Suppression of Peptidoglycan Remodeling
By [Frank Zhao]
Introduction
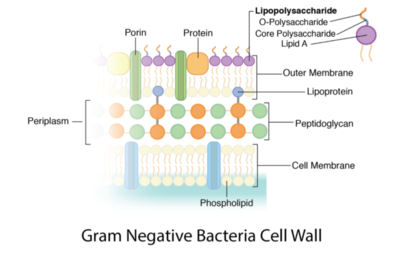
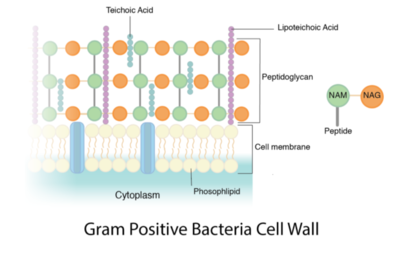
Peptidoglycan is a polymer made of sugars and amino acids that forms a layer outside the plasma membrane of bacterial cells. Peptidoglycans are cross-linked by peptides, serving as the cell wall that protects bacteria from the environment. Peptidoglycan limits the volume of the cell and thus it generates turgor pressure as the water rushes into the cell.[1] Gram-positive bacteria obtain a thicker peptidoglycan layer than Gram-negative bacteria but this layer is crucial to the survival of both types of bacteria.[2]
Many antibiotics target this component to kill the bacteria. For example, penicillin binds to penicillin-binding proteins and inhibits the synthesis of peptidoglycan, weakening the cell wall of bacteria. It has been discovered that a new mechanism could be used by antibiotics on peptidoglycan— blocking the action of autolysins, a peptidoglycan hydrolase that is essential for remodeling the bacterial cell wall during growth.[3] Studying how this new mechanism works could give us insight into developing new antibiotics against resistant strains.
Structure and Function
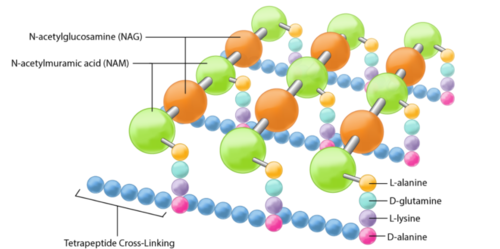
The peptidoglycan layer is located in the bacterial cell wall, consisting of chains of two alternating amino sugars, N-acetylglucosamine (an amide derivative of the monosaccharide glucose) and N-acetylmuramic acid (a combination of N-acetylglucosamine and phosphoenolpyruvate). The alternating amino sugars are linked by Teichoic acids, which are chains of phosphodiester-linked glycerol or ribitol.[1] Parallel chains of amino sugars can also be connected by cross-bridges made of extensions of peptides to form tetrapeptides, with m-Diaminopimelic acid binding D-Alanine.[4]
Peptidoglycan sacculi cross-linked by peptides form a closed, sac-shaped structure that surrounds the cytoplasmic membrane. Peptidoglycan sacculi have been purified from Escherichia coli by incubation with boiling 4% sodium dodecyl sulfate.[5] While a bacterial cell contains a significant number of different kinds of molecules, there is only one peptidoglycan molecule in each bacterial cell, which contributes to 0.8% of the mass of the whole cell.[1]
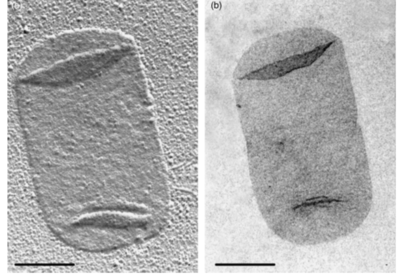
The content of peptidoglycan sacculi could be different among bacterial species. In Gram-positive bacteria, such as Staphylococcus aureus, the tetrapeptide cross-bridges are made of L-alanine, D-glutamine, L-lysine, and D-alanine. In Gram-negative bacteria, on the other hand, the cross-bridges are constituted of L-alanine, D-glutamic acid, m-diaminopimelic acid, and D-alanine. Gram-positive bacteria typically have many layers (3-20) of peptidoglycan chains outside the cytoplasmic membrane, whereas Gram-negative bacteria often have one or two layers of peptidoglycan between the inner and outer membrane.[1]
Peptidoglycan plays a crucial role in the survival and reproduction of bacteria. It is the main component of the bacterial cell wall, forming a strong net-like structure that maintains the shape of the cell. Since bacteria are unicellular organisms, it is extremely important to protect the only cell from the outside environment. When the bacteria experience a significant osmotic pressure difference across the cell membrane, the cell envelope could prevent the cell from shrinking or expanding too much so that it lyses and dies.[6]
Biosynthesis
Include some current research, with at least one figure showing data.
Inhibitive Antibiotics Targeting Peptidoglycan

Corbomycin and Complestatin
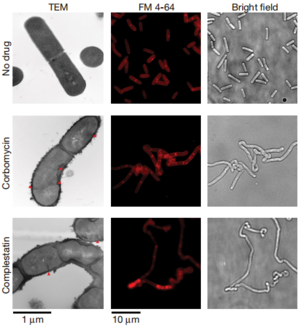
Conclusion
References
- ↑ 1.0 1.1 1.2 1.3 Slonczewski, J., and Foster J. W.. Microbiology: An Evolving Science. New York
- ↑ Anon. n.d. “Bacteria.” Basic Biology. Retrieved April 8, 2021
- ↑ [1] Culp, Elizabeth J., Nicholas Waglechner, Wenliang Wang, Aline A. Fiebig-Comyn, Yen-Pang Hsu, Kalinka Koteva, David Sychantha, Brian K. Coombes, Michael S. Van Nieuwenhze, Yves V. Brun, and Gerard D. Wright. 2020. “Evolution-Guided Discovery of Antibiotics That Inhibit Peptidoglycan Remodelling.” Nature 578(7796):582–87. doi: 10.1038/s41586-020-1990-9.
- ↑ [2] Bruslind, Linda. n.d. “Bacteria: Cell Walls.” in General Microbiology. Oregon State University.
- ↑ [3] Vollmer, Waldemar, Didier Blanot, and Miguel A. de Pedro. 2008. “Peptidoglycan Structure and Architecture.” FEMS Microbiology Reviews 32(2):149–67. doi: 10.1111/j.1574-6976.2007.00094.x.
- ↑ [] Al-Nabulsi, Anas A., Tareq M. Osaili, Reyad R. Shaker, Amin N. Olaimat, Ziad W. Jaradat, Noor A. Zain Elabedeen, and Richard A. Holley. 2015. “Effects of Osmotic Pressure, Acid, or Cold Stresses on Antibiotic Susceptibility of Listeria Monocytogenes.” Food Microbiology 46:154–60. doi: 10.1016/j.fm.2014.07.015.
Authored for BIOL 238 Microbiology, taught by Joan Slonczewski, 2021, Kenyon College.
