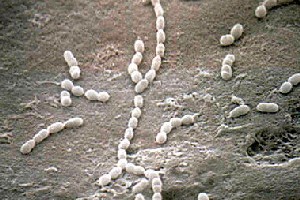
Streptococcus gordonii
A Microbial Biorealm page on the genus Streptococcus gordonii
Classification
Higher order taxa
Bacteria; Firmicutes; Lactobacillales; Streptococcaceae; Streptococcus
Species
|
NCBI: Taxonomy |
Streptococcus gordonii
Description and significance
The genus Streptococci are gram positive, mesophilic, nonmotile cocci that grow in pairs or bead like chains. Organisms within the genus comprise both pathogenic bacteria, such as S.pneumoniae and S. pyogenes, and non-pathogenic species that inhabit the mouth, skin, intestine and upper respiratory tract of humans including S. gordonii and S. mutans (1).
S. gordonii is part of the group viridians of Strepotococci, nonpathogenic commensal streptococci, which are integral members of the human oral flora. These organisms colonize tooth surfaces by creating biofilms in the human mouth, also known as dental plaque. S.gordonii plays an integral role in initiating colonization by creating surfaces for other colonizers to adhere to. Eventually dental biofilms lead to periodontal disease and dental cavities which are two of the most common diseases in developed nations (4).
S. gordonii also causes bacterial endocarditis by entering the blood stream usually after oral trauma. S. gordonii colonizes platelet-fibrin thrombi, blood clotting agents, in damaged heart valves or endocardium leading to damage and dysfunction of the heart valves. Endocarditis can be treated with antibiotic therapy and may cause death (5).
Genome structure
Although the full genome has not yet been determined, some interesting chromosomal regions have been noted. Glucosyltransferase (GTF) enzymes synthesize glucan polymers which are essential for adhesion to the tooth surface, Upstream from the GTF gene, gtfG, S. gordonii has a positive regulatory determinant, rgg, which codes for products that increase gtfG transcription. Similar determinants to rgg have been found in related bacteria like S. oralis and S. sanguis which are all pioneer colonizers in the human oral cavity. Rgg-like determinants also exist in other streptococci and lactococcal species which regulate various proteins with different functions. The data indicates that rgg-like genes are essential streptococcal regulatory determinants and the complete genome sequencing of S. gordonii may reveal further insight on the rgg determinant function (8).
Cell structure and metabolism
The human oral cavity provides a limited and varying source of nutrition for microbes inhabiting the oral micro flora (3). Oral streptococci, including S. gordonii, rely on sugars derived mainly from carbohydrates as an energy source. Fructose, a major component in the human diet, can be obtained via glucosyltransferases and from fructans via fructanases. Oral streptococci depend mainly upon the phosphotransferase system (PTS) to transport carbohydrates by means of phosphorylation and translocation through the membrane. The phosphoenolpyruvate-dependent PTS is the main pathway of transportation of sugars especially at low sugar concentrations (3).
The mammalian extracellular matrix is abundant in glycosaminoglycans (GAGs) consisting of recurring beta-linked dissaccharide units. When glycosaminoglycans are degraded, beta-linked disaccharides are released. S. gordonii ferments these beta-glucoside sugar substrates, including “cellobiose, arbutin, salicin and esculin,” to produce energy. In a recent study, beta-glucoside metabolism-encoding genes were expressed in Streptococcus gordonii colonization of saliva-coated hydroxyapatite (sHA) and impaired heart valves in rabbits (2). The beta-glucoside metabolism genes contained “a binding protein-dependent sugar transport and metabolism” and two phosphoenolpyruvate-dependent phosphotransferase systems (PTS). Several putative regulons contain these genes encoding for essential enzymes for metabolism. Bgl contains genes encoding subunits of a PTS enzyme II permease while bfr genes encode beta-glycosidase (BglF). Another regulon, esc, enables the metabolism of beta-glucoside esculin and oligochitosaccharides by containing genes encoding for regulation (EscR), transport (EscP), and metabolism (EscA) of PTS (2).
Additional PTS operons have been identified such as fruK, encoding for enzymes that phosphorylate fructose-1-phosphate to fructose 1,6-bisphosphate and fruR, encoding for catabolite reprossion. fruK and fruI are also thought to facilitate biofilm formation (3).
Ecology
Human teeth, which are non-shedding, moist and warm, are the perfect environment for biofilm development (2). In dental plaque, biofilm formation begins with pioneer organisms which attach to tooth surfaces in the human oral cavity. Streptococus gordonii is one of these pioneer organisms which initiate colonization and assist the further colonization of other organisms by creating a film on which bacteria may adhere to. Some of these later colonizers are periodontal pathogens such as Actinobacillus actinomycetemcomitans, P. gingivalis and Bacteroides forsythus (2).
In a recent study, S. gordonii was found to contain essential genes that facilitate the accrual of free floating P. gingivalis cells into the beginnings of a functioning biofilm. These genes are integral components in extracellular capsule biosynthesis, intercellular or intracellular signaling, biofilm architectural development and maintenance of adhesive proteins (7).
Initially, Streptococcus gordonii initiates colonization through formation of a monospecies biofilm. The human tooth is covered by pellicle containing lipids and proteins, including salivary agglutinin glycoprotein. The receptors for salivary agglutin glycoprotein located on S. gordonii and other pioneer colonizers recognize and bind to the pellicle (2). S. gordonii cells, bound to the surface of the tooth, then initiate a signal transduction pathway, known as BrfAB, which regulates adhesive activity. The S. gordonii monospecies biofilm then acts as a binding site for attachment of other more pathogenic organisms such as P. gingivalis by a process called coaggregation. Coaggregation is the process in which specific bacteria become interconnected by specific adhesions. Specifically, the long fimbriae (FimA) of P. gingivalis binds to glyceraldehydes-3-phosphate dehydrogenase (GAPDH) contained in the S. gordonii surface. The short fimbriae (Mfa) of P. gingivalis allows the cells to interact with the streptococcal SspA/B (antigen I/II) adhesions via an 80 amino acid binding epitope of SspA/B (BAR) (2).
Human volunteers who have introduced P. gingivalis into their mouths have shown that P. gingivalis is found solely in areas of streptococcal rich plaque. In in vitro studies, P. ginigivalis was also shown to coadhere with S. gordonii and this binding interaction promotes degradation of dentinal tubules by the otherwise non-dentin invasive P. gingivalis. Furthermore, biofilms between P. gingivalis and other streptococci, such as Streptococcus mutans and Streptococcus cistatus, are nonexistent. S. gordonii, therefore, may influence the constituents of oral biofilms by the specificity of adherence and signaling mechanisms (2).
Pathology
Although S. gordonii initiates dental plaque and the colonization of other pathogenic bacteria on tooth surfaces, S. gordonii is not directly pathogenic in the oral cavity. On the other hand, once S. gordonii enters the blood stream via oral bleeding it can colonize damaged heart valves causing endocarditis in humans (4). Blood platelets, cell fragments that facilitate blood clotting, bind to fibrinogen on the damaged heart valves and endocardium, the heart’s inner lining, and form platelet-fibrin thrombi. This platelet-fibrin thrombi can become colonized by S. gordonii causing damage to the heart valves and dysfunction of the heart (4).
During bleeding of the oral cavity more than seven hundred bacterial species may enter the blood stream although of these seven hundred, the oral streptococci are the most common causes of endocarditis (2). S. sanguis, S. oralis, and S. gordonii are the top three enodcarditis causing pathogens. It is curious that oral streptococci are efficient in binding to blood platelets especially since the blood stream is not their natural habitat. Recent studies have addressed this phenomenon and concluded that oral streptococci have adapted specialized mechanisms to recognize and bind with sialic-acid containg structures in the mouth, their natural habitat, which also allows for efficient interaction with platelet sialoglycoprotein GPI-alpha, located on the platelet membrane. In S. gordonii, Hsa and GspB proteins facilitate adhesion with both sialylated salivary molecules like mucin, MG2, as well as platelet surface proteins, GPI-apha and GPIIb. Clearly, an evolutionary adaptation in one habitat has allowed S. gordonii and other oral streptococci to invade another habitat (4).
Bacterial endocarditis occurs in humans who most often have artificial heart valves, heart disorders, or hypertrophic cariomyopathy (5). Dental surgery, oral trauma, urologic or gynecologic surgery, skin infections and intravenous drug use will increase the chances of endocarditis. Some symptoms of bacterial endocarditis include “fatigue, loss of appetite, night sweats, chills, headaches, joint discomfort, and tiny pinpoint-sized hemorrhages on the chest and back, fingers, or toes.” Treatment consists of intravenous antibiotic therapy and sometimes oral antibiotics for several weeks (5).
Current Research
Streptococcus gordonii is good candidate for a “live bacterial mucosal vaccine vector.” Since S. gordonii is a commensal bacteria whose natural habitat is the oral cavity, its use as a vaccine will ensure the safety of humans (6). The effieciency of S. gordonii in colonizing the oral cavity creates great potential for the stimulation of the mucosal immune system. S. gordonii can also be easily controlled genetically to produce a “fusion construct” that will enhance its stability. Another advantage is that heterologous antigens may be attached to the cell wall of S. gordonii thereby expressing a number of viral and bacterial antigens. Some examples of possible antigens are the E7 protein of human papillomavirus, the B subunit of Escherichia coli heat-labile toxin and the tetanus toxin fragment (6).
In a 2005 study of S. gordonii’s vaccine ability, researchers proved that the vector strain could be quickly eliminated with an antibiotic which is a necessary requirement of a functional vaccine should safety issues occur (6). S. gordonii strain SP204(1-1) was administered to the nose and mouth of 120 healthy subjects and cultures were detected by one day in 98% of the subjects. S. gordonii was cleared within one week of the treatment in 82% of the subjects. This temporary characteristic may be beneficial for a group A streptococcal vaccine strain since it would minimize the possibility of transmission to susceptible contacts and decrease constant low-level antigenic stimulation which may cause immunologic tolerance. The bacterial inoculum did not cause any major side-effects. The most common symptoms were mild headaches, nasal congestion and sore throat. Another advantage is that S. gordonii may deliver antigens directly to the mucosal surface thereby maximizing antigen-specific secretory immunoglobulins. S. gordonii is a very promising vaccine vector against respiratory pathogens although further research is need to determine the efficiency of stimulating the immune system and the survival rate of S. gordonii with specific antigens attached to their peptidoglycan cell wall (6).
In another current study, the transcriptional profiling using microarrays are used to determine the epithelial cell transcriptional responses to S. gordonii and more pathogenic oral organisms such as P. gingivalis and Aggregatibacter actinomycetemcomitans (9). It was found that the oral pathogens initiated vast changes in the gingival epithelial cell transcriptome which were specific to the pathogen species. For the commesal oral bacteria, S. gordonii and F. nucleatum, the transcriptional responses were largely different than the response produced by pathogenic oral bacteria. One transcriptome response was labeledl the HIGK response which included changes in biological pathways for cell development and morphogenesis. The HIGK response was significant when P. gingivalis was present while minimal in the presence of S. gordonii (9).
S. gordonii was found to repress proinflammatory cytokines. One hypothesis for the function of proinflammatory cytokine repression is that it may be beneficial to limit possible tissue destruction that a proinflammatory response would transpire. Additionally, it is hypothesized that commensal species can direct host cells to limit responses to more pathogenic organisms (9).
In another study, Glucotransferases (Gtfs), glucose producing enzymes, were tested for interaction with salivary amylase which may facilitate plaque formation. Salivary alpha-amylase was added to culture supernatants of S. gordonii (10). A protein complex of amylase, amylase-binding proteins (AbpB and AbpA) and glucotransferase precipitated from the solution which was labeled as rAbpA. Both amylase and rAbpB was shown to interact with Gtf-B, the glucotransferase produced by S. gordonii, by causing an increase in sucrase and transferase component activity of Gtf-B. This interaction was verified by enzyme-linked immunosorbent assay (ELISA). Several other oral streptococci including S. mitis, S. salivarius and S. parasanguinis are amylase-binding. Furthermore, amylase-binding bacteria only colonize oral cavities that contain amylase suggesting that amylase-binidng presents ecological afavantages for colonization (10).
References
(2) Kilic, A., Tao, L., Zhang, Y., Lei, Y., Khammanivog, A., Herzberg, M. "Involvement of Streptococcus gordonii Beta-Glucoside Metabolism Systems in Adhesion, Biofilm Formation, and In Vivo Gene Expression". Journal of Bacteriology. 2004. Volume 186. p. 4246-4253.
(3) Loo, C., Mitrakul, K., Voss, I., Hughes, C., Ganeshkumar, N. "Involvement of an Inductible Fructose Phosphotransferase Operon in Streptococcus gordonii Biofilm Formation". Journal of Bacteriology. 2003. Volume 185. p. 6241-6254.
(7) Kuboniwa, M., Tribble, G., James, C., Kilic, A., Tao, L., Herzberg, M., Shizukuishi, S., Lamont, R. "Streptococcus gordonii utilizes several distinct gene functions to recruit Porphyromonas gingivalis into a mixed community." Molecular Microbiology. 2006. Volume 60. p. 121-139.
(10)Chaudhuri, B., Rojek, J., Vickerman, M., Tanzer, J., Scannapieco, F. "Interaction of Salivary alpha-Amylase and Amylase-Binding-Protein A (AbpA) of Streptococcus gordonii with Glucosyltransferase of S. gordonii and Streptococcus mutans." BMC Microbiology. Volume 7. p. 60-65.
Edited by Tanya Budiarto [1] student of Rachel Larsen