Streptococcus pyogenes: Difference between revisions
No edit summary |
No edit summary |
||
| Line 1: | Line 1: | ||
{{Uncurated}} | |||
==Classification== | ==Classification== | ||
Revision as of 19:13, 19 August 2010
Classification
Kingdom: Bacteria Phylum: Firmicutes Class: Bacilli Order: Lactobacillales Family: Streptococcaceae Genus: Streptococcus Species: Pyogenes
Description and Significance
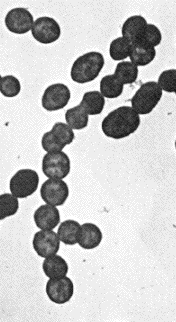
Streptococcus pyogenes is a gram-positive bacterium that usually grows in pairs or chains. It has been classified as a beta-hemolytic streptococcus because when cultured on a blood agar plate all the red blood cells are ruptured by the bacteria (1). Furthermore, it has been classified using Lancefield serotyping as group A, because it displays antigen A on its cell wall. Therefore, this bacterium is commonly called the beta-hemolytic group A streptococcus, or GAS. (2) A picture of this clearly shows how the organism grows.
Streptococcus pyogenes, also known as the flesh eating bacteria, is the most pathogenic bacterium in the whole genus (2). The name pyogenes comes from the word pyogenic, which is a classification for the streptococci that are associated with pus formation. The effects of this microbe range from mild illnesses such as strep throat and impetigo to more serious diseases such as scarlet fever, glomerulonephritis, and necrotizing fasciitis (3). However, if strep throat is untreated, it will lead to rheumatic fever, which is pretty rare now in the United States but was a more serious problem before the 20th century.
S. pyogenes usually begins infection on the surface of the skin or in the throat. From there, the bacterium begins to spread into deeper areas of the skin, which can potentially lead to life-threatening diseases (4). Because of its versatility in the human host, researchers have struggled developing a vaccine that would stop bacterial infection. This is why so much research is being done trying to understand the different cellular components of this organism as well as sequencing the genome. The more characteristics that are discovered and analyzed, the better the understanding we will have to fight this disease. Much progress has been made, because the genome for a strain has already been sequenced, and many conclusions have been drawn from the study.
Genome Structure
The genome of an M1 strain of Streptococcus pyogenes has been sequenced, and was found to contain 1,852,442 base pairs and about 1,752 predicted protein-encoding genes. (6) Also, more than 40 putative virulence associated genes have been identified, and this could explain why Streptococcus pyogenes is considered responsible for a wider variety of human disease than any other bacterial species. (6) GAS was found to produce a large number of extracellular proteins that increase the virulence of the organism by triggering a severe nonspecific immunological response in the human host (6).
The genome sequence was determined using the whole-genome shotgun approach. It used strain SF370, which was originally isolated from a patient with a wound infection. (6) The genome is a circular chromosome with an average G+C content of 38.5%, and 83% of the genes were transcribed in the clockwise direction and 76% of the genes were transcribed in the counterclockwise direction. (6) However, the replication termination site (ter) has still not been found, but it appears that the site is not 180 degrees from oriC. (6)
Cell Structure and Metabolism
One unique property of Streptococcus pyogenes is that it has a protein called protein F, which is a fibronectin binding protein that allows it to adhere to respiratory epithelial cells (7). This protein is an important virulence factor because by binding to the epithelial cells, the organism is able to stick to the cells of the host tightly, and not leave. Another characteristic of Streptococcus pyogenes is the M protein, which allows it to resist phagocytosis (8). The M protein has a fibrillar coiled-coil design, which “offers the organism several distinct advantages, ranging from antigenic variation to multiple functional domains” (8)
In addition, Streptococcus pyogenes is covered with an outer hyaluronic acid capsule (9). This capsule is required in order for the organism to be resistant to phagocytosis (5), which is vital in order for it to survive in its hosts. Finally, the “capsule may be an important adherence factor in the pharynx, since it binds CD44 on epithelial cells.” (10)
In another study, the regulation of anions such as Pi (inorganic phosphate) was examined in many different microorganisms. It was found that in Streptococcus pyogenes, “32Pi, uptake was catalyzed most rapidly in starved cells and was inhibited by addition of exogenous glycolytic and nonglycolytic energy sources such as glucose and arginine, respectively”. (11) This finding was very interesting because the mechanism of the regulation in S. pyogenes is actually the opposite in many other bacteria (11). The research reports two primary methods of regulation, the first being substrate depletion and the other being cellular ATP. This study is significant because phosphate is very important in regulating the control of many metabolic enzymes. For example, the phosphotransferase system uses a phosphate to transfer glucose into the bacteria by converting it to glucose-6-phosphate.
Ecology
All organisms interact uniquely in different environments or with other organisms. In a research study, it was found that within the same streptococcus genus, some species could actually inhibit the growth of other species. (12) The organism Streptococcus salivarius was identified to cause significant growth inhibition on Streptococcus pyogenes through a 22 amino acid lantibiotic (antibiotic peptide) designated salivaricin A. (12)
Another study that has been done was between the species Streptococcus pneumoniae and Streptococcus pyogenes. The study involved exploring the relationship of erythromycin resistance between the two species in different hospitals in Spain. (13) This is because more people have been using antibiotics in areas where there is a higher resistance of S. pneumoniae. (13) So, the researchers concluded that because the relationship between consumption and resistance is ecologically related, and both S. pneumoniae and S. pyogenes are respiratory pathogens, “if the rise in the consumption of antimicrobial agents increases resistance patterns for both S. pyogenes and S. pneumoniae, the prevalence of resistance in both species may be related.” (13) From the data, it was found that the two species have a very high correlation for resistance. Further studies must be done to draw conclusions, but this has been a step in the right direction in terms of determining how S. pyogenes interacts with other organisms.
Pathology
Streptococcus pyogenes is an unusually successful pathogen because of many different properties. First of all, the organism is able to adhere to the cells of its host with strong adhering mechanisms, which is important because the organism would be easily removed by mucus or salivary fluid. (5) The more adhesions there are, the stronger the adherence will be.
Not only does Streptococcus pyogenes adhere to its host cells, but it also invades them (5). Laboratories have shown that “group A streptococci have the potential to invade human epithelial cells at frequencies equal to or greater than classical intracellular bacterial pathogens.” (5) There are two proteins necessary for invasion, which are the M protein and the SfbI, which is a fibronectin-binding protein.
Streptococcus pyogenes affects its hosts in many different ways and causes a large range of diseases. The Center for Disease Control and Prevention (CDC) reports some symptoms of the diseases caused by Streptococcus pyogenes. They range from mild to severe, such as fever, severe pain, dizziness, and red rash at wound site. The symptoms at first might not seem severe, but in some cases the diseases have caused death to certain individuals.
Application to Biotechnology
So far, many of the enzymes released by Streptococcus pyogenes that have been studied are usually involved in how the organism interacts with the host immune system. For example, the cysteine protease SpeB is an enzyme that cleaves the immunoglobulin IgG, which allows S. pyogenes to fight against the host immune system (14). Another popular immunomodulating enzyme studied is C5a peptidase, ScpA, which is a serine endopeptidase that “cleaves the specific complement factor C5a, [and this] inhibits recruitment of phagocytic cells to the infectious site.” (14) Since these enzymes all modulate the immune system of the host and are used to help the organism survive, they are most likely not very useful in the field of biotechnology.
However, there has been a patent on two specific proteins that have been found to prevent diseases caused by Streptococcus pyogenes. The two proteins are “designated Cpa1 and Cpa49 which are obtained from the collagen binding region in Streptococcus pyogenes. The isolated proteins of the present invention have been observed to bind to collagen, and thus can be utilized in methods of treating or preventing streptococcal infection through the inhibition of the ability of the bacteria to bind to collagen.” (15) This is a perfect example of how a compound extracted from an organism can be used to fight against itself and protect victims from invasion.
Current Research
Much of the research on Streptococcus pyogenes relate to why it is able to fight against the human immune system so well. It has been a very hot field and the studies have shown progression over the years. A lot has been discovered, but it is still far from the amount of knowledge needed to fight against infections.
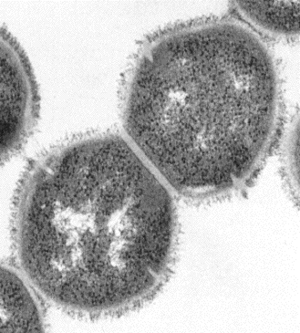
In 2005, there was a research report that showed that Group A Streptococcus (GAS) actually have pili. This finding is unusual because normally only Gram-Negative bacteria have pili. Prior to this, the cell surface of Gram-positive bacteria was rarely studied, and not much was known about them. (16) The finding showed that Streptococcus pyogenes showed the presence of “long, surface-exposed, pilus-like structures composed of members of a family of extracellular matrix-binding proteins.” (16) In Gram-negative bacteria, pili are important virulence factors because they are used to attach to target cells. The results of this study showed that the pili in GAS are composed of the same family proteins, which suggests that they are virulence factors in Gram-positive bacteria as well. (16) Furthermore, the pili were also found to have effective antigens that will protect against lethal infections. (16)
In 2006, an article reported that another reason for the failure of antibiotics against Streptococcus pyogenes could be because of biofilm formation. To test this, 289 Streptococcus pyogenes strains were isolated and their abilities to form biofilms were analyzed. (17) The experiment showed that most of the strains (90%) were able to form biofilms. The data also “indicated that biofilm-forming isolates entered epithelial cells with significantly lower efficiency than biofilm-negative strains.” (17) This suggests that if a certain strain could not enter epithelial cells to escape antibiotics, it would instead form biofilms as a means to survive in its host. (17) The conclusions of this research are very important because it shows that Streptococcus pyogeneshas alternative methods of surviving in its host. (17)
In February, a paper was published that addressed one of the hottest topics related to Streptococcus pyogenes. This topic is the FCT genome region, which stands for fibronectin-collagen-T-antigen. This region is the region that allows Streptococcus pyogenes to adhere to epithelial cells of its host, as described earlier. The paper describes many of the research that has been done on this region, and the conclusions that have been made. The adhesion has been determined to be protein F1 or F2, and a binding protein called Cpa was also found. (18) The paper predicts that “Cpa or protein F1 primes/facilitates skin or throat infections, respectively, although the presence of some skin or throat strains without the cpa or prtF1 genes indicated the existence of alternative infection strategies.” (18) The research done was mostly on the genetic linkage between different strains, and from the results “decide which combination of markers shows the strongest association to the clinical impact of the S. pyogenes isolates.” (18) As research continues to advance, more and more is being understood about how Streptococcus pyogenes invades and affects humans. Hopefully, solutions to this organism will be found soon.
References
Classification taken from NCBI, http://www.ebi.ac.uk/newt/display?search=160490
1. NIH Fact Sheet "Group A Streptococcal Infections", March 1999.
2. Todar, Kenneth. Todar’s online textbook of Bacteriology, University of Wisconsin-Madison Department of Bacteriology, 2002
3. Prescott, Harley and Klein. Microbiology, Sixth Edition. McGraw Hill Higher Education, NY, 2005
4. Facklam, R. What happened to the streptococci: overview of taxonomic and nomenclature changes. Clinical Microbiology Reviews, v.15, 2002 5. Cunningham, M. W. Pathogenesis of group A streptococcal infections. Clinical Microbiology Reviews, v.13, Jul 2000 6. J.J. Ferretti et al, 2001. Complete Genome Sequence of an M1 Strain of Streptococcus pyogenes Proc. Natl. Acad. Sci. USA. 98, 4658-4663
7. Hanski, E., P. A. Horwitz, and M. G. Caparon. 1992. Expression of protein F, the fibronectin-binding protein of Streptococcus pyogenes JRS4, in heterologous streptococcal and enterococcal strains promotes their adherence to respiratory epithelial cells. Infect. Immun. 60:5119-5125
8. Fischetti, V. A. 1989. Streptococcal M protein: molecular design and biological behavior. Clin. Microbiol. Rev. 2:285-314
9. Kendall F E, Heidelberger M, Dawson M N. 1941. A serologically inactive polysaccharide elaborated by mucoid strains of group A hemolytic streptococcus. J Biol Chem.118:61–69
10. Schrager H M, Alberti S, Cywes C, Dougherty G J, Wessels M R. 1998. Hyaluronic acid capsule modulates M protein-mediated adherence and acts as a ligand for attachment of group A streptococcus to CD44 on human keratinocytes. J Clin Investig. 101:1708–1716
11. Reizer J, Saier MH Jr. 1987. Mechanism and regulation of phosphate transport in Streptococcus pyogenes. J Bacteriol. 169(1):297–302.
12. Upton, M., J. R. Tagg, P. Wescombe, and H. F. Jenkinson. 2001. Intra- and interspecies signaling between Streptococcus salivarius and Streptococcus pyogenes mediated by SalA and SalA1 lantibiotic peptides. J. Bacteriol. 183:3931-3938
13. Gomez-Lus, R., J. J. Granizo, L. Aguilar, E. Bouza, A. Gutierrez, and J. Garcia-de-Lomas. 1999. Is there an ecological relationship between rates of antibiotic resistance of species of the genus Streptococcus? The Spanish Surveillance Group for Respiratory Pathogens. J. Clin. Microbiol. 37:3384-3386
14. Collin, M., and Olsen, A. 2003 Extracellular Enzymes with Immunomodulating Activities: Variations on a Theme in Streptococcus pyogenes, Infecion and Immunity. 71, 2983-2992
15. Podbielski, Andreas. 2004. Collagen-binding proteins from streptococcus pyogenes. PatentStorm, US Patent 6777547
16. Mora, M., G. Bensi, S. Capo, F. Falugi, C. Zingaretti, A. G. Manetti, T. Maggi, A. R. Taddei, G. Grandi, and J. L. Telford. 2005. Group A streptococcus produce pilus-like structures containing protective antigens and Lancefield T antigens. Proc. Natl. Acad. Sci. USA 102:15641-15646
17. Baldassarri et al. 2006, Therapeutic Failures of Antibiotics Used To Treat Macrolide-Susceptible Streptococcus pyogenes Infections May Be Due to Biofilm Formation, J Clin Microbiol. 44(8): 2721–2727
18. Podbielksi, Andreas. 2007. Flexible Architecture of Streptococcus pyogenes FCT Genome Region: Finally the clue for Understanding Purulent Skin Diseases and Long Term persistence, J. Bacteriol. 189: 1181-1184
KMG