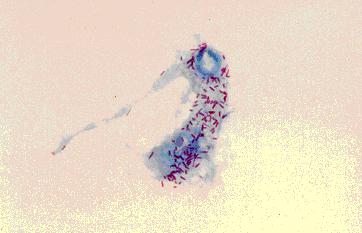
Sodalis glossinidius
A Microbial Biorealm page on the genus Sodalis glossinidius
Classification
Higher order taxa
Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacteriales; Enterobacteriaceae (1)
Species
Sodalis glossinidius (1)
Description and significance
Sodalis glossinidius is a Gram-negative, nonspore-forming, rod-shaped, filamentous bacteria. It has flagella expressed during immature host developmental stages, though it is non-motile.(10) It is one of three endosymbionts for the tsetse fly (Glossina spp.), which are all maternally transmitted to progeny.(6,7) It has a mutualistic relationship with the tsetse fly. This secondary endosymbiont resides inter- and intracellularly in the midgut, fat body and haemolymph of the insect. S. glossinidius was the first true insect endosymbiont to be isolated and cultured, achieved in 1999 using a mosquito (Aedes albopictus) feeder cell culture system and a sample from the haemolymph of Glossinia morsitans morsitans.(7) The type strain M1T was isolated.(7) S. glossinidius can grow intracellularly in Aedes albopictus or axenically in semidefined solid medium containing already enzimatically digested proteins as a nitrogen source. This microaerophilic bacterium grows optimally with 5% oxygen and 95% carbon dioxide at 25 °C. Colony morphology is uniform, with defined edges, and off-white.(7)
The tsetse fly is a vector for Trypanosoma brucei gambiense and Trypanosoma brucei rhodesiense, parasitic protozoa that cause African trypanosomiasis (commonly known as African Sleeping Sickness), an epidemic fatal disease plaguing sub-Saharan Africa.(9) S. glossinidius is important enough to have its genome sequenced because this it is a key player involved in the host-parasite relationship, possibly enabling the creation of tsetse flies that are incapable of transimtting trypanosomes and thus bringing an end to African sleeping sickness (See Ecology for more details).(6) The genome of S. glossinidius also provides an example of an organism that has undergone degenerative adaptations in order to adopt a mutualistic relationship with its host (See Genome structure for more details).(3)
Genome structure
Describe the size and content of the genome. How many chromosomes? Circular or linear? Other interesting features? What is known about its sequence? Does it have any plasmids? Are they important to the organism's lifestyle?
Sodalis glossinidius has one circular chromosome approximately 4 Mb.(10) This micro-organism has a relatively inactive biochemical profile (2,432 protein coding sequences, indicating %51 reduced coding capacity) compared to other organisms in the family Enterobacteriaceae.(3,7,10) this is likely due to its evolutionary path away from free-existence. The genome reduction and function loss undergone by S. glossinidius is less than other intracellular pathogens and obligate symbionts, shown by its ability to be cultured in vitro, but its genome size is still smaller than closely related free-living enterics.(2) Further suggesting that it is in the process of becoming more dependent on the tsetse host cell. Though most organisms of the Family Enterobacteriaceae are able to produce catalase, S. glossinidius is not. This is thought to be why this micro-organism requires a microaerophilic environment.(7) Genes dealing with transcription, translation, regulation, and nucleic acid and amino acid biosynthetic pathways were retained. On the other hand, 972 pseudogenes (mostly caused by single point mutations) are homologs of known proteins that deal defense or for transporting and metabolizing carbohydrates and inorganic ions, reflecting the mutualistic relationship with the host.(2,10)
The genome of S. glossinidius as an A+T content of only 45%, whereas endosymbiont genomes are typically AT-rich. The A+T content of intracellular pathogen R. prowazekii is 71% and 75% for the mutualist Buchnera. The AT-rich genomes of intracellular pathogens and obligates may be due to the loss of function of DNA repair and recombination functions, suggesting that S. glossinidius has retained these genes.(2) The 16S rRNA of S. glossinidius shares high sequence identity with other secondary endosymbionts of insects. It also was found to be almost identical within the species when isolated from different Glossina spp. This suggests recent acquisition by the tsetse as an enteric and possible horizontal symbiont transfer.(7)
Extrachromosomal DNA consists three are plasmids, pSG1 (82 kb), pSG2 (27 kb), and pSG4 (11 kb).(4) pSG1, pSG2, and pSG4 all had putative RepA protein. Plasmid pSG1 codes for the production and transport of siderophores which aid in the acquisition of iron.(4) This CDS is located between transposase open reading frames (ORFs) indicating horizontal gene transfer.(4) Plasmid pSG1 also contains ORFs with homology to effectors, toxins, hemolysins, and protease fitness factors.(4) There were several regions with homology to conjugative transfer pilus genes suggesting that conjugation has been important for S. glossinidius.(10) Tra1 region on pSG1 had ORFs similar to transfer regions of plasmids. On the same plasmid is Tra2, but both regions were disrupted by mutation. On plasmid pSG2 was a region with homology to pilus ORFs.(10)
Cell structure and metabolism
Describe any interesting features and/or cell structures; how it gains energy; what important molecules it produces.
S. glossinidius is a Gram-negative and rod-shaped.(7) The lipopolysaccharide in this Gram-negative bacteria seems to be expressed without the O-antigen. This reduces the micro-organism's resistance to host immuniity, another indicator of the mutualistic relationship between host and symbiont.(5) Extrachromosomal DNA encodes siderophores for the acquisition of iron.(4) Though S. glossinidius is non-motile, there are 90 flagellar-related coding sequences on the chromosome. The first of two distinct clusters encodes a complete flagellar apparatus including motility proteins (MotA and MotB).(10) A second gene cluster codes for an incomplete flagellar apparatus.(10) Three phylogenetically distinct type-III secretion systems (TTSS) referred to as Sodalis symbiosis regions (SSRs) designated SSR-1, SSR-2, and SSR-3.(10) While core components of the needle structure were conserved, regions coding for some effectors or regulators have been modified or eliminated.(10)
The TTSS and flagellar components were expressed during early larval development. They are thought to allow S. glossinidius to enter host cells and ensure the establishment and maintenance of symbiosis in intra-uterine progeny.(10)
S. glossinidius has retained functional pathways for glycolysis, gluconeogenesis, the TCA cycle and the pentose phosphate pathway of free-living bacteria.(10) Many energy metabolism and carbon compound assimilation genes are missing in the genome. This may indicate an adaptation to the low concentration and lack of diversity of carbohydrates available due to vertebrate blood being the only nutrient of the tsetse host.(2,5) The pathways to synthesize all amino acids except alanine are present.(10) Most genes involved in energy production and conversion via anaerobic respiration have been lost. Which is expected in the constant microaerophilic environment of the tsetse midgut and haemolymph.(10) In its iron-rich environment, iron transport across the cytoplasmic membrane is achieved through active transport using an ABC transporter.(10) It also has an intact FeoA ferrous iron importer. In addition, plasmid pSG1 has a system of siderophores to transport and store iron.(4)
In vitro, strain M1T showed growth and acid production more vigorous when either N-acetyl-d-glucosamine or raffinose was provided as a carbon source than when utilizing glucose. S. glossinidius produces a-galactosidase (involved in raffinose catabolism) and b-N-acetylglucosaminidase (involved in N-acetyl-d-glucosamine catabolism). Polymerized N-acetyl-d-glucosamine(chitin) is present in the exoskeleton and gut peritrophic membrane of the tsetse fly, but the presence of raffinose is not known.(7)
Ecology
Describe any interactions with other organisms (included eukaryotes), contributions to the environment, effect on environment, etc.
When plated in lab, the beginning of the streak where the inoculum was heavy, produced many large colonies that merged. Toward the end of the streak, where innoculum was thinner, colonies were less abundant smaller colonies. (7) Like other microaerophilic bacteria, S. glossinidius has population-dependent growth where growth rate increases with total respiratory capacity. (7)
about if endosymbionts are gone, larvae cant develop
In the tsetse ¯y (Glossina spp.) P- and S-endosymbionts coexist in the gut lumen, with P-endosymbionts occupying specialized mycetocyte cells in the anterior portion of the insect gut and S-endosymbionts occupying midgut epithelial cells (Huebner & Davey, 1974; Pinnock & Hess, 1974). While the role of each micro-organism has not been clearly de®ned, collectively their presence is known to be essential for egg production and larval development in the insect (Nogge, 1981). Elimination of the bacterial endosymbionts with antibiotics, lysozyme and speci®c antibodies leads to reproductive abnormalities and growth retardation in the aposymbiotic host (Hill & Campbell, 1973; Nogge, 1976, 1978; Pinnock & Hess, 1974; Southwood et al., 1975). (7)
Sodalis glossinidius is one of three endosymbionts of the tsetse fly. [3]
The genome of entomopathogenic Photorhabdus encodes many adhesins, toxins, hemolysins, proteases, and lipases and contains a wide array of antibiotic synthesizing genes. These products are likely to play a role in the elimination of competitors, as well as in colonization, invasion, and degradation of the host insect cadaver (Duchaud et al. 2003). Because maternally transmitted Sodalis is a mutualist microbe with no known adverse impact on host biology, its chromosomally encoded TTSSs, hemolysin, lipases, and adhesions may fulfill functions different from those reported in pathogenic (10)
After hybridization to DNA macroarrays of the closely related Escherichia coli, it was found that its genome has a chitinase gene and pathogenicity island genes, both of which E. coli lacks.(2) In vitro, colonies of G. m. morsitans were thought to have been more susceptible to trypanosomes because of chitinase production from S. glossinidius.(7) TALK ABOUT PATHOGENIC ISLAND GENES FROM "DEGENERATIVE" ARTICLE. PURPOSE TO INVADE INSECT CELLS.
Pathology
How does this organism cause disease? Human, animal, plant hosts? Virulence factors, as well as patient symptoms.
UUUUUUSEEEEEE (8) Sodalis glossinidius is a mutualist microbe with no known adverse impact on host biology.(10) bacterial uptake due to protein secretory system [5] Because maternally transmitted Sodalis is a mutualist microbe with no known adverse impact on host biology, its chromosomally encoded TTSSs, hemolysin, lipases, and adhesions may fulfill functions different from those reported in pathogenic microbes. (10)
TTSS in 11 and 10
It has been difficult to study the functional role of the obligate endosymbionts in tsetse, as attempts to eliminate them have resulted in retarded growth of the insect and a decrease in egg production, preventing the aposymbiotic host from reproducing (19, 26, 32). The ability to reproduce, however, could be partially restored when the aposymbiotic flies were given a blood meal supplemented with B-complex vitamins (thiamine, pantothenic acid, pyridoxine, folic acid, and biotin), suggesting that the endosymbionts may play a role in metabolism that involves these compounds (25). While the functional significance of Sodalis is unknown, it has been implicated in the susceptibility of tsetse for trypanosome transmission (2)
Application to Biotechnology
Does this organism produce any useful compounds or enzymes? What are they and how are they used?
S. glossinidius has lost much of its biochemical profile.(7) Genetic material for compounds and enzymes have drifted into pseudogene status due to its specialized environment, the tsetse fly.(3) There are no known biotechnological benefits to society.
Current Research
Enter summaries of the most recent research here--at least three required
Some of the recent research on Sodalis glossinidius:
"The endosymbionts of tsetse flies: manipulating host–parasite interactions" [6]
References
1. "Sodalis glossinidius". NCBI Taxonomy Browser. 26 August 2007. [1]
2. Akman, L., Rio, R., Beard, C., and Aksoy, S. “Genome Size Determination and Coding Capacity of Sodalis glossinidius, an Enteric Symbiont of Tsetse Flies, as Revealed by Hybridization to Escherichia coli Gene Arrays.” Journal of Bacteriology. 2001. Volume 183.15 p. 4517-4525.[2]
3. Dale, C., Jones, T., and Pontes, M. "Degenerative Evolution and Functional Diversification of Type-III Secretion Systems in the Insect Endosymbiont Sodalis glossinidius." Molecular Biology and Evolution. 2005. Volume 22.3 p. 758-766. [3]
4. Darby, A., Lagnel, J., Matthew, C., Bourtzis, K., Maudlin, I., and Welburn, S. "Extrachromosomal DNA of the Symbiont Sodalis glossinidius." Journal of Bacteriology. 2005. Volume 187.14 p. 5003-5007. [4]
5. Thomson, N., Crossman, L., and Bentley, S. "Bacterial home from home." Nature Reviews Microbiology. 2006. Volume 4 p. 168-170. [5]
6. Dale, C., and Welburn, S. "The endosymbionts of tsetse flies: manipulating host–parasite interactions." International Journal for Parasitology. 2001. Volume 31.5-6 p. 627-630. [6]
7. Dale, C., and Maudlin, I. "Sodalis gen. nov. and Sodalis glossinidius sp. nov., a microaerophilic secondary endosymbiont of the tsetse fly Glossina morsitans morsitans." International Journal of Systematic Bacteriology. 1999. Volume 49 p. 267–275. [7]
8. Aksoy, S., and Rio, R. "Interactions among multiple genomes: Tsetse, its symbionts and trypanosomes." Insect Biochemistry and Molecular Biology. 2005. Volume 35.7 p. 691-698. [8]
9. Maudlin, I. "African trypanosomiasis." Annals of Tropical Medicine & Parasitology. 2006. Volume 11.8 p. 679–701. [9]
10. Toh, H., Weiss, B., Perkin, S., Yamashita, A., Oshima, K., Hattori, M., and Aksoy, S. "Massive genome erosion and functional adaptations provide insights into the symbiotic lifestyle of 'Sodalis glossinidius in the tsetse host." Genome Research. 2006. Volume 16 p. 149-156. [10]
11. Galán, J., and Wolf-Watz, H. "Protein delivery into eukaryotic cells by type III secretion machines." Nature. 2006. Volume 444 p. 567-573. [11]
Edited by Janet Melnyk, student of Rachel Larsen