Rubrobacter radiotolerans
Classification
Higher order taxa
Domain: Bacteria
Phylum: Actinobacteria
Class: Actinobacteria
Subclass: Rubrobacteridae
Order: Rubrobacterales
Family: Rubrobacteraceae
Genus: Rubrobacter
Species
Rubrobacter radiotolerans RSPS-4 (Type Strain RRD 42256)
Description and significance
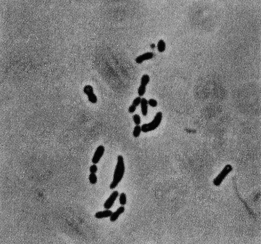
The strain RSPS-4 of Rubrobacter radiotolerans was isolated from hot spring runoff in São Pedro do Sul, Portugal (Egas et al., 2014). The spring was at the surface and had a pH of 8.9 and a temperature of 50°C. From this and isolation it seems to have optimal conditions in semi-salty water with temperatures of 45°C (hot springs). The strain was warranted to the the Spanish Type Culture Collection (CETC) under the code CETC 8386 (Egas et al., 2014). Rubrobacter radiotolerans (at first, Arthrobacter radiotolerans) was originally found in a water sample taken from a hot spring in Tottori Prefecture, Japan. It was discovered after putting the sample through gamma-irradiation (Strain P-1) (Yoshinaka et al., 1973). It is a gram-positive bacterium high in G+C content that is non-sporulating and non-motile. In earlier stages the bacteria seems to be shaped mostly as pleomorphic rods and in late stages seems to be mostly coccoid-shaped or shaped as short rods (Yoshinaka et al., 1973). R. radiotolerans are smooth, opaque, and contain pinkish to red pigmented carotenoids. The major pigmented components are bacterioruberin and monoanhydrobacterioruberin which have 13 conjugated double bonds that are characteristic of halophilic bacteria. Bacterioruberin has four tertiary OH groups and monoanhydrobacterioruberin has three tertiary OH groups (Saito et al., 1994). A third major component, deinoxanthin, has also been identified in contributing to their reddish pigment (Asgarani et al., 2000).
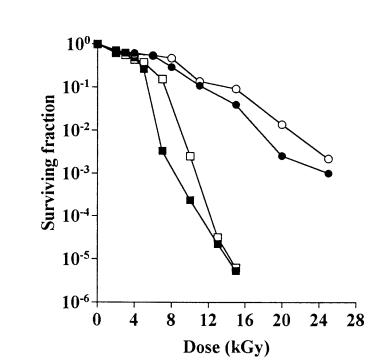
R.radiotolerans are extremely resistant to UV, thermal, and gamma radiation, even more so than its relative Rubrobacter xylanophilus (Ferreira et al., 1999). Gamma radiation is typically restricted to locations contaminated with nuclear waste as high doses are not naturally found in the biosphere. One hypothesis for such a characteristic is that it evolved in response to environmental challenges such as desiccation or reactive oxidative stress. In addition, such radiation resistance is often found in organisms that do not produce spores, such as R.radiotolerans (Egas et al., 2014).
The ability to recover from such high doses of radiation seems to result from a combination of several mechanisms. Within its carotenoids, the components deinoxanthin and bacterioruberin are attributed to radiation resistance and act as an effective antioxidant in vitro. Additionally, bacterioruberin has also been attributed to resistance to hydrogen peroxide. A DNA repair enzyme, known as R-endonuclease, aids in DNA repair activity by recognizing radiation-induced DNA lesions. Thymine glycol and ring fragmentation products such as urea residues, which blocks DNA replication and is premutagenic, and abasic sites, which can be lethal and mutagenic, are cleaved by this repair enzyme in order to prevent their cytotoxic and mutagenic effects on cells (Asgarani et al., 2000). A third mechanism involves the manganese superoxide dismutase (SOD), an enzyme that forms homo-tetramerization of 24,000 Da-monomer while maintaining enzymatic activity against potassium cyanide and hydrogen peroxide.
16S Ribosomal RNA Gene Information
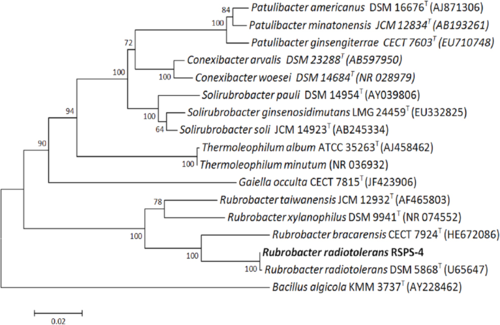
The strains RSPS-4 and P-1 (Type strain DSM 5868) have extremely similar 16S rRNA sequences, with only two components differing (Egas, et. al, 2014).
The sequencing of the 16S ribosomal RNA data of DSM 5858 can be found at:
NCBI database
(ncbi database) https://www.ncbi.nlm.nih.gov/nuccore/NR_119230.1
Genome Structure
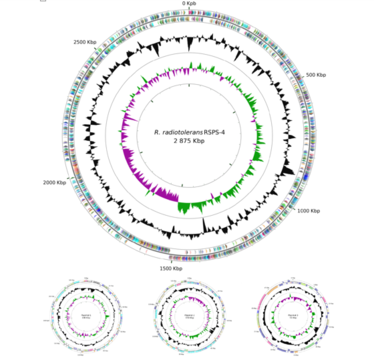
The total complete genome size is 3,267,233 bp. The genome consists of a one circular chromosome of 2,875,491 bp with a G+C content of 66.91% and three circular plasmids of 190,889 bp, 149,806 bp, 51,047 bp, holding 3,214 predicted protein coding genes, 46 tRNA genes, and one rRNA operon (strain RSPS 4) (Egas et al., 2014).
The sequencing was conducted in Penzberg, Germany at Roche Diagnostics GmbH (Egas et al., 2014).
Cell structure and metabolism
R. radiotolerans are a smooth, opaque bacterium shaped as pleomorphic rods and, later in life, is coccoid-shaped or shaped as short rods (Yoshinaka et al., 1973). Their pigmented carotenoids are pinkish to red colored. The major pigmented components are bacterioruberin and monanhydrobacterioruberin, which contain 13 conjugated double bonds that are a characteristic of halophilic bacteria. There are four tertiary OH groups in bacterioruberin and three in monanhydrobacterioruberin (Saito et al., 1994). Deinoxanthin, a third pigmented component, has also been identified (Asgarani et al., 2000).
Rubrobacter radiotolerans have multiple mechanisms that result in it being radiotolerant. Deinoxanthin and bacterioruberin are linked to radiation resistance as antioxidants and, additionally, bacterioruberin has also been linked to hydrogen peroxide resistance (Asgarani et al., 2000). R-endonuclease, a DNA repair enzyme, helps by recognizing radiation-induced lesions and manganese supreroxide dismutase maintains enzymatic activity against potassium cyanide and hydrogen peroxide (Asgarani et al., 2000).
You can find a table of different metabolic pathways, including unique gene counts and which genes that belong to certain pathways at:
(Wattam et al., 2014).
Ecology and Pathogenesis
Rubrobacter radiotolerans are extremophiles with some interesting characteristics. It is thermotolerant and therefore can live in locations with temperatures ranging from 30-55℃ in areas like hot springs. Due to its radiotolerance to UV-radiation and gamma radiation, it can reside in places that have been contaminated (Egas et al., 2014).
This bacterium contains no pathogenicity (Egas et al., 2014). The table listed below represents the specialty genes identified in R.radiotolerans strain RSPS-4. Specialty genes refers to the highlighted sets of genes that are of interest to infectious disease researches. Such genes include antibiotic resistance, virulence factors, drug targets, and human homologs. The genetic data collected on R.radiotolerans only identified human homologs, “bacterial proteins that share high sequence similarity with human proteins.” If this species were pathogenic, it would contain virulence factors that would “enable bacteria to establish itself on or within a host organism and enhance its potential to cause disease” (Wattam et al., 2014). You can find the table listing these specialty genes associated with Rubrobacter radiotolerans at https://www.patricbrc.org/portal/portal/patric/SpecialtyGeneList?cType=genome&cId=42256.3&kw=. (Wattam et al., 2014).
Current Research
Egas and colleagues published a paper in regards to the complete genome sequence of Strain RSPS-4 and its analysis (Egas et al., 2014). Terato and colleagues published research about Rubrobacter radiotolerans’ radioresistant characteristics (Terato et al., 2011). Wattam and colleagues are using new technology to research bacteria such as Rubrobacter radiotolerans (Wattam et al., 2014).
References
Asgarani, E., Terato, H., Asagoshi, K., Shahmohammadi, H. R., Ohyama, Y., Saito, T, & Ide, H. (2000). Purification and characterization of a novel DNA repair enzyme from the extremely radioresistant bacterium Rubrobacter radiotolerans. Journal of radiation research, 41(1), 19-34.
Egas, C., Barroso, C., Froufe, H. J. C., Pacheco, J., Albuquerque, and M. S. da Costa (2014). Complete genome sequence of the Radiation-Resistant bacterium Rubrobacter radiotolerans RSPS-4. Standards in Genomic Sciences, 9(3), 1062-1075.
Ferreira, A., Nobre, M., Moore, E. et al. (1999). Characterization and radiation resistance of new isolates of Rubrobacter radiotolerans and Rubrobacter xylanophilus. Extremophiles 3: 235. doi:10.1007/s007920050121
Kausar, Javed, et al. (1997) "16S rRNA gene sequence of Rubrobacter radiotolerans and its phylogenetic alignment with members of the genus Arthrobacter, gram-positive bacteria, and members of the family Deinococcaceae." International Journal of Systematic and Evolutionary Microbiology 47.3: 684- 686.
Saito, T., Terato, H. & Yamamoto, O. (1994). Pigments of Rubrobacter radiotolerans Arch. Microbiol. 162: 414. doi:10.1007/BF00282106
Terato, H., Suzuki, K., Nishioka, N., Okamoto, A., Shimazaki-Tokuyama, Y., Inoue, Y., & Saito, T. (2011). Characterization and radio-resistant function of manganese superoxide dismutase of Rubrobacter radiotolerans. Journal of radiation research, 52(6), 735-742.
Wattam, A.R., D. Abraham, O. Dalay, T.L. Disz, T. Driscoll, J.L. Gabbard, J.J. Gillespie, R. Gough, D. Hix, R. Kenyon, D. Machi, C. Mao, E.K. Nordberg, R. Olson, R. Overbeek, G.D. Pusch, M. Shukla, J. Schulman, R.L. Stevens, D.E. Sullivan, V. Vonstein, A. Warren, R. Will, M.J.C. Wilson, H. Seung Yoo, C. Zhang, Y. Zhang, B.W. Sobral (2014). “PATRIC, the bacterial bioinformatics database and analysis resource.” Nucl Acids Res 42 (D1): D581-D591. doi:10.1093/nar/gkt1099. PMID: 24225323.
Yoshinaka, Taeko, Keiji Yano, and Hikoyuki Yamaguchi (1973). "Isolation of highly radioresistant bacterium, Arthrobacter radiotolerans nov. sp." Agricultural and Biological Chemistry 37.10: 2269-2275.
Author
Amber Parcel & Katie McCullough
Students of Hidetoshi Urakawa at Florida Gulf Coast University