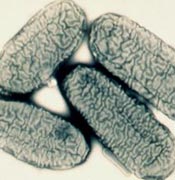
Synergistes jonesii
Classification
Kingdom: Bacteria
Phylum: Synergistetes
Class: Synergistia
Order: Synergistales
Family: Synergistaceae
Genus: Synergistes
Species: S. jonesii
Description
Synergistes jonesii is a bacteria that was originally isolated from the rumen of a Hawaiian goat by Dr. Raymond J. Jones, after whom it was named. It is characterized as a gram-negative vibrio, or rod-shaped, bacterium and is known for being 0.6-0.8µm in diameter and 1.2-1.8µm in length. Given its habitat within the rumen it is considered a thermophilic bacteria, which follows its categorization in the sub-section of Synergistetes: Thermanaerovibrio. Also attributed to its habitat is the fact that S. jonesii is anaerobic. Another important aspect is that it contains no cili or flagella making it a non-motile bacteria. One of the most intriguing facets of S. jonesii is its ability to degrade toxic pyridinediols that would otherwise harm the ruminant host. S. jonesii contains high hydrogenase activity which allows for the anaerobic degredation of pyrindinediols when also in the presence of methyl viologen or in the presence of alpha-ketoacids under hydrogen or nitrogen gas.
Genome Structure
Not much information has been gathered on the genome structure of Synergistes jonesii, aside from what can be deduced from observed metabolic processes. One of the major characteristics is that the genome contains genes that code for proteins that allow for the catabolism of arginie and histidine. The genome of S. jonesii is comprised of a 58% GC content.
Related Bacterial Species
Synergistes jonesii is in a sub-section of Synergistetes known as Thermanaerovibrio. This classification means that the bacteria under this sub-section are anaerobes that live in warm environments, and have vibrio, or rod-like, shapes. In addition to Synergistes jonesii, this sub-section of Synergistetes includes Thermanaerovibrio velox, Thermoanaerovibrio acidaminovorans, and Cloacibacillus evryensis. Like S. jonesii, C. evryensis and T. acidaminovorans degrade amino acids. [1] [2] [3]
Metabolism
Synergistes jonesii is a chemoorganotroph which relies on organic chemicals for its source of energy and carbon. Because S. jonesii is strictly anaerobic, the primary metabolic pathway of this microbe is fermentation. Pyridinediols, arginine, and histidine are metabolized by the microbe for energy and carbon. Arginine is metabolized using the arginine deaminase pathway and it is still uncertain as to how histidine is degraded by the cell. When arginine is metabolized by the arginine deaminase pathways, its products are carbon dioxide, acetate, butyrate, citrulline, and ornithine. When histidine is metabolized, the products are carbon dioxide, acetate, butyrate, citrulline, and ornithine, and indirectly formate and propionate.
Energy
Unfortunately, there is still quite a bit that is unknown about the nutritional requirements for Synergistes jonesii outside of the fact that this bacteria is capable of of utilizing the amino acids arginine and histidine for growth. Arginine and histidine were found to be the primary source of energy for the S. jonesii.
Nitrogen
There is still a lot that is unknown about the primary source of nitrogen for Synergistes jonesii. What is known about the miroorganism is that it does not reduce nitrate and therefore does not obtain its nitrogen from this source. The most likely source of the Nitrogen that this microbe utilizes is from the metabolizing arginine and histidine. When both of these substrates are metabolized, ammonia is one of the products of the pathways.
Carbon
Synergistes jonesii is unique in that it is currently the only know rumen microorganism to utilize arginine and histidine as a major or source of energy and carbon yielding substrates. A third amino acid that S. jonesii seemed capable of metabolizing was glycine, although to a lesser extent from arginine and histidine. Arginine was found to be degraded in S. jonesii by the arginine deaminase pathway and produced carbon dioxide, acetate, butyrate, citrulline, and ornithine. Histidine metabolism also produced carbon dioxide, acetate, butyrate, citrulline, and ornithine, as well as providing carbon for formate and propionate. Due to the fact that the amino acids arginine and histidine are the major source of carbon for Synergistes jonesii, it may allow the microorganism to compete in the rumen and occupy an individualized niche within the community.
Ecology
The only known habitat for Synergistes jonesii is within the rumen. This microbe was first isolated from the rumen of a goat in Hawaii and has since been found in the rumen of cattle. S. jonesii is not a common microbe is ruminant populations, but some geographic distributions have been shown.
References
(1) Data extracted from the "16S rRNA-based LTP release 111 (full tree)". http://www.arb-silva.de/fileadmin/silva_databases/living_tree/LTP_release_111/LTPs111_SSU_tree.pdf
(2) "Thermanaerovibrio". Bacterio.net. http://www.bacterio.net/thermanaerovibrio.html
(3) "Cloacibacillus". Bacterio.net. http://www.bacterio.net/cloacibacillus.html
(4) "Amino acid utilization by the ruminal bacterium Synergistes jonesii strain 78-1". SpringerLink. http://link.springer.com/article/10.1007/BF00250272. Accessed 20 April 2014.
(5) Rincón, M.T., M.J. Allison, F. Michelangeli, Y. De Sanctis, and M.G.Domínguez-Bello. 1998. "Anaerobic degradation of mimosine-derived hydroxypyridines by cell free extracts of the rumen bacterium Synergistes jonesii." FEMS Microbiology Ecology 27: 127-132. http://onlinelibrary.wiley.com/doi/10.1111/j.1574-6941.1998.tb00530.x/pdf. Accessed 20 April 2014.
(6) "Corrigendum". ac.els-cdn.com. http://ac.els-cdn.com/S0723202011802626/1-s2.0-S0723202011802626-main.pdf?_tid=040cfe92-c8e7-11e3-bd6c-00000aacb360&acdnat=1398038208_0ffc7eba11d04e6f73321fd893cc4472. Accessed 20 April 2014.