2008/gibson_chel/Classification.htm
- Walker, D.H. (1996) Rickettsiae. Medical Microbiology, 4th Edition, Chapter 38. Retrieved from https://www.ncbi.nlm.nih.gov/books/NBK7624/
- Uchida, T. et al. (1992). Rickettsia Japonica Sp. Nov., The Etiological Agent Of Spotted Fever Group Rickettsiosis In Japan. International Journal of Systematic Bacteriology, 42.2, 303-305. Retrieved from http://ijs.microbiologyresearch.org/content/journal/ijsem/10.1099/00207713-42-2-303.
- Akter, A. et al. (2017). Extremely low genomic diversity of Rickettsia japonica distributed in Japan. Genome Biology and Evolution, 9(1). 124–133. Retrieved from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5381555/
- Izzard, L., et al. (2018). Isolation of a divergent strain of Rickettsia japonica from Dew's Australian bat Argasid ticks (Argas (Carios) dewae) in Victoria, Australia. Ticks And Tick-Borne Diseases, 9(6) 1484-1488. Retrieved Sept. 17, 2018, from https://www.sciencedirect.com/science/article/pii/S1877959X18300694
- Climate Change: Global Temperature | NOAA Climate.gov. (2018). Climate.gov. Retrieved from https://www.climate.gov/newsfeatures/understanding-climate/climate-change-global-temperature
- Hanaoka, N., et al (2009). Diagnostic assay for Rickettsia japonica. Emerging Infectious Diseases 15(12): 1994–1997. Retrieved from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3044520/
- Dong, X., et al (2012). Genomic Analysis of Rickettsia japonica Strain YHT. J Bacteriol 194(24): 6692. Retrieved from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3510609/
- Rickettsia. (2018). En.wikipedia.org. Retrieved from https://en.wikipedia.org/wiki/Rickettsia
- Choi, Y., et al (2005). Spotted Fever Group and Typhus Group Rickettsioses in Humans, South Korea. Emerging Infectious Diseases, 11(2): 237-244. Retrieved from https://wwwnc.cdc.gov/eid/article/11/2/04-0603_article
- G. Venkatesan., et al. (2010). Viral Zoonosis: A Comprehensive Review. Asian Journal of Animal and Veterinary Advances, 5, 77-92. Retrieved from https://scialert.net/abstract/?doi=ajava.2010.77.92
- Uchiyama T., et al. (2006). The major outer membrane protein rOmpB of spotted fever group rickettsiae functions in the rickettsial adherence to and invasion of Vero cells. Microbes Infect, 8(3), 801-9. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/16500128
- Weiss, E. (1973). Growth and physiology of rickettsiae. Bacteriological Reviews, 37(3), 259. Retrieved from https://www.ncbi.nlm.nih.gov/pmc/articles/
- Gray, Lawrence R, et al. (2013). Regulation of pyruvate metabolism and human disease. Cellular and molecular life sciences, 71(14), 2577-604.
- Driscoll, T, et al. (2017). Wholly Rickettsia! Reconstructed Metabolic Profile of the Quintessential Bacterial Parasite of Eukaryotic Cells. Mbio, 8(5), 00859-17
- Burgdorfer W, et al. (1975). Mechanisms of transovarial infection of spotted fever Rickettsiae in ticks. Annals of the New York academy of Sciences, 266, 61–72. Retrieved from https://nyaspubs.onlinelibrary.wiley.com/doi/abs/10.1111/j.1749-6632.1975.tb35088.
- Paddock CD, et al. (2014). Phylogeography of Rickettsia rickettsii genotypes associated with fatal Rocky Mountain spotted fever. Am J Trop Med Hyg, 91, 589–597. Retrieved from http://www.ajtmh.org/content/journals/10.4269/ajtmh.14-0146
- Inokuma, H., et al. (2011). Evaluation of Rickettsia japonica pathogenesis and reservoir potential in dogs by experimental inoculation and epidemiologic Survey. Clinical and Vaccine Immunology, 18(1), 161–166. Retrieved Sept. 17, 2018, from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3019780/
- Mahara, F. (1997). Japanese Spotted Fever: Report of 31 Cases and Review of the Literature. Emerging Infectious Diseases, 3(2), 105-111. Retrieved from https://wwwnc.cdc.gov/eid/article/3/2/97-0203_article
- Ogawa A, et al. (2018). What is Japanese red spot fever. National Institute of Infectious Disease. Retrieved from https://www.niid.go.jp/niid/ja/biorisk-guidance/392-encyclopedia/448-jsf-intro.html