Sputnik virophage
Classification
Viruses; unclassified virophages
Species
Sputnik Virophage
synonym: Acanthamoeba virus CD-2008
Description and Significance
The Sputnik virophage is a satellite virus of the giant mimivirus. In 2008, while studying Mamavirus, it was observed within the Mamavirus factory and Mamavirus capsids 4. Sputnik is 740Å in diameter, icosahedral in shape with a lipid bilayer 1. The discovery of the Sputnik virophage was actually what prompted the term “virophage” to exist, separating it from the other satellite viruses as it is the only satellite virus known to affect its hosts reproduction. Although there is some debate if the Sputnik virophage is merely a satellite virus or indeed a new class of virus deserving of a new term, the idea of a virus becoming the parasite of another was a new concept in the world of virology. A supporting point for those in favor of the term virophage, is that traditional satellite viruses use the host cell transcription machinery and then the other associated virus assists the satellite virus in virion assembly 7. As mentioned earlier, this is not the case for Sputnik, only the Acanthamoeba polyphaga mimivirus (APMV) virus factories are used for transcription, replication, and virion assembly, completely independent of the amoebae host's nucleus.
Genome Structure
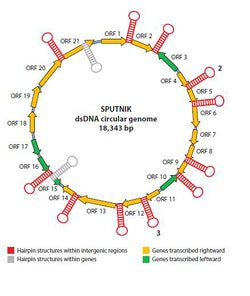
The sputnik virophage has a 18,343 nucleotide genome arranged consisting of double stranded DNA arranged in a single circular chromosome. It contains 21 predicted genes, of which eight have homologous sequences (Figure 1). Three of these have been compared to a putative intergrase complex, with homologues in both bacterial and archaeal genomes. Slightly larger than a human mitochondrial genome, it does not contain the necessary genes to replicate without co-infecting with Acanthamoeba polyphaga mimivirus which is postulated to contain the viral machinery that sputnik needs to replicate itself. Because of its novel chimeric genetic structure and replication cycle it has been placed into an as of yet unidentified family of viruses. 4
Cell Structure, Metabolism and Life Cycle
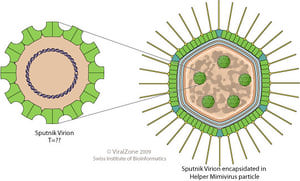
The Sputnik Virophage is an icosahedral, un-enveloped capsid with a diameter of 50 nm (740 Å). The triangulation number of the capsid is T=27 1 with a total of 1620 capsid proteins 2. There is a lipid membrane located beneath the capsid proteins. Studies have shown that the most abundant protein of Sputnik is the V20 protein which is hypothesized to self-assemble and compose the icosahedral capsid 3. Studies of the capsid proteins of Sputnik have identified the major capsid protein of the virophage, protein V20, to have two consecutive jelly roll domains. The double jelly roll domains (two eight-stranded antiparallel β-barrels) are organized into capsomers and then arranged in hexagonal arrays. The capsomers of Sputnik are made up of three monomeric double jelly roll domains that create a pseudohexagonal shape. Sputnik also has fiber-like structures on its surface that are hypothesized to help stabilize the individual capsomers or aid in host recognition and attachment 1.
Sputnik is a virophage of the APMV family and is able to infect the common host Acanthamoeba without association with its mimivirus host. However, viral lysis was not observed within seven days post-infection without the mimivirus host 4. The genome is composed of dsDNA, structural (capsid) protein and non-structural capsid proteins (DNA polymerase, genome packaging ATPase), however it is unable to produce its own viral factory within the Acanthamoeba hose and instead utilizes the giant viral machinery that is produced by APMV 5. Recent studies of Mimivirus have shown that Sputnik infection is not present when the Mimivirus is bald, so it thought that the fibers on the Mimivirus capsid are involved with adhesion of the virophage and host virus.6
Sputnik particles have been identified within the amoeba cell as soon as 30 minutes post-infection. Between 6 and 8 hours the Sputnik virions are observed in the amoeba, however there is a lag phase between Sputnik and Mimivirus production. Mimivirus is not observable within the cell until 8 hours post-infection. Co-infection of Mimivirus with the Sputnik Virophage is deleterious to Mimivirus production and results in production of abnormal Mimivirus particles. Side effects of co-infection that have been observed are an 11% increase in Mimivirus capsid thickness, multiple Mimivirus capsid layers and asymmetric accumulation of capsid fibrils. Co-infection with Sputnik decreases the yield of infectious mimivirus particles by 70% 4 and decreases cell lysis at 24 hours by 13% 6.
Ecology and Pathogenesis

The Sputnik virophage was first discovered in water from a cooling tower in Paris after inoculation of AMPV. The Sputnik virophage uses AMPV as a means to replicate and does not use any of the host’s machinery to replicate, making it a parasite of APMV (Figure 3). APMV is a large virus containing a linear 1,185-kilobase chromosome with the ability to encode its own “virus factory” and up to 900 proteins inside an amoeba host 4. After APMV infects an amoebae of the Acanthamoeba genus, the Sputnik virophage uses this virus factory to assist with its replication while hindering the production of properly functioning APMV virions, decreasing its ability to produce infective virions by as much as 70% 1.
The Sputnik virophage can be found where there are amoebae of the Acanthamoeba genus and where AMPV infects these amoebae. The AMPV virus needs to be present in the host for the Sputnik virophage to replicate, therefore the environment of both AMPV and the amoebae ultimately determine the location of the Sputnik virophage. The host amoebae are found in marine environments and moist soils and in water systems they are important members of the microbial community especially since they often inhabit biofilms 8. The AMPV virus was originally located in a water cooling tower in England 7.
The Sputnik virophage appears to play an important role in horizontal gene transfer between its hosts, giant viruses. It is thought that during the infection of AMPV and Sputnik virophage in an amoeba, horizontal gene transfer can occur. Due to its large genome and many proteins it is believed that AMPV can replicate unfamiliar DNA and incorporate it into its own genome; this could especially be the case with Sputnik virophage because it uses AMPV’s replication machinery 4. This would help explain why AMPV has genes of eukaryotic, prokaryotic, and unknown origin 7. Evolutionary genes of archaeal, mimvirus, and a novel origin comprise the Sputnik virophage itself. The Sputnik virophage contains an integrase gene, yet the origin of this is unclear 4. The Sputnik virophage is a conundrum of mix and matched genes that could allow insight into horizontal gene transfer between viruses and evolutionarily probably contribute to the environment greatly.
Although there is some thought that AMPV could be a human pathogen associated with pneumonia when infecting Legionella-like amoebae, there is no clear evidence for that APMV nor Sputnik virophage is associated directly to pneumonia or any other human aliment 7. Therefore, Sputnik virophage does not appear to be a threat to humans, but rather pathogenic to AMPV by decreasing its production of viable virions.
References
[3] - The UniProt Consortium. B4YNG0 (CAPSD_SPTNK). Universal Protein Knowledge Base. September 2011.
Author
Page authored by Patrick Ropp, Tori Roznowski, Rosalyn Schloemer, and Emily Schmitt-Matzen, students of Prof. Jay Lennon at Michigan State University.
